Potri.013G032100 [POPLAR]

| External link |
|
|||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.013G032100.1 pacid=42811399 polypeptide=Potri.013G032100.1.p locus=Potri.013G032100 ID=Potri.013G032100.1.v4.1 annot-version=v4.1
ATGGAAGAGAGTAATAGCAATAACCACATAGCAAACAATGTGGCTCGAGCAATTGCTGCTGCTCTTGACTGGAACTCTACTCCTGATGCTCGCAAAGCTG
CTGTGTCTTTCCTTGAATCCATCAAAGCAGGAGATGTACGAATTTTGGCCAGCTCATCATTCGTTCTTGTCAAAAAGGACTGGTCTTCTGAAATTCGGTT
GCATGCTTTCAAAATGCTGCAGCATTTGGTACGGTTGCGATGGGAGGAATTGAGCCCAACAGAACGCAGGAACTTTGCGAATGCTGCTGTTGAGTTGATG
GCTGAAATTGCAAATTCCTGTGAAGAATGGGTGCTGAAAAGTCAGACAGCTGCCCTTGTTGCTGAGATTGTCAGAAGAGAAGGCTTAGAACTGTGGAAAG
AACTGCTTCCATCTCTGGTTTCCCTATCTAGTCAGGGCCCCATACAAGCTGAGTTGGTTTCGATGACACTGAGGTGGCTTCCTGAAGATATTACAGTTCA
CAATGAAGATTTAGAAGGTGATCGGCGTAGGTTATTGTTGCGAGGGCTTACTCAGTCATTGCCTGAAATGCTGCCACTACTATACACCTTACTTGAGAGA
CATTTTGGAGCTGCATTGAGCGAAGCAGGGAGGCAGCAACTGGACATCGCAAAACAGCATGCAGCAACAGTAACAGCGACTTTGAATGCTGTCAACGCTT
ATGCTGAGTGGGCGCCTTTGCAGGATCTAGCCAAATATGGCATAATTTATGGCTGTGGTGTCATGCTGTCTTCTCCTGACTTTCGTCTTCATGCCTGTGA
ATTTTTCAAACTCGTCTCACAAAGGAAGAGACCTGCCGATGCCTCTGCTTCTGAATTTGATTCTGCAATGCGTAATATCTTTCAGATCATGATGAATGTC
TCAAGAGATATTTTGTACAAAACTGTCTCAAGTGCTGGAGTTATGGATGAGAGTGAATTTGAGTTTGCAGAATATATATGCGAGAGTATGGTGTCTTTGG
GTTCTTTTAACTTTCAATGCATTTCTGGTGACAACACTATACTCTCTCTTTATTTACAGCAGATGCTGGGCTTTTTTCAACATTTTAAGCTAGCTCTTCA
TTATCAATCCCTGCTATTTTGGCTGGTGCTAATGAGAGATTTAATGTCAAAGCCGAAAGTTACTGCATATTCAGCAGACGGGTCTGCTTTTAACAGTGCA
GGCTCTAGTTCTGGACAGGTTGATGATGAAAAAAGGAGGACTTTAAGCCTTGTGGATGATGATATTTGTGTTGTAATTCTGGATATCTCTTTCCAGCGCT
TGCTTAAGAAAGAAAAGGTTTTCTCTGGGAATTCCTTTTCTCCTGGAACATTGGAGTTGTGGAGTGATGATTTTGAAGGCAAGGGTGACTTTGGCCAATA
TCGTTCCAAGCTGACCGAGCTGATGAGACTTGTTGCTTCTTTTAAGCCCCTAATAGCTGGTGCTAAAATCTCTGAAAGAATTCTTTCAATCATTAAGAGC
ATTCCAAATTCTCAAATACCTGTTCAGGACTTAGCTGTGATGGAAAGCATGCAAGTGGCTTTAGAAAATGTTGTCAATGCAGTTTTTGATGGATCAAATG
GCTATGCGGCGGTCAGTTCGGAAGTTCACCTAGCGTTGTGTAGAGTTTTTGAAGATTTACTTCAGCAACTCCTTTCTTTGAAATGGACGGAGCCAACTCT
TGTGGAAATCCTAGGGCACTATTTAGACGCGCTGGGTCCATTTTTGAAGTATTTTCCAGATGCGGTTGGTGGTGTCATTAATAAATTATTCGAGCTTCTA
ATGTCAATCCCATTTGTTGTTAAGGATCCTTCAGTAAGCAGTGCACGCCATGCAAGGTTACAGATTTGTACGTCATTTATTCGAATAGCCAAATCTGCTG
ACAAAAGTGTTCTACCTCATATGAAAGGAATAGCTGATACCATGGCATATATGCAAAGAGAGGGCTCTCTGCTTCGTGGTGAACATAATCTGCTAGGTGA
AGCATTTCTTGTTATGGCTTCAGCTGCCGGGACTCAACAACAGCAAGAAGTTTTGGCCTGGTTGCTTGAACCTTTGAGCCAACAGTGGACACAGCTAGAG
TGGCAAAATAATTATTTGTCTGAGCCACTAGGTCTGATCCGTTTGTGCTCTGAGACAGCATTCATGTGGTCAATTTTCCACACTGTGACATTCTTTGAGA
AGGCACTTAAGAGGAGTGGAATCAGAAAAGGCAGTCTGAATTTGCAAAGCATCTCAACAGCAAGTACTATACATCCAATGGCTTCTCATCTGTCCTGGAT
GCTGCCACCCCTCTTGAAACTTCTTCGTGCTGTGCACTCACTTTGGTCTGCATCCATATCTCAAATGTTACCTGGAGATATCAAAGCTGCCATGACCATG
GGCAATGCTGAACGATATAGTCTCCTTGGGGAAGGAAACCCCAAATTGTCCAAAGGTTCCTTAAACTTCATAGATGGATCTCATATTGATACAAGTAGGG
AAGGACACACCGAAACAAATGAAGCTGATATACGGAATTGGTTGAAAGGTATCAGAGATAGTGGATACAATGTATTGGGCCTATCGATGACCATTGGTGA
TCCATTTTTTAAATGTTTGGATGTTCATTCTGTTGGTGTGGCACTACTGGAAAACATACAGTCAATGGAGTTCAGACATACAAGGCAGCTTGTCCATTCA
GCTTTGATACCTTTGGTTAAACATTGTCCTATGGAAATGTGGGAGGTCTGGCTGGAAAAGCTCTTGCATCCATTATTTATCCATGTACAGCAAGCTCTTA
CCTTTTCTTGGTCCAGTCTCCTGCATGAGGGTAAAGCAAAGGTCCCAGATGTCCTTGGCATCCTTGCTGAGGCAGACTTGAAAGCAGAAGTAATGGAAGA
GAAACTGCTTCGAGACCTAACTCGTGAGATGTGCGTTCTTCTCTCTACAATAGCTTCGCCCGGACTAAATACCGGGCTTCCTACTTTAGAACAATCTGGC
CATGCTATCCGTGTGGATGCATCTTCTTTGAAGGAATTGGACGCATTTGCATCAAATTCTATGGTTGGTTTTCTTTTGAAGCACAATGGTTTGGCTGTTC
CAGCATTACAGATTTGTTTAGAAGCTTTCACATGGACAGATGGTGAAGCAGTATCAAAAGTTTTGTCCTTTTGTGCTAGTGTGATTCTTCTAGCAATCTC
AGCTAATAATGTTCAACTCAGAGAGTTTGTTTCTAAAGATCTGTTCTCTGCAATTATCAAAGGTTTGGCGCTCGAGTCAAATGCATTTATCAGTGCTGAT
CTAGTTGGTTTCTGTCGAGAAATATTCATGCATCTCTGTGATAGAGATCCAGCTCCAAGGCAGGTTCTGCTTTCTCTCCCTTGTATTAAACCCCAGGATT
TGGTTGCCTTTGAAGAAGCTTTGACCAAGACAGCTAGCCCTAAGGAACAAAAGCAACATATGAAGAGCTTGCTTTTATTAGCTACTGGAAACATGCTAAA
AGCGCTAGCTGCTCAGAAGAGTGTAAACATTATCACAAATGTTACAATGAGGCCTCGCAGCTCGGTAAATGCTCCAGAAACCAGAATTGATGAGGGGGAT
ACTATAGGATTGGCAGCCATTTTGTGA
|
|||||||||||||||||||||||||
|
AA sequence
|
>Potri.013G032100.1 pacid=42811399 polypeptide=Potri.013G032100.1.p locus=Potri.013G032100 ID=Potri.013G032100.1.v4.1 annot-version=v4.1
MEESNSNNHIANNVARAIAAALDWNSTPDARKAAVSFLESIKAGDVRILASSSFVLVKKDWSSEIRLHAFKMLQHLVRLRWEELSPTERRNFANAAVELM
AEIANSCEEWVLKSQTAALVAEIVRREGLELWKELLPSLVSLSSQGPIQAELVSMTLRWLPEDITVHNEDLEGDRRRLLLRGLTQSLPEMLPLLYTLLER
HFGAALSEAGRQQLDIAKQHAATVTATLNAVNAYAEWAPLQDLAKYGIIYGCGVMLSSPDFRLHACEFFKLVSQRKRPADASASEFDSAMRNIFQIMMNV
SRDILYKTVSSAGVMDESEFEFAEYICESMVSLGSFNFQCISGDNTILSLYLQQMLGFFQHFKLALHYQSLLFWLVLMRDLMSKPKVTAYSADGSAFNSA
GSSSGQVDDEKRRTLSLVDDDICVVILDISFQRLLKKEKVFSGNSFSPGTLELWSDDFEGKGDFGQYRSKLTELMRLVASFKPLIAGAKISERILSIIKS
IPNSQIPVQDLAVMESMQVALENVVNAVFDGSNGYAAVSSEVHLALCRVFEDLLQQLLSLKWTEPTLVEILGHYLDALGPFLKYFPDAVGGVINKLFELL
MSIPFVVKDPSVSSARHARLQICTSFIRIAKSADKSVLPHMKGIADTMAYMQREGSLLRGEHNLLGEAFLVMASAAGTQQQQEVLAWLLEPLSQQWTQLE
WQNNYLSEPLGLIRLCSETAFMWSIFHTVTFFEKALKRSGIRKGSLNLQSISTASTIHPMASHLSWMLPPLLKLLRAVHSLWSASISQMLPGDIKAAMTM
GNAERYSLLGEGNPKLSKGSLNFIDGSHIDTSREGHTETNEADIRNWLKGIRDSGYNVLGLSMTIGDPFFKCLDVHSVGVALLENIQSMEFRHTRQLVHS
ALIPLVKHCPMEMWEVWLEKLLHPLFIHVQQALTFSWSSLLHEGKAKVPDVLGILAEADLKAEVMEEKLLRDLTREMCVLLSTIASPGLNTGLPTLEQSG
HAIRVDASSLKELDAFASNSMVGFLLKHNGLAVPALQICLEAFTWTDGEAVSKVLSFCASVILLAISANNVQLREFVSKDLFSAIIKGLALESNAFISAD
LVGFCREIFMHLCDRDPAPRQVLLSLPCIKPQDLVAFEEALTKTASPKEQKQHMKSLLLLATGNMLKALAAQKSVNIITNVTMRPRSSVNAPETRIDEGD
TIGLAAIL
|
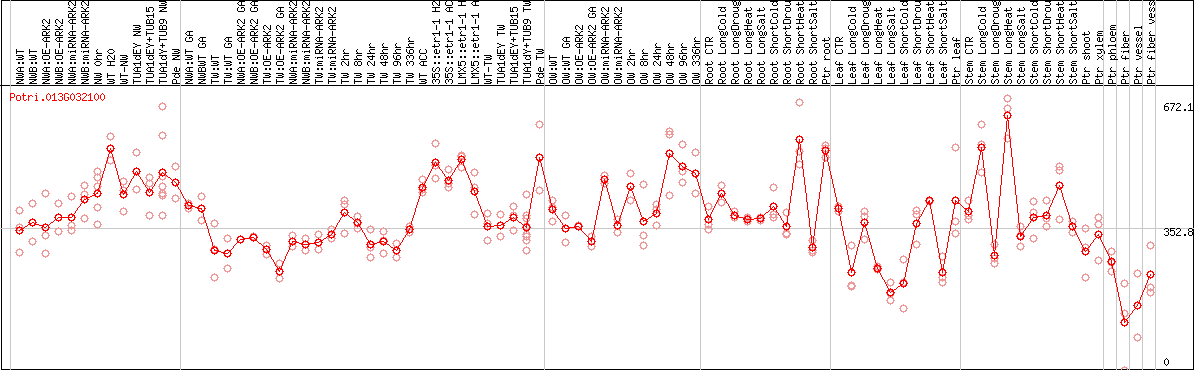
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.013G032100 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
