XRN4.2 (Potri.013G035600) [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | XRN4.2 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.013G035600.1 pacid=42810982 polypeptide=Potri.013G035600.1.p locus=Potri.013G035600 ID=Potri.013G035600.1.v4.1 annot-version=v4.1
ATGGGTGTACCGGCGTTCTACAGGCTTCTCGCAGACCGATACCCGCTATCAATTTCGGATGTGATCGAAGAGGAACCTCAAGAGGATTCGAATGGAAACT
CCAAACCAATCGATGTTTCAAAGCCTAACCCTAACGGTATTGAGTTTGACAATCTGTATTTGGATATGAACGGGATTATCCATCCCTGCTTCCATCCTGA
AGGCAAGCCTGCACCTGCTACTTATGATGATGTGTTCAAGTCCATATTTGCTTACATTGATCATTTGTTTGCTTTGGTTCGTCCAAGGAAACTACTTTTT
ATGGCAATTGATGGGGTTGCTCCTAGAGCTAAAATGAACCAGCAACGGTCGCGACGGTTTAGAGCTGCAAAAGATGCTGCTCAAGCAGAAGCTGAAGAAG
AAAGATTGAGGAAAGAATTTGAGGCTGAGGGGGAGCTTTTATCCGTTAAAGAAAAGCCTGAAACATTTGATTCTAATGTTATCACTCCTGGCACTCAATT
TATGGCTGCTCTTTCGACTGCGCTTCAGTATTATATACAAAGTAGATTGAATCACAATCTGGGCTGGCAGAATACCAAGGTTATTCTTTCTGATTCAAAT
GTTCCTGGTGAGGGAGAGCACAAAATCATGTCTTATATAAGATTGCAACGCAACCTTTCCGGTTTCAATCCAAACACACGCCATTGTCTGTATAGTCTGG
ATGCAGATTTGATTATGCTGTCATTAGCCACGCGTGAGGTTCACTTTTCCATATTAAGAGAGATAGTTACACTTCCCGGGCAGCAGGATAAATGTTTTCT
CTGCGGTCAAGCTGGTCATCTTGCAGCTGAATGTCGTGGCAAGCAAGGAGATGATGCTTTAGATTGGCATGTGGTAGATGATACCCCAATTCACAAGAAG
AAATATCAGTTCCTCAACATCTGGGTGCTGCGAGAATATTTGCAATATGATCTGGACATTCTGAACCCACCATTTGCTATCGACTTTGAACGAATAGTGG
ATGACTTTGTGTTCCTGTGTTTTTTTGTTGGCAATGACTTCCTTCCGCATATGCCAACATTGGAAATTAGGGAGGGTGCTATAAGCTTACTGATGCATAT
TTATAGAAGGGAGTTCTCTGCAATGGGTGGTTATCTTACTCTTGCGGGTGAGGTTTTCCTAGATAAAGTAGAACACTTTATTCAGTGTGTTGGTGTTTAC
GAGGAGCAAATCTTTCAGAAGAGGACTCGTATACAACAGGCTTTCGATAATAATGAAGAGATGAAACTCAAAGCAAGAAGAGAGTCCTCTGAAGTGATAC
AAGCACCAGTTGTAGATAAGGTTAAATTAGGGGAACCTGGATACAAAGAGAGATACTATGCTGAGAAATTTGAGTTGTCAAATCAAGAGGAGATTGACAA
AGTTAGAAAGGAAGTGGTATTGAAGTATGTGGAAGGATTGTGTTGGGTCTGTCATTATTACTTCCAAGGTGTTTGCTCTTGGCAATGGTTTTATCCGTTC
CATTATGCTCCATTTGCTTCAGATCTTAAGGATCTGGGTGAGGTGGAATTAAACTTTTTTATTGGAGAGCCATTCAAACCCTTTGATCAACTTATGGGAA
CGCTACCAGCTGCAAGTTCTAATGCTTTGCCAAAAGAATACAGGAAATTGATGACCAATCCATCTTCACCAATTCATAGATTCTTTCCCTCAGATTTTGA
AATTGACATGAATGGCAAGCGCTTTGCTTGGCAGGGTATTGCCAAATTGCCATTCATTGATGAGAGGAAATTGCTTGCCCAGACAAAGAAGCTTGAAAGC
ACTTTAACGGAGGAAGAGCAGATCAGGAATAGAGTGATGCTTGATTTGCTATATATTCATCCTGTTCATCCATTAGCTCAACTAGTTATTTCATACTATC
AACAGAACGATCATTTATCTGAAGGTGAAAGATTTGCTTGGGAAATTGACACAAGAGCCAGTGGGGGGATGAATGGTTGTCTTTGGTTATATGAGAGAAA
TGTGCGGAGAAGTGTGGTCCCCTCTCCTATACTTGGATTGCCAGCCTTAGAGGGTAACCAAGTTTTAAATATCACATTTCTAAATCCTAAAAACCGTGCA
CACATCCCAGAAATTCCTGAGGGTGTAGTCATGCCTGAAAAGATCGTAAAACCTGTAGATCTTAAACCATTCCCCACATTATGGCATGAAGATAATGGCC
GGAGACAGCAAGGCAGGGAAAGGCCACAAGTACAACGAGCAATAGCTGCCCCCTTCCTGGGGGATGCAGCTCATCGTCTTGTCAAGAACACCCTTAATAT
CAAACCAAATGGTTCATCCTCTAGAGTGTTTGACCAGCAACTATATCATAGTATCCCCGGCAATTACACATTCTACAGACCAAGACCAGCTGGACCAGCT
GGTTATGGAAGGGGCTATTGGGATGATCCAAACTATCATTATGCACAGCACAGTAACCAGCAAGGCCTGATGAGTAATCCTAGGTATCGATCTTTGTCAA
ATGGTGTGCAAAGCAACCGACACAACTTTAGGACTCAAGACGGAGTACAGTACCATCAACAGTATCATAACTTGAGTACTGGGGTGTCTGCTCTGACTGT
AGAAGAAAATATCAGAAGCAGGGCACCTGCAGTGATCTCACCTCGAATGCCGAATCCAGGAAACACTCCAAACTTACAAAATCAGGCCGAGCAGAACACA
GGTCTACTTTCATCACCACCCACCAATTGGATCAACAAAACAGCTGCAGGAAACACTGGAATGTACTTCAAACAAAAATCAACATCGATAGGTCCAAATG
AGAAGCAGGTGAAGCAGGTTTATCAGGTTAAGACACAAGTAGCTCAGGAGACGCCAGATATTCAAGCTCAGGAGACGCCAGATTTAAAACAGCAGTAA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.013G035600.1 pacid=42810982 polypeptide=Potri.013G035600.1.p locus=Potri.013G035600 ID=Potri.013G035600.1.v4.1 annot-version=v4.1
MGVPAFYRLLADRYPLSISDVIEEEPQEDSNGNSKPIDVSKPNPNGIEFDNLYLDMNGIIHPCFHPEGKPAPATYDDVFKSIFAYIDHLFALVRPRKLLF
MAIDGVAPRAKMNQQRSRRFRAAKDAAQAEAEEERLRKEFEAEGELLSVKEKPETFDSNVITPGTQFMAALSTALQYYIQSRLNHNLGWQNTKVILSDSN
VPGEGEHKIMSYIRLQRNLSGFNPNTRHCLYSLDADLIMLSLATREVHFSILREIVTLPGQQDKCFLCGQAGHLAAECRGKQGDDALDWHVVDDTPIHKK
KYQFLNIWVLREYLQYDLDILNPPFAIDFERIVDDFVFLCFFVGNDFLPHMPTLEIREGAISLLMHIYRREFSAMGGYLTLAGEVFLDKVEHFIQCVGVY
EEQIFQKRTRIQQAFDNNEEMKLKARRESSEVIQAPVVDKVKLGEPGYKERYYAEKFELSNQEEIDKVRKEVVLKYVEGLCWVCHYYFQGVCSWQWFYPF
HYAPFASDLKDLGEVELNFFIGEPFKPFDQLMGTLPAASSNALPKEYRKLMTNPSSPIHRFFPSDFEIDMNGKRFAWQGIAKLPFIDERKLLAQTKKLES
TLTEEEQIRNRVMLDLLYIHPVHPLAQLVISYYQQNDHLSEGERFAWEIDTRASGGMNGCLWLYERNVRRSVVPSPILGLPALEGNQVLNITFLNPKNRA
HIPEIPEGVVMPEKIVKPVDLKPFPTLWHEDNGRRQQGRERPQVQRAIAAPFLGDAAHRLVKNTLNIKPNGSSSRVFDQQLYHSIPGNYTFYRPRPAGPA
GYGRGYWDDPNYHYAQHSNQQGLMSNPRYRSLSNGVQSNRHNFRTQDGVQYHQQYHNLSTGVSALTVEENIRSRAPAVISPRMPNPGNTPNLQNQAEQNT
GLLSSPPTNWINKTAAGNTGMYFKQKSTSIGPNEKQVKQVYQVKTQVAQETPDIQAQETPDLKQQ
|
DESeq2's median of ratios [POPLAR]
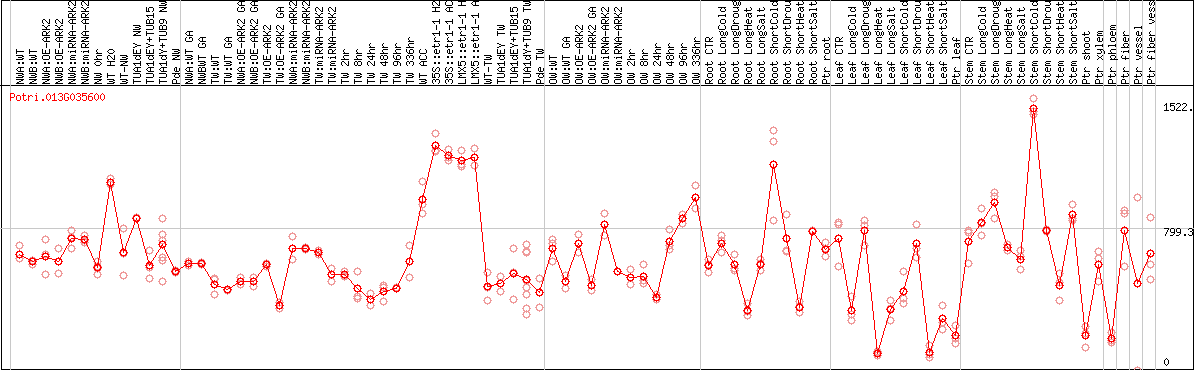
Coexpressed genes
Potri.013G035600 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
