CEF.2 (Potri.013G035800) [POPLAR]

| External link |
|
||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | CEF.2 | ||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| ||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| ||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.013G035800.8 pacid=42811866 polypeptide=Potri.013G035800.8.p locus=Potri.013G035800 ID=Potri.013G035800.8.v4.1 annot-version=v4.1
ATGGCATCTAATGTGCCTCCAGGGGCACCTAGACAACCACCACCGCCTAGTTATAACCCAAATTATCAGCAAAACCCGAATATTTTATCCGATAATTTTC
AGAATTTGAATCTCAATCGGCCCTCTTCCATGCCGAATTCAGCTCCTAGGCCTTCGCCATTTGGCCAACCACCGCCATTTCCCTCCTCAGCTCCTTCACC
ACAATTTTCCCGGCCTGGTGCACCACCTCCTGGTGTTGTCCCTAGACCTTCAGTGCCTCCCTCGGGATTACCTCCATCCACTTTCCCCCCAAATGTTACC
CCTGGTAGACCTACTGGGCCGCCTTTCAGTCAGCCGCAGCCTTTCAGTCAGACGCAGCCTTTCAGTCAGCCGCAGCCTTTCAGTCAGCCGCAGCCTTTCA
GTCAGACGCAGCCTTTTGGGTCCAGACCGCCTCCGGGGTCATTCCCGTCTTCTGCAAGTAGTGGTCTGGCAATGGGTCCTGTCTCTGGTGCACCACCTCA
AGGCTCACTTGTACCGCCATTGGGTTCTCGCCCTAGTCCTGCTGCACCATCTTCTTCACCTTTGAGTATGCCCCCTTCCAGTTCATTTGGTGGTTTAATG
AGTAATGGGCCACCAGCACCAGCTTTCCAGAGTGCTCCACGTTTTCCTTCCCCTGTTAGTGCACCACAGCAACCGCCCATGGGTCCTCCACCAACAGTGG
GGTTGGCTCGCGCACCGCCTCAAAGCATGCGCCACTTAATGGGTAGTTCGGGGATTAGTGGCCAGCCAGTTGCTCCCTTTTCAGCACCACCTCAAGGGAC
ACCGTTTTCAGCTCAGCATGGGATGGCACCACCACCAGTAGGTTCTCCTTTCGCACCACAGATGCAACCGCAGTCTGTAGCTCAACCTCCACCAATTCCT
GGTTCTGCTCAACCACCTAGAATGTTTGGAATGCCACCACCACTACCAAATCAAATGACTGCTATATCACCTGTTATGGGTCAAACTGGAAGTCCTCTAT
CAGGAGCATCAAAGATTGATCCCAATCAAATTCCCCGGCCCATTCCAGGTTCCTCTGTGATCCTGCACGATACTCGTGCGGGGAATCAGGCCAACCCACC
ACCGCCTGCTACAAGTGACTACATTGTTTCAGACACTGGGAATTGCAGTCCACGGTACATGAGGTGCACCATCAATCAGATTCCATGCACGGTTGATCTG
TTATCAACATCTGGCATGCCGTTGGCTTTGTTGGTTCAACCTCTGGCCCTTCCTCATCCATCAGAGGATCCTGTCCAGGTTGTTGATTTTGGAGAAAGTG
GTCCTGTTCGTTGTTCTCGTTGCAAAGGCTACATAAATCCCTTTATGAAATTCATTGATCAAGGGAGGCAGTTCATTTGTAATTTGTGTGGGTTTACTGA
TGAGACTCCCCGTGACTACCACTGCAATCTAGGTCCAGATGGTAGGCGTAGAGATGCTGATGAAAGGCCCGAGCTATGCAGGGGAACAGTTGAATTTGTT
GCCACAAAAGAATACATGGTCCGTGATCCCATGCCGGCTGTGTATTTTTTTCTTATTGATGTATCCATGCATGCCATACAAACTGGTGCGACTGCTGCAG
CTTGCAGTTCAATTAATCAAGTTATTGCTGATCTTCCGGAAGGGCCACGGACAATGGTGGGCATTGCAACTTTTGATTCAACAATCCATTTTTACAATCT
GAAACGGGCATTACAACAGCCACTGATGCTTATTGTACCTGATATACATGATGTTTATACTCCTCTGCAAACTGATGTTATTGTTCCGGTTTCCGAGTGC
CGTCAACATTTAGAGCTATTATTGGATAGTATTCCAACTATGTTTCAGAACAGTAGAATCGTTGAATCAGCCTTCAGTGCAGCAATCAAGGCAGCTTTTT
TGGCAATGAAAAATACTGGAGGGAAGCTTTTGGTATTTCAATCAGTCTTGCCTTCTGTTGGCATTGGAGCCCTTTCTGCTCGAGAGGCTGAGGGAAGAAG
TAACATCTCTGCAGGGGAGAAGGAGGCCCATAAATTACTCCAACCAGCAGACAAAACATTGAAGGAAATGGCAATTGAATTTGCTGAATATCAGGTCTGT
GTTGATGTGTTCATCACTACCCAGACTTATGTCGACATTGCTTCTATCTCTGTCATCCCTAAAACAACTGGAGGGCAGGTTTATTATTACTATCCTTTCT
CTGCTGTTTCTGATCCAGCAAAGCTGTACAATGATCTCAGATGGAATGTCACACGGCCTCAAGGGTTTGAGGCAGTGATGCGTGTGAGGTGCAGCCAGGG
CATTCAAGTTCAAGAATACCATGGAAACTTTTGCAAGCGCATTCCAACAGATATTGACCTAGCTGCGATTGATTGTGACAAAACTATTATGGTTACATTA
AAACATGATGATAAGTTGCAGGATGGCTCAGAATGTGCTTTCCAGTGTGCCCTCCTCTACACCACCGTGTATGGACAAAGAAGAATTCGTGTTACAAATT
TATCATTACCATGCACAAACAATTTAAGTAATCTGTTCCGCTTGGCTGACTTGGATTCCCAATTTGTGTGTTTCTTGAAGCAAGCGGCAAGCGAAATCCC
TTCCAACCCACCCTTGGTTATCCGAGACCGAGTGACAAACTTTTGCATAAACATTCTGCTTTCATATCGCAAATTTTGTGCGACAGTATCATCATCAGGA
CAACTCATCCTTCCAGAAGCACTTAAACTTCTGCCCCTGTACACCCTCGCATTAATTAAAAGTACAGGATTGAAACTCGATGGGAGGATAGACGATCGGT
CTTTTTGGATCAATTATGTTTCTTCTGTTTCTACTCCTTTAGCGATTCCATTGGTGCACCCTAGGATGATTGCAATCCACGACCTTGATTCACAGGAAGC
AATTGGATCTCTTATCCCCCCTGCACTTCCTCTTTCTAGTGAATATGTGAATGACAATGGAGTTTATCTTCTTGAGAATGGTCAGGATGTTTCAATATAT
ATTGGGAACTCGGTTAATCCAGATATCTTGCAAAAACTGTTTGGCATTTCATCTGTTGCTGAAATTCCAACTCAGTATGTCCTGGAGCAATATGACAATT
CACTATCAAAGAAACTAAATGATGTGGTGAACGAGATACGACGCCAAAGATGTTCTTTCCTGCGTTTAAAGTTGTGTAAAAAAGGAGATCCATCAGGAAT
GACATTCTTCTCATATTTGGTTGAAGACAAGGTTCCAGCTGGCACTCTCTCCTATGTAGAATTTCTAGTGCAGGTCCATCGGCAAATTCAAGTAAAAATG
TCTTCATAA
|
||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.013G035800.8 pacid=42811866 polypeptide=Potri.013G035800.8.p locus=Potri.013G035800 ID=Potri.013G035800.8.v4.1 annot-version=v4.1
MASNVPPGAPRQPPPPSYNPNYQQNPNILSDNFQNLNLNRPSSMPNSAPRPSPFGQPPPFPSSAPSPQFSRPGAPPPGVVPRPSVPPSGLPPSTFPPNVT
PGRPTGPPFSQPQPFSQTQPFSQPQPFSQPQPFSQTQPFGSRPPPGSFPSSASSGLAMGPVSGAPPQGSLVPPLGSRPSPAAPSSSPLSMPPSSSFGGLM
SNGPPAPAFQSAPRFPSPVSAPQQPPMGPPPTVGLARAPPQSMRHLMGSSGISGQPVAPFSAPPQGTPFSAQHGMAPPPVGSPFAPQMQPQSVAQPPPIP
GSAQPPRMFGMPPPLPNQMTAISPVMGQTGSPLSGASKIDPNQIPRPIPGSSVILHDTRAGNQANPPPPATSDYIVSDTGNCSPRYMRCTINQIPCTVDL
LSTSGMPLALLVQPLALPHPSEDPVQVVDFGESGPVRCSRCKGYINPFMKFIDQGRQFICNLCGFTDETPRDYHCNLGPDGRRRDADERPELCRGTVEFV
ATKEYMVRDPMPAVYFFLIDVSMHAIQTGATAAACSSINQVIADLPEGPRTMVGIATFDSTIHFYNLKRALQQPLMLIVPDIHDVYTPLQTDVIVPVSEC
RQHLELLLDSIPTMFQNSRIVESAFSAAIKAAFLAMKNTGGKLLVFQSVLPSVGIGALSAREAEGRSNISAGEKEAHKLLQPADKTLKEMAIEFAEYQVC
VDVFITTQTYVDIASISVIPKTTGGQVYYYYPFSAVSDPAKLYNDLRWNVTRPQGFEAVMRVRCSQGIQVQEYHGNFCKRIPTDIDLAAIDCDKTIMVTL
KHDDKLQDGSECAFQCALLYTTVYGQRRIRVTNLSLPCTNNLSNLFRLADLDSQFVCFLKQAASEIPSNPPLVIRDRVTNFCINILLSYRKFCATVSSSG
QLILPEALKLLPLYTLALIKSTGLKLDGRIDDRSFWINYVSSVSTPLAIPLVHPRMIAIHDLDSQEAIGSLIPPALPLSSEYVNDNGVYLLENGQDVSIY
IGNSVNPDILQKLFGISSVAEIPTQYVLEQYDNSLSKKLNDVVNEIRRQRCSFLRLKLCKKGDPSGMTFFSYLVEDKVPAGTLSYVEFLVQVHRQIQVKM
SS
|
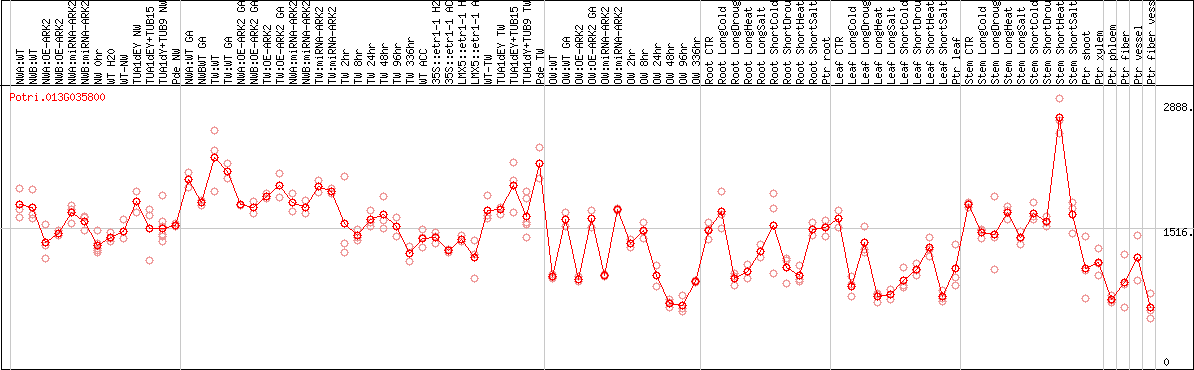
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.013G035800 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
