Potri.013G036600 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.013G036600.1 pacid=42811005 polypeptide=Potri.013G036600.1.p locus=Potri.013G036600 ID=Potri.013G036600.1.v4.1 annot-version=v4.1
ATGATGTCCTTCCTACAGAAACCATTCTCAAAGTTTCTCCAAGTCCTTTTTTTTCTCCTTCTAGTGTGCATTCCTTCTTTCTTTGCCTTTCATCCTAATT
CCTCTGCAACTTCATTTGGTGCTGCCACATATGTAGCTGCTGAAGGTAATAAAGAAGCGGAGGCTCTCCTAAAATGGAAAGCAAGTCTTGATGACAACCA
CAGCCAATCTGTCCTGTCTTCTTGGGTTGGCAGCAGCCCTTGCAAGTGGCTTGGGATCACTTGTGACAATTCTGGAAGTGTCGCGGGTTTCAGCCTTCCA
AATTTTGGTCTGAGAGGTACACTTCATAGTTTCAACTTCTCATTCTTCCCCAATCTTCTCACTCTCAATCTCGGGAACAATTCACTCTACGGAACTATTC
CACTTGAGATGGGATTGTTAACAAGTCTTAACTTTCTCTACCTTGATAAAAATAATCTCACTCGTAGGATCCCTTTTTCTATTGGAAATTTGAGAAACTT
ATCCATTCTCAATCTCAAGAATAACAAACTCTCTGGTTCCATCCCTTCTTCTATTGGGAACATGACTTTGCTCACTCGACTTGATCTGAACAATAATAAT
TTATCTGGATCTGTTCCTCGGGAAATAGGACAACTTGAATCCCTTGTTGAATTGAAATTGTCTTCAAATAATTTCACTGGCCATTTACCACGAGACTTGT
GCCTTGGTGGGTTGCTTGTAAATTTTACAGCCGCCAACAATCATTTTTCTGGTCCAATCCCTAAAAGCTTGCGAAATTGCACTAGTCTATTTAGATTCAG
GCTTGACGGGAACCAATTGTCAGGGAATATTTCTGAAGATTTTGGCCTCTACCCAAACTTGAACTATGTGGACCTAAGTCACAATGATTTGTCCGGTGAG
CTTAAGTGGAAATGGGGAGGTTTTCACAACTTGGCGTGCCTATTATTGTCAAACAATAATATCTCCGGTGAGATACCATCAGAGCTTGGAAAGGCTACTA
GGCTACAAATAATTGATCTCTCGTCAAATCTCCTAAAGGGGACAATTCCAAAAGAATTGGTGCAGTTGAAGGCGTTGTACAAGCTTACTCTTCATAACAA
CCATCTTTGTGGTGTCATTCCCTTTGAAATCCAAATGCTATCTCGACTTCAATCCCTTAATTTAGCTTCTAATAATCTTGGTGGATCAATCCCAAAACAA
CTGGGACAGTGCTCAAATTTATTACAGTTGAACTTGAGTCATAATAAGTTTACAGGGAGCATCCCATCTGAGATAGGCTTACTGCATTTACTTGGACATC
TAGATCTCAGTGGGAATCTTCTGGCAGGAGAGATACCATCACAAATTGGGCAATTGAAACGGTTAGAAACGATGAACCTCTCTCACAACAAACTTTCCGG
TTTGATTCCAACTGCTTTTGTAGATTTGGTGAGCTTGACAGCTGTAGACATCTCCTACAATGAACTAGAGGGCCCTATCCCCGAAATCAAAGGCTTCACT
GAAGCATTTATGAATAATAGTGGTCTCTGTGGAAATGTCAGTGGTCTAAAGCCTTGTACACTTCCTACAAGCAGGAGAAAGAGCAACAAAATTGTCATTT
TAATTCTTTTTCCTCTCCTGGGCAGTCTACTTCTACTACTTATCATGGTTGGGTGTTTATACTTTCACCACCGAACAAGTAGAGACAGAATTTCCTGTTT
AGGAGAGCGACAAAGTCCACTCAGTTTTGCAGTATGGGGCTATCAGGAAGAGATCCTCCATGACACTATCATCCAGGCCACTAATAATTTCAACTCCAAT
AACTGTATAGGGAAAGGGGGATACGGAATTGTTTATCGAGCCATGTTGCCAACAGGTCAGGTGGTTGCCGTGAAGAAACTTCACTCATCAAGAGAGGGCG
AGCTTATGAATATGAGAACTTTTAGAAATGAGATTCACATGTTAATAGATATTCGACATCGAAATATTGCGAAGTTATATGGATTTTGTTCATTAATAGA
GCACTCTTTTTTAGTTTACGAGTTCATAGAAAGGGGAAGTTTGAAGATGAATTTATCTATCGAAGAACAGGCAATGGACTTGGATTGGAATAGAAGGCTT
AATGTTGTTAAAGGAGTGGCCAATGCATTATCCTATTTGCACCATGATTGCTCGCCTCCAATCATTCATCGAGACATTTCCAGCAGCAATGTTCTTTTAG
ATTTGGAATTCGAGGCTCATGTTTCGGATTTTGGGACAGCTCGGCTCTTGATTCCTGACTCGACCAACTGGACCTCATTTGCTGGCACCTTCGGATACAT
AGCTCCAGAGCTAGCTTACACAATGAGAGTGAACGAGAAGTGCGATGTCTACAGCTTTGGAGTGGTCACAATGGAAGTAATAATGGGAATGCATCCGGGT
GATCTCATCTCATCTCTTTCTGCATCAGCATTTTCCTCCTCGTCGTGCTCACAAATCAACCAGCATGCATTGTTGAAGGATGTGATAGACCAGCGTATCC
CACTTCCTGAAAATCGAGTTGCAGAAGGTGTGGTCTACATTATCAAAATAGCATTTGAATGCTTGCTTGCCAATCCCCAATCTAGGCCAACCATGCGACA
AGTAGCTTCGAAGCTCATAGCTCGATGGCCTCCGCTGTCAAAGTCATTCTCCGCAATAACATTGGAAGATCTGATGCCTCAAACAACTATGACAGGCTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.013G036600.1 pacid=42811005 polypeptide=Potri.013G036600.1.p locus=Potri.013G036600 ID=Potri.013G036600.1.v4.1 annot-version=v4.1
MMSFLQKPFSKFLQVLFFLLLVCIPSFFAFHPNSSATSFGAATYVAAEGNKEAEALLKWKASLDDNHSQSVLSSWVGSSPCKWLGITCDNSGSVAGFSLP
NFGLRGTLHSFNFSFFPNLLTLNLGNNSLYGTIPLEMGLLTSLNFLYLDKNNLTRRIPFSIGNLRNLSILNLKNNKLSGSIPSSIGNMTLLTRLDLNNNN
LSGSVPREIGQLESLVELKLSSNNFTGHLPRDLCLGGLLVNFTAANNHFSGPIPKSLRNCTSLFRFRLDGNQLSGNISEDFGLYPNLNYVDLSHNDLSGE
LKWKWGGFHNLACLLLSNNNISGEIPSELGKATRLQIIDLSSNLLKGTIPKELVQLKALYKLTLHNNHLCGVIPFEIQMLSRLQSLNLASNNLGGSIPKQ
LGQCSNLLQLNLSHNKFTGSIPSEIGLLHLLGHLDLSGNLLAGEIPSQIGQLKRLETMNLSHNKLSGLIPTAFVDLVSLTAVDISYNELEGPIPEIKGFT
EAFMNNSGLCGNVSGLKPCTLPTSRRKSNKIVILILFPLLGSLLLLLIMVGCLYFHHRTSRDRISCLGERQSPLSFAVWGYQEEILHDTIIQATNNFNSN
NCIGKGGYGIVYRAMLPTGQVVAVKKLHSSREGELMNMRTFRNEIHMLIDIRHRNIAKLYGFCSLIEHSFLVYEFIERGSLKMNLSIEEQAMDLDWNRRL
NVVKGVANALSYLHHDCSPPIIHRDISSSNVLLDLEFEAHVSDFGTARLLIPDSTNWTSFAGTFGYIAPELAYTMRVNEKCDVYSFGVVTMEVIMGMHPG
DLISSLSASAFSSSSCSQINQHALLKDVIDQRIPLPENRVAEGVVYIIKIAFECLLANPQSRPTMRQVASKLIARWPPLSKSFSAITLEDLMPQTTMTG
|
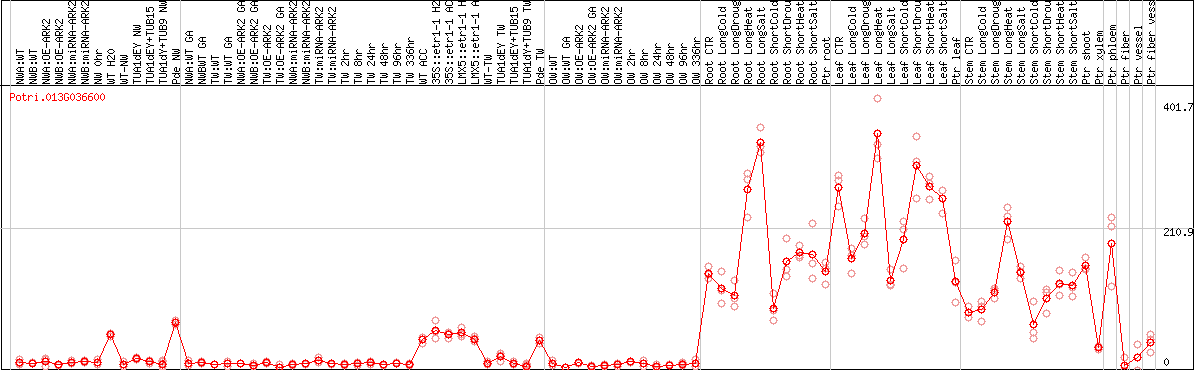
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.013G036600 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
