External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G45060 45 / 8e-06
Disease resistance protein (TIR-NBS-LRR class) family (.1)
AT1G72840 42 / 0.0001
Disease resistance protein (TIR-NBS-LRR class) (.1), Disease resistance protein (TIR-NBS-LRR class) (.2)
AT1G69550 42 / 0.0002
disease resistance protein (TIR-NBS-LRR class) (.1)
AT4G09430 41 / 0.0003
Disease resistance protein (TIR-NBS-LRR class) family (.1)
AT4G28560 40 / 0.0003
RIC7
ROP-interactive CRIB motif-containing protein 7 (.1)
AT4G09360 40 / 0.0004
NB-ARC domain-containing disease resistance protein (.1)
AT5G18370 40 / 0.0008
Disease resistance protein (TIR-NBS-LRR class) family (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.013G037599
178 / 8e-52
AT5G36930 627 / 0.0
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Potri.T005501
66 / 1e-12
AT5G36930 612 / 0.0
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Potri.019G003285
64 / 6e-12
AT5G36930 640 / 0.0
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Potri.T011750
63 / 7e-12
AT5G36930 551 / 2e-176
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Potri.001G066500
63 / 1e-11
AT5G36930 462 / 2e-146
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Potri.019G007884
60 / 1e-10
AT5G36930 654 / 0.0
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Potri.019G004599
59 / 3e-10
AT5G36930 627 / 0.0
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Potri.019G003942
57 / 8e-10
AT5G36930 591 / 0.0
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Potri.007G143300
57 / 1e-09
AT5G36930 519 / 2e-162
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10039850
47 / 3e-06
AT5G36930 573 / 0.0
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Lus10025626
47 / 3e-06
AT3G46530 87 / 5e-18
RECOGNITION OF PERONOSPORA PARASITICA 13, NB-ARC domain-containing disease resistance protein (.1)
Lus10018616
47 / 3e-06
AT5G36930 578 / 0.0
Disease resistance protein (TIR-NBS-LRR class) family (.1), Disease resistance protein (TIR-NBS-LRR class) family (.2)
Lus10028075
45 / 1e-05
AT3G07040 168 / 1e-43
RESISTANCE TO PSEUDOMONAS SYRINGAE 3, RESISTANCE TO P. SYRINGAE PV MACULICOLA 1, NB-ARC domain-containing disease resistance protein (.1)
Lus10005017
43 / 9e-05
AT4G08850 562 / 0.0
Leucine-rich repeat receptor-like protein kinase family protein (.1.2)
PFAM info
Representative CDS sequence
>Potri.013G037566.1 pacid=42811070 polypeptide=Potri.013G037566.1.p locus=Potri.013G037566 ID=Potri.013G037566.1.v4.1 annot-version=v4.1
ATGAACCTGCAGAGCTGCGCTAGTCTTGAGAAATTGCCGGAGAAATTGGGTAATATGAAAGTCTTAACTGATCTTCTATTAGATGATACCGGAGTTCAGA
ATCTACCCTCTTCCACTGGAATTTTGAAGAAGCTCAAAAAGTTATTGGTGCGTGGACGTTGTCCTGGTTTTAGTCTTAATAGTGCATGTCCTCGTCTTCC
TTCAAATGCCTTTCACTCGAAAGAGCTTGCTATCAAGAAACAACCTTTCCTGCCACCCTCCCTCTCTGGTTTAAGCTCATTGACAACACTCGATATTAGT
AATCGCCATTTATCAATCAATGATATTTCCATCAATCTTGGGAGTTTATCCTCTCTCCAGGATCTGAATTTGGCAGTAAATGATTTCTCCGAGTTGCCTG
CCGGCATTGGCAACCTTGCGAAGCTAGAGAAATTGGACTTGTCATGGTGTAGAAATCTTCTGTTCATCTCAGAGATCCCCTCTGGTGGCACGTTATTGCA
GATCATTGGAAAAGGTGTCAATTCAGTCAAAAACAGCGCCGGATCTGTTACTGGGTGA
AA sequence
>Potri.013G037566.1 pacid=42811070 polypeptide=Potri.013G037566.1.p locus=Potri.013G037566 ID=Potri.013G037566.1.v4.1 annot-version=v4.1
MNLQSCASLEKLPEKLGNMKVLTDLLLDDTGVQNLPSSTGILKKLKKLLVRGRCPGFSLNSACPRLPSNAFHSKELAIKKQPFLPPSLSGLSSLTTLDIS
NRHLSINDISINLGSLSSLQDLNLAVNDFSELPAGIGNLAKLEKLDLSWCRNLLFISEIPSGGTLLQIIGKGVNSVKNSAGSVTG
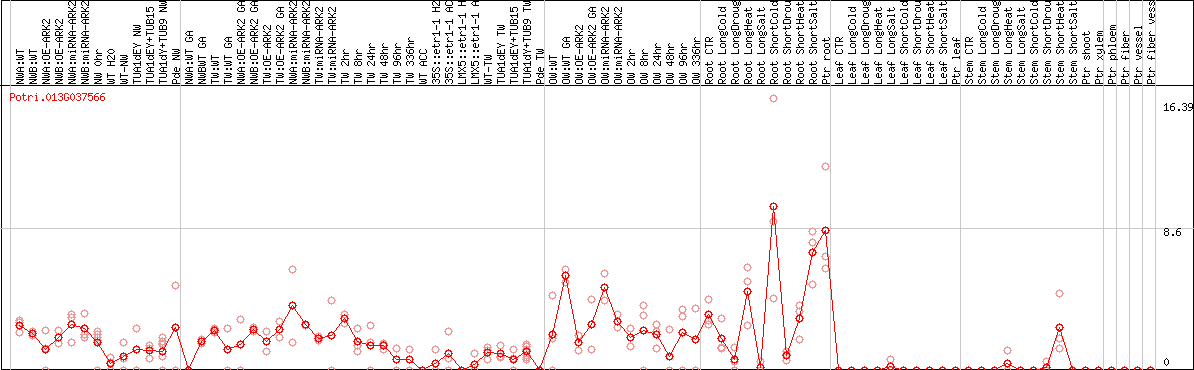
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.013G037566 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.