External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G04750 169 / 5e-50
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT5G43790 140 / 2e-40
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT3G46790 137 / 1e-38
CRR2
CHLORORESPIRATORY REDUCTION 2, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT3G05240 136 / 3e-38
MEF19
mitochondrial editing factor 19 (.1)
AT1G20230 136 / 6e-38
Pentatricopeptide repeat (PPR) superfamily protein (.1)
AT1G34160 134 / 1e-37
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT3G26782 134 / 2e-37
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT3G49170 134 / 3e-37
EMB2261
embryo defective 2261, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT4G33990 133 / 6e-37
EMB2758
embryo defective 2758, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
AT5G44230 130 / 4e-36
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.005G053600
243 / 3e-78
AT3G04750 759 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.016G065500
150 / 2e-43
AT5G06540 828 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.002G220300
144 / 1e-40
AT1G11290 559 / 0.0
CHLORORESPIRATORY REDUCTION22, Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.003G223900
137 / 6e-39
AT5G56310 508 / 1e-176
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.002G258766
137 / 1e-38
AT1G06150 705 / 0.0
EMBRYO DEFECTIVE 1444, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2)
Potri.004G125500
136 / 3e-38
AT3G29230 829 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.013G058900
136 / 5e-38
AT4G13650 1282 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Potri.003G081700
136 / 6e-38
AT4G16835 852 / 0.0
Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Potri.005G012600
135 / 6e-38
AT2G29760 540 / 0.0
ORGANELLE TRANSCRIPT PROCESSING 81, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10000117
144 / 4e-42
AT4G21065 404 / 5e-139
Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2)
Lus10006628
143 / 8e-42
AT4G21065 399 / 7e-137
Tetratricopeptide repeat (TPR)-like superfamily protein (.1), Tetratricopeptide repeat (TPR)-like superfamily protein (.2)
Lus10039388
143 / 9e-41
AT5G43790 506 / 8e-176
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10030053
143 / 2e-40
AT1G08070 884 / 0.0
ORGANELLE TRANSCRIPT PROCESSING 82, EMBRYO DEFECTIVE 3102, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10030225
137 / 1e-39
AT5G08510 472 / 2e-165
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10005987
138 / 3e-39
AT5G08510 602 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10033026
139 / 7e-39
AT1G08070 921 / 0.0
ORGANELLE TRANSCRIPT PROCESSING 82, EMBRYO DEFECTIVE 3102, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10017446
137 / 5e-38
AT1G11290 1152 / 0.0
CHLORORESPIRATORY REDUCTION22, Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10013149
136 / 5e-38
AT1G56690 946 / 0.0
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10028837
132 / 1e-37
AT1G11290 650 / 0.0
CHLORORESPIRATORY REDUCTION22, Pentatricopeptide repeat (PPR) superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0020
TPR
PF01535
PPR
PPR repeat
Representative CDS sequence
>Potri.013G041150.1 pacid=42812597 polypeptide=Potri.013G041150.1.p locus=Potri.013G041150 ID=Potri.013G041150.1.v4.1 annot-version=v4.1
ATGATAACAGGATTTGCTTTCCATGGTTATGGAAGCAAGGTTTTGCAACTATGGAAGATGCAGGAAGATGCAAGTCCAGACAACGTGACTTTTGTTTCTG
TTCTTTCAGCTTGTAGCCATAGTGGACTTGCAGATCAAGGGATTAAAACTTTCAGTTCTGTGAAAGACCATGGCATTGAACCAGTAGTCGAGCACTATGG
ATGTTTGGTAGATCTTTTAGCACGACCAGGGAGGTTGTCTGAGGAAAAAGATATCATTAATCAGATGCTCATGAAACCAAGTCGTTCCATCTGGGGGGGA
ATACTAAATGCTTGCCAAGTTCAAGAGGATGTGGAAACGGCGGAGATAGCTTCCAGGGAGTCGCTAAATTTAGACCCTGAGGAAGAAGGTGGATACACGT
TGCTGTCTAACATAAATGCAGCTAGTGGAAGGTGA
AA sequence
>Potri.013G041150.1 pacid=42812597 polypeptide=Potri.013G041150.1.p locus=Potri.013G041150 ID=Potri.013G041150.1.v4.1 annot-version=v4.1
MITGFAFHGYGSKVLQLWKMQEDASPDNVTFVSVLSACSHSGLADQGIKTFSSVKDHGIEPVVEHYGCLVDLLARPGRLSEEKDIINQMLMKPSRSIWGG
ILNACQVQEDVETAEIASRESLNLDPEEEGGYTLLSNINAASGR
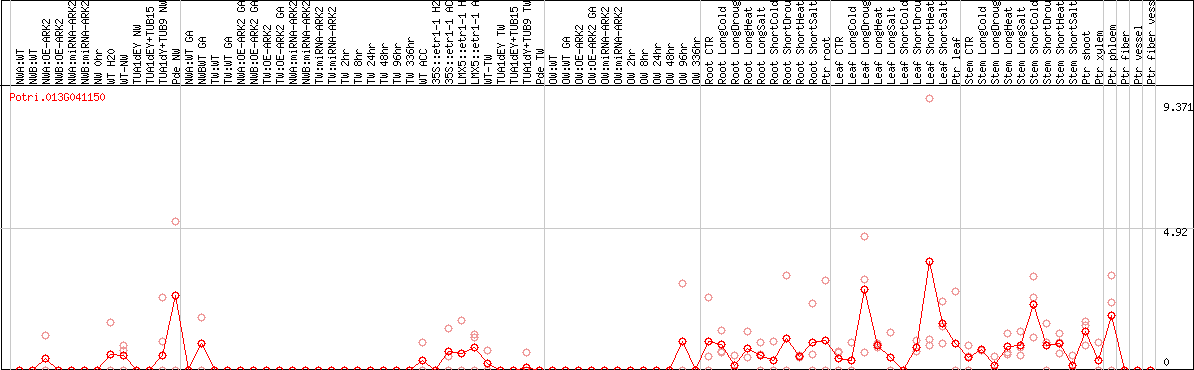
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.013G041150 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.