CHR923 (Potri.013G048500) [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | CHR923 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.013G048500.2 pacid=42811504 polypeptide=Potri.013G048583.1.p locus=Potri.013G048500 ID=Potri.013G048500.2.v4.1 annot-version=v4.1
ATGGAGGATAAACACGAGGAAGTTGAGGACATTGAGAGTGGTTTAAGTGATTCATTTATTGACGACGACGACGAAGATTCTAATAATGATGATGATAACG
AACCTTCTACATCTGGACAAGATGATGGGACCCGTATTCAGGAACCTTTGACCGACCAAGAAGTAGAGGAGTTGGTTGCTGAGTTCCTGGAAGTTGAGAG
TAAGGCTGCAGAGGCTCAAGAAGCACTGGAAAAGGAGTCTCTTGCAAAAGTAGAGAGTGATGTGAGAGAGGAGTTGGCACGAAGTCTCCAGGGTGATGAT
CTGGAGGCAGCTGTTGAAGATGAGATGGCTACTTTCAGGGAAGAGTGGGAAAATGTGCTCGATGAGCTCGAGACTGAGAGTTATCATTTGTTGGAACAAC
TTGATGGGACTGGTATTGAGCTGCCAAACCTTTATAAGTGGATTGAGAGCCAGGCTCCAAACAGTTGCTGCACTGAAGCTTGGAAAAGAAGGGCACACTG
GGTGGGAACTCAGATGACCAAGGAAACGACTGATACAGTAGCTGATGCTGAGAAATATCTTCAAATCCATAGGCCTGTTAGACGAAAACATGGTAAATTA
TTGGAGGAGGGTGCAAGTGGATTCCTGCAGAAAAAACTTGCAATGGATGGAAGCGAGGCTATTGCAGAAAATAGAGAAGTAGATTGGGCCTCTATGAAGA
AACTATTCTCAACTAGCAGTTCTGAGGATGTTGCTTCATTTGGCAGCAAGCATTGGGCTTCTGTTTACCTTGCCAACACTCCCCAGGAAGCTGCACTTAT
GGGACTTAAATTTCCTGGAGTCAATGAGGTTGAGGAGATTGAAGACATTGATGGGAATTCCACTGATCCATTTGTTGCAGAGGCCATAGCAAATGAAAAG
GAATTAGTTCTTTCAGAGGAACAGAGAAAAAATTACAGAAAGGTCAAAGAGGAAGATGATGCAAAAATTGATCAGAAACTTCAACTTCGTTTGAAACAAA
GGAGACGTCTTAAGAGATGTAAACAGGGTGTCAGCTCAGTCGTCCAGGAAATGGGAACAAACATGGCTGAGTCATTGCCTCTGGATGACAATCATCATGA
GGTCACATGCCAAGATTTGAAGAAAGACGTGTGTGAAAATTCTGGAGACCTTGATATGGAACAGTTGATGAGTGAGAGCAATTCAGTTTTCCCAGAATCT
GATGCCTCTGAACCCAGAAGGTCCAAGCGTCCAAATGAGAGTGAGGACCTAAGCATCAATAATAAGAAGATTCGGACTGTTATCATAGACAGTGACAATG
AAGCAGACATTTTGGAGGATAAATCAGTACACGGCATTAAAGTTGAAGACCAATCAACTTTGCTAGAAAACATTGGGGACCCTTCTGCTGGCTGCAATCC
TTCACAGGGTTCAAGTGAGAAGTTTCAGTGCACTGCCTGTGATAAAGTTGCTGTTGAAGTGCACTCACATCCACTTTTGAAAGTAATTGTTTGCAAAGAT
TGTAAATTCTTAATGGAGGAAAAAATGCATGTGAAGGATCCTGACTGTTCTGAGTGTTACTGTGGATGGTGTGGAAAAAACAATGACTTGGTAAGTTGTA
GGTCATGCAGAACATTATTCTGCACTGCTTGTATAAAAAGGAACATCGGTGAAGAGTACTTGTATAAGGTTCCAGTTTCTGGTTGGCAATGTTGTTGCTG
CTCTCCAAGTCTGCTACAAAGACTTACATCACAGTTAGAGAAAGCCATGGGGTCTGGAGATATAATGGTTTCAAGTTCTGATAGTGATTCAGACTCTTCA
GATACAAATGACGGTGTTACAATCAGCTCTAAGAGAAAGAAGCAAAAGAAGATTAGAAGAATCATTGATGATGCTGAATTAGGAGAAGAGACCAAGAGAA
AAATTGCTATTGAAAAGGAACGTCAAGAACGCTTGAAATCCCTGAAAGTCAAGTTTTCTGACAAATCCAAGATGATGAATTTTGCAAGCTGTAGCGGAAA
CTTACCCGAAGGTGCTAGTGTTGAAGTGATTGGTGATGCCACAACTGGTTATATTGTGAATGTTGCGAGGGAAAAAGGTGAAGAAGCTGTGAGGATTCCT
CCAAGCTTATCATCTAAATTGAAAGCTCATCAGGTGGCAGGGATAAGATTTCTATGGGAAAATATTATACAGTCAATTAGAAAAGTGAAGTCTGGGGATA
ATGGTCTTGGTTGTATTTTAGCGCATACAATGGGCCTTGGTAAAACTTTTCAGGTCATAGCTTTTCTGTACACTGCCATGAGAGGTGTTGATCTGGGTTT
AAGAACAGCACTAATTGTGACTCCTGTGAATGTGCTGCATAACTGGCGCAAGGAGTTCATGAAGTGGACACCTTCAGAAGTAAAGCCGATCCGTGTTTTC
ATGCTGGAGGATGTGTCGAGGGAGAGAAGAGTGGAATTGCTTGCAAAGTGGAGAGCTAAAGGTGGTGTTTTCTTGATAGGCTACTCTGCCTTTAGGAACT
TATCTCTCGGAAAGAATGTGAAGGAACGAAATATGGCTAGAGAAATGTGTAGTGCCTTGCAGGATGGACCTGATATACTTGTTTGTGATGAAGCCCACAT
CATCAAGAACACCAGAGCTGAAACAACCCAGGCACTGAAACTCGTTAAGTGCCAGAGAAGGATAGCATTAACTGGATCACCTCTTCAGAACAATCTAATG
GAGTATTATTGTATGGTTGATTTTGTAAGAGAAGGTTTTCTTGGCAGCAGCCACGAGTTTCGGAACCGCTTCCAAAATCCTATAGAAAATGGACAACATA
CTAATTCAACGGTTGATGATGTAAAGATAATGAACCAAAGATCTCATATTCTCTATGAACAATTGAAAGGGTTTGTTCAAAGAATGGACATGAGTGTGGT
GAAGAAAGACCTGCCACCTAAAACTGTATTTGTAGTGGCTGTAAAGCTCTCGCCATTACAGAGGAAATTGTACAAGAGATTCCTTGATGTGCATGGGTTC
ACAAATGGTAGGGCTTCAAATGAAAAGACGAGTAAGAGCTTTTTTGCTGGTTACCAGGCACTGGCTCAGATATGGAACCATCCTGGAATTTTGCAATTGA
GGAAAGGGAGAGAATATGTTGGCAATGTTGAAAACTTTCTTGCAGATGACTGCTCTAGTGATGAAAATGTGGACTACAATACAATTGTCGAAGAGAAGTC
AAGGAATCCAAATGATTTTATTCAAGGGAAAAACGATGATGGATTCTTTCAAAAGGATTGGTGGAATGATCTTCTACTTGAAAATAATTATAAAGAGGTT
GATTACAGCGGCAAAATGGTATTGCTGCTTGACATTTTAGTCATGTCTTCTGATGTGGGTGATAAGACGTTGGTTTTTACCCAGAGCATACCCACTTTAG
ATCTGATAGAGCTTTATCTTTCAAGATTACCTCGACTTGGAAAAAAGGGAAAGTTTTGGAGAAAAGGAAAAGACTGGTATAGGCTAGATGGAAGAACAGA
GAGCTCTGAAAGACAGAGGTTGGTTGAGAGGTTCAATGATCCCAAGAATAAACGGGTCAAGTGTACTTTGATTTCTACCAGGGCTGGTTCTCTTGGGATT
AATCTCTATGCTGCTAACCGGGTGGTTATTGTTGATGGCTCTTGGAATCCAACGTATGATCTTCAGGCCATATATCGGGCTTGGAGGTATGGCCAGACTA
AGCCAGTGTTTGCTTACAGATTAATGGCACATGGAACCATGGAAGAAAAAATCTATAAGCGTCAGGTAACAAAGGAGGGACTTGCTGCAAGGGTGGTTGA
TAGACAACAAGTATATAGGACTATCTCCAGAGAAGAAATGTTGCATCTTTTTGAATTTGGTGATGATGAAAACTCAGACACACTAATTGACATTGGCCAA
GAGTATAGACAGGCAGATACCAGGAATATTTCTAGCCAAACCGCAAATTCTTTGAAACAGAATGCGTCTCGTTCTCATGGAAGCTGTGCTTCTGATAAGG
TGATGGAAAGCTTGGTTGGCAAGCACCGTCAGAGGTGGATTTTCGATTACCATGAACATGAGACTCTTTTGCAAGAGAATGAAGAGGAAAAACTAACAAA
GGAGGAGCAAGACATGGCTTGGGAAGTATATAAGAGATCATTAGAATGGGAAGAAGTGCAGCGAGTTTCCCTTGATGATTCCACTTTTGAGCGAAAGCCA
CCCATGTCAAATGGTGCATCTTCTGCCCCAGATGCAAGCAGCATACCTGTGCCAAGCATGGCCCGTCCTGCATCTGAGGCTTCCAATGGAGCACCTTCCC
AGAGCATTTTAAGGAGTCGTATGGTGCAGCGGAAATGTACTAATCTCTCTCATTTGCTGACCCTTAGAAGTCAGGGAACGAAAGCTGGGTGCACTACTAT
ATGTGGGGAATGTGCCCAGGAGATAAGCTGGGAAGATCTGAAACGGGAGGGTAAGGCAGCACGGTAA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.013G048500.2 pacid=42811504 polypeptide=Potri.013G048583.1.p locus=Potri.013G048500 ID=Potri.013G048500.2.v4.1 annot-version=v4.1
MEDKHEEVEDIESGLSDSFIDDDDEDSNNDDDNEPSTSGQDDGTRIQEPLTDQEVEELVAEFLEVESKAAEAQEALEKESLAKVESDVREELARSLQGDD
LEAAVEDEMATFREEWENVLDELETESYHLLEQLDGTGIELPNLYKWIESQAPNSCCTEAWKRRAHWVGTQMTKETTDTVADAEKYLQIHRPVRRKHGKL
LEEGASGFLQKKLAMDGSEAIAENREVDWASMKKLFSTSSSEDVASFGSKHWASVYLANTPQEAALMGLKFPGVNEVEEIEDIDGNSTDPFVAEAIANEK
ELVLSEEQRKNYRKVKEEDDAKIDQKLQLRLKQRRRLKRCKQGVSSVVQEMGTNMAESLPLDDNHHEVTCQDLKKDVCENSGDLDMEQLMSESNSVFPES
DASEPRRSKRPNESEDLSINNKKIRTVIIDSDNEADILEDKSVHGIKVEDQSTLLENIGDPSAGCNPSQGSSEKFQCTACDKVAVEVHSHPLLKVIVCKD
CKFLMEEKMHVKDPDCSECYCGWCGKNNDLVSCRSCRTLFCTACIKRNIGEEYLYKVPVSGWQCCCCSPSLLQRLTSQLEKAMGSGDIMVSSSDSDSDSS
DTNDGVTISSKRKKQKKIRRIIDDAELGEETKRKIAIEKERQERLKSLKVKFSDKSKMMNFASCSGNLPEGASVEVIGDATTGYIVNVAREKGEEAVRIP
PSLSSKLKAHQVAGIRFLWENIIQSIRKVKSGDNGLGCILAHTMGLGKTFQVIAFLYTAMRGVDLGLRTALIVTPVNVLHNWRKEFMKWTPSEVKPIRVF
MLEDVSRERRVELLAKWRAKGGVFLIGYSAFRNLSLGKNVKERNMAREMCSALQDGPDILVCDEAHIIKNTRAETTQALKLVKCQRRIALTGSPLQNNLM
EYYCMVDFVREGFLGSSHEFRNRFQNPIENGQHTNSTVDDVKIMNQRSHILYEQLKGFVQRMDMSVVKKDLPPKTVFVVAVKLSPLQRKLYKRFLDVHGF
TNGRASNEKTSKSFFAGYQALAQIWNHPGILQLRKGREYVGNVENFLADDCSSDENVDYNTIVEEKSRNPNDFIQGKNDDGFFQKDWWNDLLLENNYKEV
DYSGKMVLLLDILVMSSDVGDKTLVFTQSIPTLDLIELYLSRLPRLGKKGKFWRKGKDWYRLDGRTESSERQRLVERFNDPKNKRVKCTLISTRAGSLGI
NLYAANRVVIVDGSWNPTYDLQAIYRAWRYGQTKPVFAYRLMAHGTMEEKIYKRQVTKEGLAARVVDRQQVYRTISREEMLHLFEFGDDENSDTLIDIGQ
EYRQADTRNISSQTANSLKQNASRSHGSCASDKVMESLVGKHRQRWIFDYHEHETLLQENEEEKLTKEEQDMAWEVYKRSLEWEEVQRVSLDDSTFERKP
PMSNGASSAPDASSIPVPSMARPASEASNGAPSQSILRSRMVQRKCTNLSHLLTLRSQGTKAGCTTICGECAQEISWEDLKREGKAAR
|
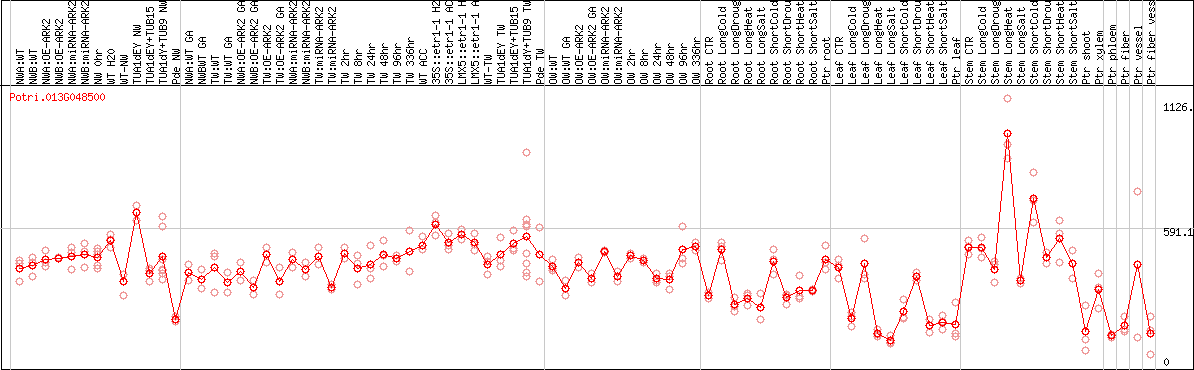
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.013G048500 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
