External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G15120 239 / 4e-78
Protein of unknown function (DUF1637) (.1)
AT5G39890 230 / 5e-75
Protein of unknown function (DUF1637) (.1)
AT1G18490 207 / 9e-66
Protein of unknown function (DUF1637) (.1)
AT2G42670 180 / 6e-56
Protein of unknown function (DUF1637) (.1), Protein of unknown function (DUF1637) (.2)
AT3G58670 175 / 6e-54
Protein of unknown function (DUF1637) (.1), Protein of unknown function (DUF1637) (.2), Protein of unknown function (DUF1637) (.3)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.019G038851
409 / 2e-145
AT5G39890 225 / 6e-73
Protein of unknown function (DUF1637) (.1)
Potri.004G129400
235 / 1e-76
AT5G39890 338 / 1e-117
Protein of unknown function (DUF1637) (.1)
Potri.015G055600
216 / 2e-69
AT1G18490 341 / 2e-118
Protein of unknown function (DUF1637) (.1)
Potri.017G079400
214 / 3e-69
AT5G39890 300 / 4e-103
Protein of unknown function (DUF1637) (.1)
Potri.012G057400
208 / 1e-66
AT1G18490 329 / 3e-114
Protein of unknown function (DUF1637) (.1)
Potri.004G058000
197 / 2e-62
AT3G58670 320 / 1e-111
Protein of unknown function (DUF1637) (.1), Protein of unknown function (DUF1637) (.2), Protein of unknown function (DUF1637) (.3)
Potri.011G067300
193 / 7e-61
AT2G42670 316 / 8e-110
Protein of unknown function (DUF1637) (.1), Protein of unknown function (DUF1637) (.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10013620
325 / 4e-112
AT5G39890 225 / 7e-73
Protein of unknown function (DUF1637) (.1)
Lus10003861
322 / 1e-110
AT5G39890 229 / 4e-74
Protein of unknown function (DUF1637) (.1)
Lus10015360
223 / 1e-71
AT5G39890 316 / 6e-108
Protein of unknown function (DUF1637) (.1)
Lus10027254
210 / 1e-66
AT1G18490 324 / 1e-111
Protein of unknown function (DUF1637) (.1)
Lus10038962
209 / 1e-66
AT1G18490 332 / 1e-114
Protein of unknown function (DUF1637) (.1)
Lus10017645
192 / 1e-60
AT3G58670 390 / 2e-139
Protein of unknown function (DUF1637) (.1), Protein of unknown function (DUF1637) (.2), Protein of unknown function (DUF1637) (.3)
Lus10020711
189 / 1e-59
AT3G58670 386 / 8e-138
Protein of unknown function (DUF1637) (.1), Protein of unknown function (DUF1637) (.2), Protein of unknown function (DUF1637) (.3)
Lus10033603
189 / 3e-59
AT3G58670 385 / 3e-137
Protein of unknown function (DUF1637) (.1), Protein of unknown function (DUF1637) (.2), Protein of unknown function (DUF1637) (.3)
Lus10029828
182 / 2e-54
AT3G58670 322 / 4e-109
Protein of unknown function (DUF1637) (.1), Protein of unknown function (DUF1637) (.2), Protein of unknown function (DUF1637) (.3)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0029
Cupin
PF07847
PCO_ADO
PCO_ADO
Representative CDS sequence
>Potri.013G064200.1 pacid=42811745 polypeptide=Potri.013G064200.1.p locus=Potri.013G064200 ID=Potri.013G064200.1.v4.1 annot-version=v4.1
ATGACAATCGAAGCTACGGTGGAGCCAAGGAGAGAACCAACTGCACATGTAAACAGGCTTGGATTTGCCAAGAGGCCTACTAAGAGGAAGAGATCCAAGA
AGACTAAAAAGTGTGCACCGACAATGGCACTGCAGGATCTGTATGTGTCGTGTAAAGAAGTGTTTAAAGGTCCTGGAACTGTTCCTTTGCATCAAGATGT
CAAGAGACTCTGCCACATGCTTGATAACATGAAGCTTGAAGACTTTGGGCTAAGTTGCAAGCTGGAATTCTTCAATCCTAAAGCTGCAGTTAGAGGGACT
CCAAGAGTCACTTACACAATTGTTTACGAGTGCGATAAGTTCTCGATGTGTGTTTTCTTCCTTCCAGCAACTGCTGTCATCCCTCTCCATAACCATCCAG
GAATGACGGTTTTTAGCAAACTTCTCATGGGGACAATGCACGTAAAATCATATGATTGGGTTGATCCTCCGGCTACAGATGAGCCTGACTCACCAGCCCA
AGTGCGATTGGCAAAACTGGAAGCAGACAGTGTGTTTACAGCTCCTTGCCACACTTCGGTATTGTATCCAACAACAGGAGGAAATATCCATCAATTCACC
GCCATTACACCTTGTGCAGTACTCGACGTGCTCGGACCTCCCTATTCTAACGAGGATGGTCGAGATTGCTCATACTACAAAGATTTTCCATACACTGCCT
TCCCAAATGGAGAGATGGGGTCGGAAGAAGAAGAGGGTGACTGTTACGCGTGGTTGGAGGAGATCACGGTGCCTGAGAATTTACAAATGTTTGTGATTAA
GTATTTAGGTCCACAGGTTGATGATAGTAGTAGCTAG
AA sequence
>Potri.013G064200.1 pacid=42811745 polypeptide=Potri.013G064200.1.p locus=Potri.013G064200 ID=Potri.013G064200.1.v4.1 annot-version=v4.1
MTIEATVEPRREPTAHVNRLGFAKRPTKRKRSKKTKKCAPTMALQDLYVSCKEVFKGPGTVPLHQDVKRLCHMLDNMKLEDFGLSCKLEFFNPKAAVRGT
PRVTYTIVYECDKFSMCVFFLPATAVIPLHNHPGMTVFSKLLMGTMHVKSYDWVDPPATDEPDSPAQVRLAKLEADSVFTAPCHTSVLYPTTGGNIHQFT
AITPCAVLDVLGPPYSNEDGRDCSYYKDFPYTAFPNGEMGSEEEEGDCYAWLEEITVPENLQMFVIKYLGPQVDDSSS
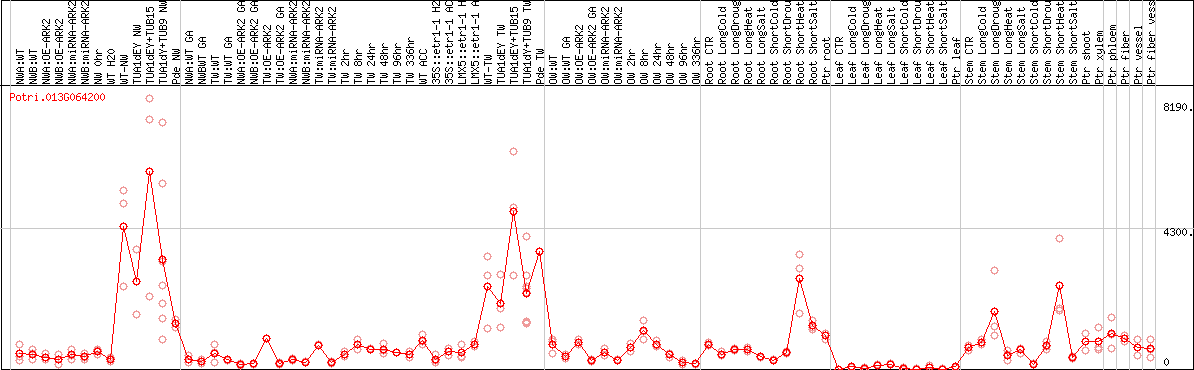
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.013G064200 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.