Potri.013G068400 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.013G068400.9 pacid=42812053 polypeptide=Potri.013G068400.9.p locus=Potri.013G068400 ID=Potri.013G068400.9.v4.1 annot-version=v4.1
ATGGCTCAGCAACAATCTTCACGTCTCAATCGGCTACTCACTTTATTAGACACTGGTTCGACACAAGCTACAAGATTGACTGCTGCGAAACAGATTGGGG
ATATTGCGAAATCGCATCCTCAAGACTTGCATTCTCTCCTGAAGAAGGTTTCTCAGAATCTCCATAGTAAAAACTGGGATACAAGAGTTGCAGCTGCTCA
TGCGATTGGGGCTATTGCTCAGAATGTCAAGCACACTTCTTTGACTGAATTATTTGCTTCTGTCGAAACAAAGATGTCTGAGATTGGGGTTTCGGGTCAT
GTTGAAGATTTGGTGGCATGCCCTAATTTTCATTCTCAAATAATATCAAACGGTTTATTTAGAAGCTTTGATATGAACAAAGTGTTGGAGTTCGGTGCTT
TGTTGGCATCTGGAGGTCAGGAATATGACATTGCAAATGACAATTCTAAGAATCCACGGGAGCGATTGGCTCGCCAGAAACAAAACCTGCGACGTCGTTT
GGGTTTAGATGTTTGTGAGCAGTTTATGGATGTCAACGATGTGATTAAAGATGAAGACCTTGTTGTTCACAGGCCAGAGTCTCAAAGAAACGGTTTAGAT
CATAGATTTTACAAACACCCGTCTGTGCATAACATCCAGCAACTTGTTGCAAGTATGGTTCCAAGTGTAATTTCCAAAAGACCAAGTGCAAGGGAGCTGA
ATCTCCTAAAGCGAAAAGCAAAAATAAATTCCAAAGACCAAGTGAAAAGTTGGTCAGAGGATGGGGATACAGAGGTTGCTTGTCCTCAGAGTACAACTCC
AAAGGGCTCAAATACAGAATCCTTTAGCTTTAAAAAGGCAGATGCTGATGAAGAAGACAACTTGGAGCATGATGGAGATGGGAGGTGGCCTTTCCATGGT
TTTGTTGAACAACTTATTGTTGATATGTTTGACCCTGTTTGGGAGGTTCGCCATGGCAGTGTCATGGCGTTGAGAGAAATCGTAACTCATCATGGTGGTT
CTGCTGGATTGGTTGTGCCTGACTTGAGCTTGGATGGTGCTCTTGATGAACTCAGAGAGAGAGAATATTCAAATACCATTAAAAGAGAGAGAGAAATTGA
TTTGAATTTGCAAGTTTTAACAGATGAGTTTGAGCCAAATCCAAAAAGACATAAATCTGAAGATGTGTCATCCCAGACAATGGATATGATGGTCTCTACT
AGCAATCTCGGTAGTAGTGACATTTGTGTAAAACTTGAGCATAGTGGATGGAATTTGCCTGTTGGGCAGGTTAATAGCCAAGTTGATATCGTTAGTTGTG
TTAAGATGGAGCCCGAATCTTACCCTAATGTTGCATCTTATTCTGCTGAAAGAGCTGTTGGCATGGTAGAGTCAAAGGGTTACCCTGAACATCAGGGTTC
TTTTATGAAGTCAAATTTGCAGAACAGTTCTCCTGAAAATTGCGAGCTGATGAATTTGGTTAAATTGGCCAGACATTCTTCTATAAAAAATAATGAATTT
CTTCAAGATTGTGCAATCCGTTTCTTATGCATATTATCACTAGATCGTTTTGGAGACTATGTATCTGACCAGGTGGTTGCTCCAGTACGTGAAACTTGCG
CCCAAGCATTAGGTGCTGCATTTAAGTATATGCATCACTCTTTAGTTTATGAAACATTGAATATTCTGCTTCAAATGCAGCGTAGGCCAGAATGGGAAAT
TCGTCATGGAAGCCTTTTGGGCATCAAGTATTTGGTTGCTGTTCGACAGGAGATGCTGCCTGATTTGCTTGGCTGCATTCTACCTGCTTGTAAAGCTGGA
CTGGAGGACCCTGATGATGATGTCAGGGCAGTTGCTGCAGATGCTCTAATACCTACTTCTGCTGCAATTGTTTCCATGAAGGGTCGGACATTGCATTCCA
TTGTTATGCTACTTTGGGATATCTTGCTTGATTTGGATGATCTAAGTCCTTCCACTAGCAGTGTGATGAATCTGCTGGCAGAAATATATTCCCAGGAAGA
GATGATTCCCAAGAAAACATCAAAAGATAAACAAGAGTTGGATTTAAATGAAGTGGTACATGTTGATGATGTTGGAGAAGGAAGAGACTTACAAGAAAAT
CCATACATGTTATCGACATTGGCACCACGATTATGGCCATTCATGAGGCATAGTATCACATCAGTCCGACATTCAGCGATTCGAACTTTGGAGCGATTGC
TTGAAGCTGGTTATAAGAGGAACATTTCTGAGCCATCCAGTGCATCATTTTGGCCCTCTTTTATCTTGGGAGATACCCTGCGTATTGTTTTTCAGAATCT
GCTGTTGGAATCAAATGATGAAATTTTGCGTTGTTCAGAGAGGGTTTGGAGGCTCCTTGTCCAGTGCCCAGCAGAGGACTTGGAAGCTGCTGCAAGTTCA
TACATGGCTTCTTGGATAGAACTTACAACTACTCCATATGGCTCACCATTAGATTCTACAAAGATGTTTTGGCCTGTGGCTCCACCTCGAAAGTCTCATT
TTAAAGCAGCTGCTAAAATGAGAGCTGTAAGGCTTGAAAATGAGTCTTGCAGCAGCATTGGCTTAGATTTTGAGAAAGAAACTATTCCTCAGCAAAGAAA
TGGGGATGCTTCTGCTAGCACAGTCAAGATAATAGTAGGTGCTGATGCTGAAATCTCAGTAACCTATACTCGAGTGATTACGGCTTCAGCTTTGGGAATG
TTCGCTTCTAAGTTGCGTGGGGATTCTATGCAACACGTAATAGATCCATTGTGGAATGCTCTCACCTCTTTATCAGGGGTTCAGCGGCAGGTAGCATCAA
TGGTTCTTATATCGTTGTTCAAAGAGATTAAAAGGAAGGAATCCTCTGAAATCCATGGGGTTATGCCTGCTTTTCCCAATCATGTTGAGAAGTTGCTGTT
CGATTTGTTATCTTGTTCTGACCCTGCACTTCCAACAAAAGATTCAGTTCTTCCATATTCTGAGCTCTCAAGGACTTATACAAAGATGCGCAATGAGGCC
AGCCAGTTATTACATGTCACCGAGTCATCTGGCATGTTCAAGAATTCATTGTCAACTATTAAAATAGATGTGGAAAAATTGAGTCCTGATGAGGCAATCA
ATTTTGCATCAAAATTACCATTATCATGCAATGATAGTGCTGGTGATGAATCTACTGGACACAACATTGTTGATGATATTGATTCATCAAAACAGCGACT
TCTGACAACTTCGGGCTATTTAAAATGTGTTCAGAGTAATTTGCATGTTACTGTCTCCGCATTAGTTGCTGCTGCAGTTGTCTGGATGTCAGAGCTTCCT
GCACGGCTGAATCCGATCATTTTGCCCCTGATGGCCTCAATCAAAAGAGAGCAGGAGGAGATATTGCAACAAAAGGCTGCTGAAGCTCTTGCAGAGCTAA
TTTCACGTTGTATTGCACGTAAGCCTGGCCCAAATGATAAGCTGATTAAGAATATTTGTAGTTTGACGTGCATGGATCCTTGTGAGACACCCCAGGCTGG
AGTCATTGGGTCCACAGAGGTTGTTGATGATCAAGATCTCCTCTCCTTTGGGATTAGTACAGGGAAACAAAAGTCAAAGGTTCATATGCTAGCTGGTGGT
GAAGACCGGTCAAGAGTTGAGGGCTTCATTAGCAGACGAGGTTCTGAACATGCATTGAAACATTTATGTGAGAAGTTTGGGGCTTATTTATTTGATAAGC
TACCAAAACTATGGGATTGCCTTGTCGAAGTTCTCAAGCCTGGAAGTCCTGCAGATGAACAACAATTTGAAAAAACTATCGCATCCATTAAGGATCCTCA
AATCCTTATAAATAATATCCAGGTGGTGCGATCTATTGCTCCATTACTGGATGAGGCATTGAAACCTAAATTGCTCACACTTCTCCCTTGCATTTTCAAA
TGTGTGCGCCATTCTCATGTTGCAGTTAGATTAGCTGCCTCAAGGTGTATTACCTCAATGGCCAAATCTATGACAACAAATGTCATGGCAGCTGTAATTG
AGGATGCCATTCCTATGCTAGGGGATGTAACGTCTGTCCATGCCAGACAAGGTGCTGGCATGCTTATTAGTTCACTTGTTCAGGGACTGGGAGTGGAGCT
GGTCCCCTATGCTAGGCTACTGGTTGTTCCTCTTCTACGTTGCATGAGTGATTGTGATCATTCTGTCAGACAGAGTGTGACACGTAGTTTTGCTGCTCTT
GTTCCTCTCCTTCCTTTAGCTCGAGGCCTAGCCCCACCTAGTGGACTAAATGAAGGGTTGGCAAGAAATGCAGAGGATGCACAATTTTTGGAGCAACTAC
TTGACAATTCCCATATTGATGATTACAAGCTCTGCACTGAATTAAAAGTGACATTGAGGAGGTATCAACAAGAAGGTATAAATTGGCTGGCCTTCTTAAA
ACGATTCAAGCTTCATGGAATTCTATGTGATGATATGGGGCTTGGTAAGACCCTGCAAGCATCAGCAATTGTGGCATCTGATGTAGCAGAGTTTCGTGCT
TTGAATAACTGTGAGGATGTTCAACCATCTTTAATTGTTTGCCCATCGACACTTGTGGGACACTGGGCCTTTGAGATAGAGAAGTACATTGATGCTTCTT
TAATTAGTACTCTACAATATAGTGGTTCTGCTCAAGAGCGCATCTGTCTTCGAGAACAATTTCTTAAGCACAATGTCATCATAACATCATATGATGTGGT
TCGCAAAGATATTGATTACCTTGGACAGTCCCTGTGGAATTACTGTATTTTAGATGAAGGGCATATAATCAAGAATGCAAAATCCAAAATTACAGCTGCC
GTGAAGCAGTTGAAAGCCCAGCATCGCCTAATACTTAGTGGAACACCAATTCAGAACAACATCATGGATTTATGGTCCCTGTTTGACTTTTTAATGCCAG
GGTTCCTTGGAACTGACAGACAATTTCAAGCCACTTATGGGAAACCTTTGCTGGCTGCTAGAGATCCTAAATGCTCTGCCAAGGATGCTGAAGCTGGGGT
ACTTGCTATGGAAGCATTGCACAAGCAGGTCATGCCTTTTCTCTTACGAAGAACAAAAGATGAAGTATTATCTGACCTGCCAGAGAAAATTATTCAAGAC
AGATATTGTGACCTGAGCCCTGTCCAGTTAAAACTTTACGAACAGTTCTCTGGTTCCCTTGTTAGACAGGAGATTTCAAGCATGGTAAAGCTGGATGATT
CAGCACAACCAGAAGGAAATAGTGCTTCACCAAAAGCTTCCACACACGTTTTTCAGGCACTTCAATACTTGTTAAAACTTTGTAGTCATCCATTGCTTGT
TGCTGGTGAAAAAATGCCCGAGTCACTTGTATGCCAGTTACATGAACTCTTACCTCCAAATTGTGACATACTTTCGGAACTTCATAAGCTTCACCACTCT
CCAAAACTAGTTGCTCTCCAGGAGATTCTGGAAGAGTGTGGAATAGGTGTGGATGCTTCAAGTTCTGATAATGCTGTGAGTGTTGGGCAGCACCGAGTTT
TGATATTTGCGCAGCACAAAGCCCTTCTTGACATTATTGAAAGGGACTTGTTTCATTCCCAAATGAAGAATGTCACATACTTGCGGCTGGATGGATCTGT
TGAGCCAGAAAAAAGATTTGATATTGTAAAAGCTTTTAATTCAGATCCTACTATTGATGCCTTGTTGCTTACGACTCATGTTGGCGGGCTTGGTTTGAAT
CTTACATCTGCTGACACCCTTGTCTTTATGGAGCATGATTGGAATCCAATGCGAGATCTTCAGGCAATGGACAGGGCACACAGATTGGGGCAAAAGAAAG
TTGTGAATGTCCATCGCCTGATCATGCGTGGCACACTGGAAGAGAAAGTTATGAGTCTCCAAAAGTTCAAAGTGTCGGTTGCCAATGCTGTTATTAATGC
AGAGAATGCCAGTCTGAAAACGATGAACACAGATCAATTGCTCGATTTGTTTGCGTCAGCTGAAACCAGAGCAAAGGGAGCTACTGCATCAAAGCGCACA
GACGGCAGCTTTGATGGAGATCCAAAATTAATGGGCACTGGGAAGGGGTTAAAAGCCATCCTTGGTGGACTGGAGGAGCTCTGGGATCAATCACAGTACA
CTGAAGAGTACAATCTGTCCCAGTTCTTGTCCAAGCTTAATGGCTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.013G068400.9 pacid=42812053 polypeptide=Potri.013G068400.9.p locus=Potri.013G068400 ID=Potri.013G068400.9.v4.1 annot-version=v4.1
MAQQQSSRLNRLLTLLDTGSTQATRLTAAKQIGDIAKSHPQDLHSLLKKVSQNLHSKNWDTRVAAAHAIGAIAQNVKHTSLTELFASVETKMSEIGVSGH
VEDLVACPNFHSQIISNGLFRSFDMNKVLEFGALLASGGQEYDIANDNSKNPRERLARQKQNLRRRLGLDVCEQFMDVNDVIKDEDLVVHRPESQRNGLD
HRFYKHPSVHNIQQLVASMVPSVISKRPSARELNLLKRKAKINSKDQVKSWSEDGDTEVACPQSTTPKGSNTESFSFKKADADEEDNLEHDGDGRWPFHG
FVEQLIVDMFDPVWEVRHGSVMALREIVTHHGGSAGLVVPDLSLDGALDELREREYSNTIKREREIDLNLQVLTDEFEPNPKRHKSEDVSSQTMDMMVST
SNLGSSDICVKLEHSGWNLPVGQVNSQVDIVSCVKMEPESYPNVASYSAERAVGMVESKGYPEHQGSFMKSNLQNSSPENCELMNLVKLARHSSIKNNEF
LQDCAIRFLCILSLDRFGDYVSDQVVAPVRETCAQALGAAFKYMHHSLVYETLNILLQMQRRPEWEIRHGSLLGIKYLVAVRQEMLPDLLGCILPACKAG
LEDPDDDVRAVAADALIPTSAAIVSMKGRTLHSIVMLLWDILLDLDDLSPSTSSVMNLLAEIYSQEEMIPKKTSKDKQELDLNEVVHVDDVGEGRDLQEN
PYMLSTLAPRLWPFMRHSITSVRHSAIRTLERLLEAGYKRNISEPSSASFWPSFILGDTLRIVFQNLLLESNDEILRCSERVWRLLVQCPAEDLEAAASS
YMASWIELTTTPYGSPLDSTKMFWPVAPPRKSHFKAAAKMRAVRLENESCSSIGLDFEKETIPQQRNGDASASTVKIIVGADAEISVTYTRVITASALGM
FASKLRGDSMQHVIDPLWNALTSLSGVQRQVASMVLISLFKEIKRKESSEIHGVMPAFPNHVEKLLFDLLSCSDPALPTKDSVLPYSELSRTYTKMRNEA
SQLLHVTESSGMFKNSLSTIKIDVEKLSPDEAINFASKLPLSCNDSAGDESTGHNIVDDIDSSKQRLLTTSGYLKCVQSNLHVTVSALVAAAVVWMSELP
ARLNPIILPLMASIKREQEEILQQKAAEALAELISRCIARKPGPNDKLIKNICSLTCMDPCETPQAGVIGSTEVVDDQDLLSFGISTGKQKSKVHMLAGG
EDRSRVEGFISRRGSEHALKHLCEKFGAYLFDKLPKLWDCLVEVLKPGSPADEQQFEKTIASIKDPQILINNIQVVRSIAPLLDEALKPKLLTLLPCIFK
CVRHSHVAVRLAASRCITSMAKSMTTNVMAAVIEDAIPMLGDVTSVHARQGAGMLISSLVQGLGVELVPYARLLVVPLLRCMSDCDHSVRQSVTRSFAAL
VPLLPLARGLAPPSGLNEGLARNAEDAQFLEQLLDNSHIDDYKLCTELKVTLRRYQQEGINWLAFLKRFKLHGILCDDMGLGKTLQASAIVASDVAEFRA
LNNCEDVQPSLIVCPSTLVGHWAFEIEKYIDASLISTLQYSGSAQERICLREQFLKHNVIITSYDVVRKDIDYLGQSLWNYCILDEGHIIKNAKSKITAA
VKQLKAQHRLILSGTPIQNNIMDLWSLFDFLMPGFLGTDRQFQATYGKPLLAARDPKCSAKDAEAGVLAMEALHKQVMPFLLRRTKDEVLSDLPEKIIQD
RYCDLSPVQLKLYEQFSGSLVRQEISSMVKLDDSAQPEGNSASPKASTHVFQALQYLLKLCSHPLLVAGEKMPESLVCQLHELLPPNCDILSELHKLHHS
PKLVALQEILEECGIGVDASSSDNAVSVGQHRVLIFAQHKALLDIIERDLFHSQMKNVTYLRLDGSVEPEKRFDIVKAFNSDPTIDALLLTTHVGGLGLN
LTSADTLVFMEHDWNPMRDLQAMDRAHRLGQKKVVNVHRLIMRGTLEEKVMSLQKFKVSVANAVINAENASLKTMNTDQLLDLFASAETRAKGATASKRT
DGSFDGDPKLMGTGKGLKAILGGLEELWDQSQYTEEYNLSQFLSKLNG
|
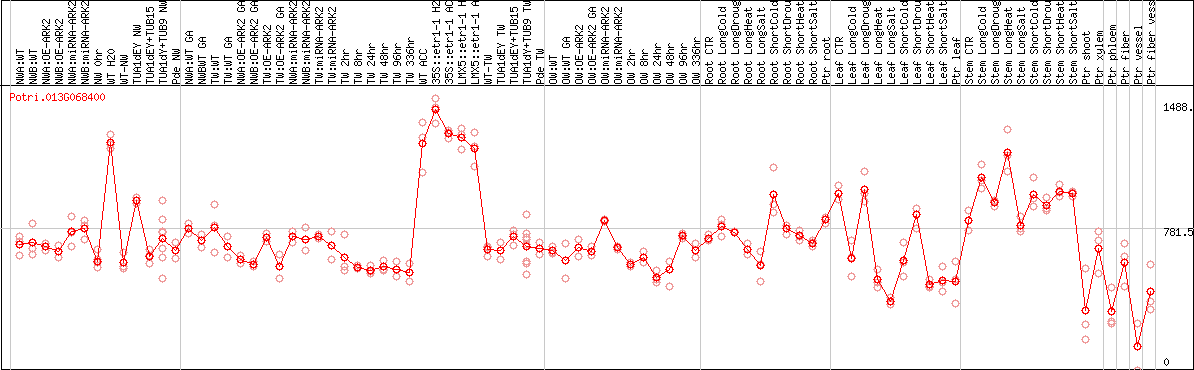
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.013G068400 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
