Potri.013G081300 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.013G081300.8 pacid=42811892 polypeptide=Potri.013G081300.8.p locus=Potri.013G081300 ID=Potri.013G081300.8.v4.1 annot-version=v4.1
ATGTCTTCCTTAAGCAGGGAACTGGTGTTTCTGATCTTGCAGTTCTTGGACGAGGAGAAGTTCAAGGAAACTGTTCATAAGTTAGAGCAAGAGTCTGGTT
TTTTCTTCAATATGAAACATTTTGAGGATCAAGTTCAGGCTGGTGAATGGGATGAGATTGAGCGATATTTATGTGGATTCACGAAGGTTGAAGATAACCG
TTACTCTATGAAGATATTCTTCGAAATAAGAAAGCAGAAATATTTGGAGGCTCTTGATAGACAGGATCGGGCAAAGGCTGTTGAGATTCTTGTTAAGGAC
CTGAAGGTTTTTGCATCCTTCAATGAAGAACTTTTCAAGGAAATAACTCAGTTGCTTACACTTGATAATTTCAGGCAAAATGAGCAGCTTTCGAAATACG
GGGACACAAAATCAGCTCGAAATATCATGCTTGTCGAACTTAAAAAGCTGATTGAAGCAAACCCGCTGTTTCGAGACAAGCTTACTTTCCCACCCTTCAA
AAGTTCGCGACTTAGAACGTTAATAAACCAGAGTCTTAATTGGCAGCATCAACTTTGTAAGAATCCTCGTTCAAACCCTGACATCAAAACTCTATTTATT
GATCATTCTTGCACTCCTACAACTGCTAATGGGGCTCATCCACCTCCTCCCTCGAACACCCCCCTTGTGGGACCGATTCCTAAGGCTGGGGCATTCCCCC
CTATTGGAGCCCATGGCCCTTTTCAACCTGTTGTTTCCCCAACTCCGGGTGCTATTGCTGGTTGGATGTCGGCCAATAATCCTTCTTTACCACACCCTGC
TGTGGCAGCGGGCCCCCCTACTCTTGTGCAGCCTTCTAGTGCAGCTGCCTTCTTGAAACACCCAAGGACTCCCACAGGAATGACAGGAATGAATTATCAG
TCAGCTGATTCAGAACACTTGATGAAGCGGATGCGCCCTGGCCAATCTGAAGAGGTTTCCTTTTCTGGTATAGCACATACTCCCAATATCTATTCCCAAG
ATGACCTTCCCAAAACTGTTGTACGGACCCTTAACCAAGGATCCAATGTCATGAGCATGGACTTTCATCCACAGCATCAAACCATCCTGCTAGTTGGGAC
AAATGTTGGTGACATTAGCCTTTGGGAAGTTGGCTCCCGGGAAAGGTTGGCACATAAACCTTTCAAAGTTTGGGATCTCTCTGCTTCTTCAATGCCACTG
CAGACAGCATTGTTGAATGATGCTGCAATATCTGTAAATCGATGTGTGTGGGGGCCAGATGGACTAATGCTTGGTGTTGCATTTTCGAAACATATAGTGC
AAATATATACTTACAATCCAACTGGAGAACCGAGGCAGCATTTAGAGATTGATGCCCATGTCGGTGGTGTGAATGATATTGCATTTGCTCATCCCAACAA
ACAACTATGCATTGTCACATGCGGTGATGACAAGATGATTAAGGTTTGGGATGCTGGAGCAGGAGGCAGACAATATATATTTGAAGGCCATGAAGCGCCT
GTGTATTCGTTGTGTCCCCATTACAAAGAAAATATCCAGTTTATTTTTTCAACAGCAATTGATGGGAAGATTAAAGCTTGGCTATATGATTCTTTGGGAT
CTAGAGTGGACTATGATGCTCCTGGACTTTGGTGCACCATGATGGCATATAGTGCAGATGGGACTAGGCTCTTCTCTTGTGGAACAAGTAAAGAAGGTGA
ATCTCACCTAGTTGAGTGGAATGAGAGTGAAGGATCTATCAAACGAACATATTTGGGTTTCAGGAAGCGTTCTTTAGATGTTGTTCAGTTTGACACAACA
AGGAGCCACTTCTTAGCTGCTGGTGATGAGTTTCAAATTAAGTTTTGGGATATGGATAATACAAACATGTTGACAGCTGTTGATGCAGATGGGGGGTTGC
CTGCTAGTCCTAGACTGAGATTTAACAAGGAAGGTTCATTGTTGGCAGTCACAACTAGTGACAATGGTATTAAAATTCTAGCCAGTAGTGATGGATTGCG
CTTGATAAGGATGTTGGAGAGCAGGGCTATTGATAAAAGTCGAAGCCCGTCAGAACCAATCAACTCTAAGCCACTGATTGTTAACGCACTAGGCTCTGTG
GCAAATGTTTCTTCTGGCCTCGCATCATCACTGGAACGCTCAGATAGAATCCAGCCTGCAGTTTCTATTGGCAATCTTGGTACCATGGACAACAGTAGGC
TTGTTGATGTTAAACCAAGGATTTCAGATGATACAGACAAATTGAAAAGCTGGAAGAGTGATATTGTAGATTCATCTCAGCTAAAAGCTTTGCGGTTGCC
TGACTCCATAGTAGCTGGCAAGGTTGTGCGGCTGATTTATACTAACTCAGGGCTGGCTCTATTGGCCCTTGCCTCTAATGCTGTCCACAAGCTGTGGAAA
TGGCAGCGCAGTGAAAGAAATCTGACAGGAAAGGCTACTGCATCAAATGCACCACAGTTGTGGCAGCCTCCTAGTGGAACTCCCATGACTAATGATATAA
ATGAGAGCAAACCAGCTGAAGAATCTGCTGCATGCATTGCACTATCCAAAAATGATTCTTATGTCATGTCTGCCTCTGGTGGGAAGGTCTCTTTGTTCAA
CATGATGACATTCAAGGTCATGACAACATTTATGTCCCCACCTCCTGCAGCCACTTTTTTAGCATTCCATCCTCAAGATAATAACATTATTGCAATTGGC
ATGGAGGATTCTACTGTGCAAATATATAATGTCAGAGTTGATGAGGTCAAAACTAAACTCAAGGGCCACCAGAATCGAATAACAGGACTTGCTTTTTCAC
AGAGTTTAAATGTGCTGGTGTCTTCTGGGGCTGATGCACAGTTGTGTGTCTGGAGCATTGATGGATGGGAGAAAAAGAAAATGAGGTTCATTCAAGCACC
ACCTAGTCGTCAATCCCCTTTGGTAGGCGAAACTAGAGTCCAATTTCATAATGACCAAGCACATCTGTTGGTTGTCCATGAAAGCCAAATTGCCATCTAT
GACAGCAAACTTGAATGTTCGCGCTCATGGTCTCCTAAAGACACCCTTGCGGCTCCCATTTCAAGTGCAATATATTCAAGTGATGGTTTCTTGGTTTATA
CGGGTTTTTGTGATGGTGCTGTTGGAGTTTTTGATGCGGACAGCTTAAGAATCAGATGCCGAATAGCACCTTCTGCTTATATACCTTCACACCCTGCAGG
CAGCACTGCCTATCCTTTGGTTATAGCTGCTCATCCATCTGAGCCCAATCAGATTGCACTTGGTATGAGTGATGGTGCAGTACATGTGGTTGAGCCATCT
GATGTGGAGATGAAGTGGGGTGGCCCATCCTCTCAAGATAACGGGACCCATCCGTCTAATACATCGAATCCTTCTCCAAGTGGTCACCTGTCAGAGCTCC
CTTCCAGGTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.013G081300.8 pacid=42811892 polypeptide=Potri.013G081300.8.p locus=Potri.013G081300 ID=Potri.013G081300.8.v4.1 annot-version=v4.1
MSSLSRELVFLILQFLDEEKFKETVHKLEQESGFFFNMKHFEDQVQAGEWDEIERYLCGFTKVEDNRYSMKIFFEIRKQKYLEALDRQDRAKAVEILVKD
LKVFASFNEELFKEITQLLTLDNFRQNEQLSKYGDTKSARNIMLVELKKLIEANPLFRDKLTFPPFKSSRLRTLINQSLNWQHQLCKNPRSNPDIKTLFI
DHSCTPTTANGAHPPPPSNTPLVGPIPKAGAFPPIGAHGPFQPVVSPTPGAIAGWMSANNPSLPHPAVAAGPPTLVQPSSAAAFLKHPRTPTGMTGMNYQ
SADSEHLMKRMRPGQSEEVSFSGIAHTPNIYSQDDLPKTVVRTLNQGSNVMSMDFHPQHQTILLVGTNVGDISLWEVGSRERLAHKPFKVWDLSASSMPL
QTALLNDAAISVNRCVWGPDGLMLGVAFSKHIVQIYTYNPTGEPRQHLEIDAHVGGVNDIAFAHPNKQLCIVTCGDDKMIKVWDAGAGGRQYIFEGHEAP
VYSLCPHYKENIQFIFSTAIDGKIKAWLYDSLGSRVDYDAPGLWCTMMAYSADGTRLFSCGTSKEGESHLVEWNESEGSIKRTYLGFRKRSLDVVQFDTT
RSHFLAAGDEFQIKFWDMDNTNMLTAVDADGGLPASPRLRFNKEGSLLAVTTSDNGIKILASSDGLRLIRMLESRAIDKSRSPSEPINSKPLIVNALGSV
ANVSSGLASSLERSDRIQPAVSIGNLGTMDNSRLVDVKPRISDDTDKLKSWKSDIVDSSQLKALRLPDSIVAGKVVRLIYTNSGLALLALASNAVHKLWK
WQRSERNLTGKATASNAPQLWQPPSGTPMTNDINESKPAEESAACIALSKNDSYVMSASGGKVSLFNMMTFKVMTTFMSPPPAATFLAFHPQDNNIIAIG
MEDSTVQIYNVRVDEVKTKLKGHQNRITGLAFSQSLNVLVSSGADAQLCVWSIDGWEKKKMRFIQAPPSRQSPLVGETRVQFHNDQAHLLVVHESQIAIY
DSKLECSRSWSPKDTLAAPISSAIYSSDGFLVYTGFCDGAVGVFDADSLRIRCRIAPSAYIPSHPAGSTAYPLVIAAHPSEPNQIALGMSDGAVHVVEPS
DVEMKWGGPSSQDNGTHPSNTSNPSPSGHLSELPSR
|
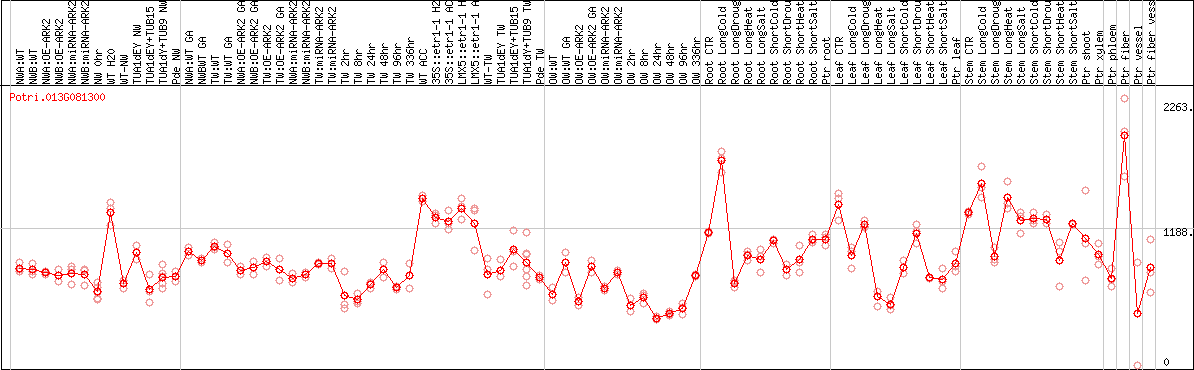
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.013G081300 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
