External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G35460 223 / 2e-72
bHLH
bHLH080
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT4G09180 210 / 1e-67
bHLH
bHLH081
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT2G42280 122 / 2e-32
bHLH
bHLH130
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2)
AT1G51140 114 / 1e-29
bHLH
bHLH122
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT1G05805 107 / 5e-27
bHLH
bHLH128
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
AT2G43140 99 / 3e-24
bHLH
bHLH129
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2)
AT2G24260 67 / 1e-12
bHLH
LRL1, bHLH066
LJRHL1-like 1 (.1)
AT4G30980 66 / 3e-12
bHLH
LRL2, bHLH069
LJRHL1-like 2 (.1)
AT5G58010 63 / 3e-11
bHLH
LRL3, bHLH082
LJRHL1-like 3 (.1)
AT4G02590 61 / 1e-10
bHLH
bHLH059, UNE12
unfertilized embryo sac 12, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.3)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.019G079900
346 / 5e-121
AT1G35460 228 / 1e-74
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.016G050500
126 / 4e-34
AT2G42280 296 / 2e-98
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2)
Potri.006G057600
121 / 1e-32
AT2G42280 278 / 3e-92
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2)
Potri.003G207200
118 / 1e-30
AT2G42280 221 / 2e-68
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2)
Potri.001G017300
115 / 6e-30
AT1G51140 237 / 7e-75
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.009G064700
109 / 1e-27
AT2G42280 225 / 5e-70
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2)
Potri.014G150600
105 / 2e-26
AT1G05805 265 / 1e-86
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.002G231700
100 / 1e-24
AT1G05805 274 / 6e-90
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Potri.001G270000
96 / 1e-22
AT2G42280 198 / 7e-60
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10025518
106 / 1e-26
AT1G05805 242 / 2e-77
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10031255
100 / 9e-26
AT2G42280 172 / 5e-53
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2)
Lus10037484
102 / 5e-25
AT2G42280 191 / 2e-57
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2)
Lus10015704
102 / 7e-25
AT2G42280 194 / 2e-58
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2)
Lus10031826
100 / 1e-24
AT1G51140 182 / 3e-54
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1)
Lus10021023
101 / 3e-24
AT3G12160 384 / 3e-132
ARABIDOPSIS THALIANA RAB GTPASE HOMOLOG A4D, RAB GTPase homolog A4D (.1)
Lus10026733
93 / 4e-23
AT2G43140 179 / 2e-56
basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2)
Lus10023829
69 / 4e-13
AT3G12160 377 / 5e-132
ARABIDOPSIS THALIANA RAB GTPASE HOMOLOG A4D, RAB GTPase homolog A4D (.1)
Lus10021846
65 / 6e-12
AT2G24260 275 / 3e-90
LJRHL1-like 1 (.1)
Lus10014726
62 / 2e-11
AT4G02590 278 / 8e-96
unfertilized embryo sac 12, basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.1), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.2), basic helix-loop-helix (bHLH) DNA-binding superfamily protein (.3)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF00010
HLH
Helix-loop-helix DNA-binding domain
Representative CDS sequence
>Potri.013G107500.1 pacid=42812342 polypeptide=Potri.013G107500.1.p locus=Potri.013G107500 ID=Potri.013G107500.1.v4.1 annot-version=v4.1
ATGCATCCAGCACCGGCAGGAAGCAGCAACACTAGTGGCGGTGGATGTGGCGGTGGCGGAGGAGGAGGTGGTGGTGGTGGTGGTGGGCTTCCTAGGCTCC
GTTCAGCACCAGCCACATGGTTACTAGCACTTCTTGAAGAAGAAGAAGAGGACCCGTTAAAGCAAAATCAGAACTTGACTCAGCTACTCACTTCAAACGC
ACCTTCAAGTAGAAATTCAGCGCCCTTCAATGCTTCCAGTGCTGCTGTGGAACCGGGTTTGTATGAAACGGGTAGTGGTTTTCAAAGGCAAAACAGCTCC
CCAGCTGATTTTCTAGGGAATTCTGGTATTGGAAGTGATCAGGGGTATTTTTCGAACTATGGAATTGCTTCAAATTATGAGTATATGCCGCCAAATATGG
AGGTTTCACCTTCTGCTAAAAGGGCTAGAGAGCTTGAGTTGCAAAACCCTCCTGCTAGATATCCACCTCCTTTGAAAGGAGCGCAAACTGGATCTCTAAG
GGCTTCTAGTTTAATAGAAATGGAGATGGACAAGCTTTTGGAGGAATCAGTGCCTTGCAAGATTCGTGCCAAGCGTGGCTGTGCTACACATCCTCGGAGC
ATTGCGGAGAGAGTTCGGAGAACTAGAATTAGTGACCGGATAAGGAAACTTCAAGAACTTGTTCCGAATATGGATAAGCAAACAAACACTGCAGATATGC
TGGAGGAGGCAGTAGACTATGTCAAGTTTCTTCAAAGGCAGATTCAGGAACTCACGGAACAGCAAAGAAAATGCAAATGCATGGCTAAAGAATAA
AA sequence
>Potri.013G107500.1 pacid=42812342 polypeptide=Potri.013G107500.1.p locus=Potri.013G107500 ID=Potri.013G107500.1.v4.1 annot-version=v4.1
MHPAPAGSSNTSGGGCGGGGGGGGGGGGGLPRLRSAPATWLLALLEEEEEDPLKQNQNLTQLLTSNAPSSRNSAPFNASSAAVEPGLYETGSGFQRQNSS
PADFLGNSGIGSDQGYFSNYGIASNYEYMPPNMEVSPSAKRARELELQNPPARYPPPLKGAQTGSLRASSLIEMEMDKLLEESVPCKIRAKRGCATHPRS
IAERVRRTRISDRIRKLQELVPNMDKQTNTADMLEEAVDYVKFLQRQIQELTEQQRKCKCMAKE
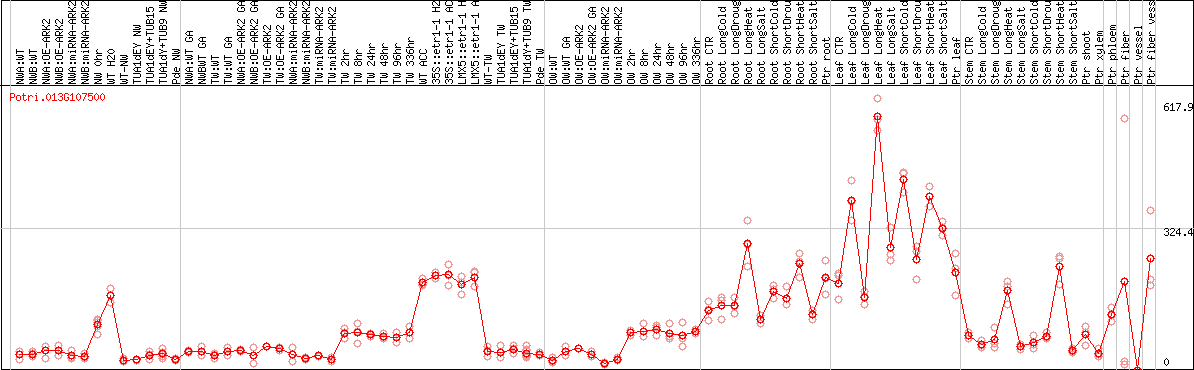
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.013G107500 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.