External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G16920 145 / 2e-38
UBC23 ,PFU2
PHO2 FAMILY UBIQUITIN CONJUGATION ENZYME 2, ubiquitin-conjugating enzyme 23 (.1)
AT3G15355 134 / 5e-35
UBC25 ,PFU1
PHO2 FAMILY UBIQUITIN CONJUGATION ENZYME 1, ubiquitin-conjugating enzyme 25 (.1)
AT2G33770 132 / 3e-34
ATUBC24, UBC24, PHO2
UBIQUITIN-CONJUGATING ENZYME 24, phosphate 2 (.1)
AT1G53025 129 / 1e-33
Ubiquitin-conjugating enzyme family protein (.1)
AT1G53023 123 / 1e-32
Ubiquitin-conjugating enzyme family protein (.1)
AT3G12775 74 / 9e-15
ubiquitin-conjugating enzyme family protein (.1)
AT3G13550 54 / 1e-08
EMB144, COP10, CIN4, FUS9
FUSCA 9, EMBRYO DEFECTIVE 144, CONSTITUTIVE PHOTOMORPHOGENIC 10, CYTOKININ-INSENSITIVE 4, Ubiquitin-conjugating enzyme family protein (.1.2)
AT2G16740 53 / 1e-08
UBC29
ubiquitin-conjugating enzyme 29 (.1)
AT4G27960 52 / 4e-08
UBC9
ubiquitin conjugating enzyme 9 (.1.2)
AT5G53300 52 / 4e-08
UBC10
ubiquitin-conjugating enzyme 10 (.1.2.3.4)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.010G146500
159 / 1e-45
AT2G33770 255 / 3e-77
UBIQUITIN-CONJUGATING ENZYME 24, phosphate 2 (.1)
Potri.011G118800
162 / 8e-45
AT3G15355 411 / 1e-135
PHO2 FAMILY UBIQUITIN CONJUGATION ENZYME 1, ubiquitin-conjugating enzyme 25 (.1)
Potri.008G104400
155 / 2e-44
AT3G15355 267 / 7e-84
PHO2 FAMILY UBIQUITIN CONJUGATION ENZYME 1, ubiquitin-conjugating enzyme 25 (.1)
Potri.001G400300
159 / 8e-44
AT3G15355 383 / 5e-125
PHO2 FAMILY UBIQUITIN CONJUGATION ENZYME 1, ubiquitin-conjugating enzyme 25 (.1)
Potri.013G130800
151 / 1e-41
AT3G15355 334 / 1e-107
PHO2 FAMILY UBIQUITIN CONJUGATION ENZYME 1, ubiquitin-conjugating enzyme 25 (.1)
Potri.004G043500
145 / 1e-38
AT2G33770 823 / 0.0
UBIQUITIN-CONJUGATING ENZYME 24, phosphate 2 (.1)
Potri.011G052600
145 / 1e-38
AT2G33770 796 / 0.0
UBIQUITIN-CONJUGATING ENZYME 24, phosphate 2 (.1)
Potri.009G136000
142 / 2e-37
AT2G16920 1069 / 0.0
PHO2 FAMILY UBIQUITIN CONJUGATION ENZYME 2, ubiquitin-conjugating enzyme 23 (.1)
Potri.004G176000
132 / 4e-34
AT2G16920 1045 / 0.0
PHO2 FAMILY UBIQUITIN CONJUGATION ENZYME 2, ubiquitin-conjugating enzyme 23 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10037778
155 / 2e-44
AT2G16920 256 / 9e-81
PHO2 FAMILY UBIQUITIN CONJUGATION ENZYME 2, ubiquitin-conjugating enzyme 23 (.1)
Lus10016921
150 / 3e-43
AT2G16920 235 / 3e-70
PHO2 FAMILY UBIQUITIN CONJUGATION ENZYME 2, ubiquitin-conjugating enzyme 23 (.1)
Lus10040748
151 / 9e-43
AT2G16920 244 / 1e-72
PHO2 FAMILY UBIQUITIN CONJUGATION ENZYME 2, ubiquitin-conjugating enzyme 23 (.1)
Lus10028151
153 / 3e-42
AT1G53025 388 / 2e-129
Ubiquitin-conjugating enzyme family protein (.1)
Lus10005854
151 / 1e-40
AT2G33770 785 / 0.0
UBIQUITIN-CONJUGATING ENZYME 24, phosphate 2 (.1)
Lus10042852
146 / 5e-40
AT1G53025 318 / 5e-103
Ubiquitin-conjugating enzyme family protein (.1)
Lus10034472
149 / 7e-40
AT2G16920 907 / 0.0
PHO2 FAMILY UBIQUITIN CONJUGATION ENZYME 2, ubiquitin-conjugating enzyme 23 (.1)
Lus10025070
148 / 2e-39
AT2G16920 1209 / 0.0
PHO2 FAMILY UBIQUITIN CONJUGATION ENZYME 2, ubiquitin-conjugating enzyme 23 (.1)
Lus10018596
143 / 7e-39
AT3G15355 339 / 3e-111
PHO2 FAMILY UBIQUITIN CONJUGATION ENZYME 1, ubiquitin-conjugating enzyme 25 (.1)
Lus10001214
141 / 1e-37
AT2G33770 508 / 9e-171
UBIQUITIN-CONJUGATING ENZYME 24, phosphate 2 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0208
UBC
PF00179
UQ_con
Ubiquitin-conjugating enzyme
Representative CDS sequence
>Potri.013G108500.1 pacid=42811836 polypeptide=Potri.013G108500.1.p locus=Potri.013G108500 ID=Potri.013G108500.1.v4.1 annot-version=v4.1
ATGGATGCTACAGAATATGAAGAAACATATGTTGAAGGAATTGAGGAGATAGAAGATGATGTAGACAGGAGGTTCAAGAGATTCAAAAGCTTCAGGGTGC
TTCAATATCCTCCTCATGACCATCAATTTAGGCACAAACAAAGACTTTGTAGCTGGATGCTTTCAGCCACCAGATTTGGTGAGCATGCAAATCCCAGGAG
CAGCTTTTCTGAAAAGATAAAAAAGGACTGGGAAACATTGGCGGCTGTTTTTTCAGATCATAGTTCGATTTTTGTTACAACTTACGAGTCGAGAATTGAT
CTCATGAGGGTTTGCATTGCTGGTTTAGAAGGAACTCCATACTGTTATGGCTTGTTCTTTTTTGACATATTCTTCCCTCCTGAGTACCCAGCTAAGCCTC
CTGAAATCTTTTACCATGCTCCTGACTTTGATTTAAGCCCTTGTTTGCATCAAGATGGAAGGGTTTCTCTCAACCTCTTGAGTCTTAATCAATGGTATAG
GTTTCGGTTAGGAGGAAAACAGCAATCATGGAATCCGGAACGATCCGATATATCGAGGGTTCTTCTCTCCATCCAACATTTGATTTTGAATGATAAACCT
TATCTGAACGAGTCTATTAGTTTTGTGTACTCTTCTAAAGAGAGATCCTCAAAATATAACAATGAGGTGTTCATGTCATCTTGTGAGGCCATGCTTGCCA
TGCTCAGGTTTTCTCCAGGTGATTGCGGTGATTTTGTGTTGGGGCATTTTCGAAAGAGAGCTCACCGGATTCTCTTGATTTATAAAGATCAAATGCTGAA
ACATAAAGGTGATAATGGTATGAAGCAGTTGTTCTTCAAGCTTGTGAAAGCTTTTGAAAGCAATGGAGCTTATTGCCAGCATCATTGTAGCAAAACAGAG
GTTGAGCAATTGAAGAAAGAGAGATTAAACACGGGAGGATTTTCCATGGAAGAATGTTCTGAATGA
AA sequence
>Potri.013G108500.1 pacid=42811836 polypeptide=Potri.013G108500.1.p locus=Potri.013G108500 ID=Potri.013G108500.1.v4.1 annot-version=v4.1
MDATEYEETYVEGIEEIEDDVDRRFKRFKSFRVLQYPPHDHQFRHKQRLCSWMLSATRFGEHANPRSSFSEKIKKDWETLAAVFSDHSSIFVTTYESRID
LMRVCIAGLEGTPYCYGLFFFDIFFPPEYPAKPPEIFYHAPDFDLSPCLHQDGRVSLNLLSLNQWYRFRLGGKQQSWNPERSDISRVLLSIQHLILNDKP
YLNESISFVYSSKERSSKYNNEVFMSSCEAMLAMLRFSPGDCGDFVLGHFRKRAHRILLIYKDQMLKHKGDNGMKQLFFKLVKAFESNGAYCQHHCSKTE
VEQLKKERLNTGGFSMEECSE
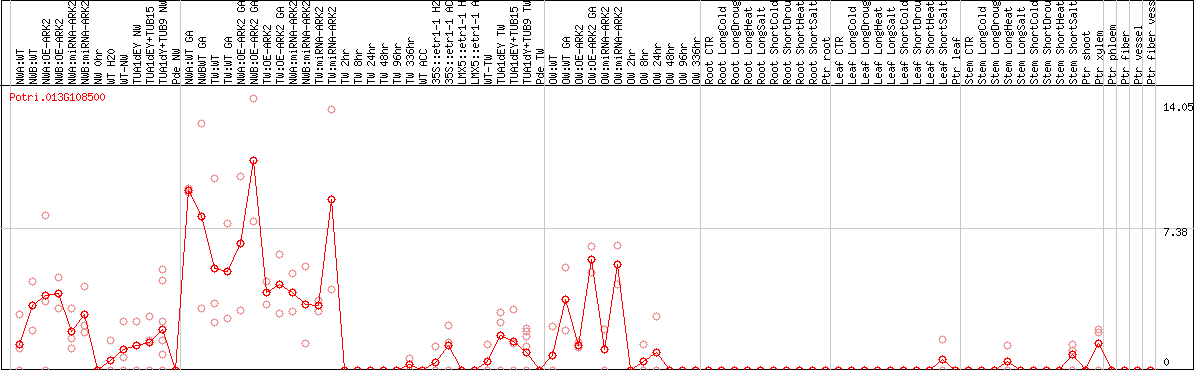
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.013G108500 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.