Potri.013G119900 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.013G119900.1 pacid=42811324 polypeptide=Potri.013G119900.1.p locus=Potri.013G119900 ID=Potri.013G119900.1.v4.1 annot-version=v4.1
ATGAGTGGAGGCAAGCCTTCTTCAAAAAGGAAAGGAGTTGAAAAATATGAAGATGCATCAGTAAATACAGAAGTAGAACTTCATGTTGAACATGAGGCTA
TTTTTCATGATCATGTACCAATTGTTTCGTATTACAACGACCGTATCACACCTCTCCTTGATGCAGTGGACAGGCTGAGGCAACTCCAGGTAATGAAAGA
AGGCATACAACTTCCAACCATTGTTGTTGTTGGCGATCAATCTTCAGGCAAATCCAGTGTTCTTGAATCATTGGCTTGTATCAATCTTCCTCGTGGCGAT
GGCATTTGCACTAGAGTGCCTCTCATTGTGAGGCTTAAACATCACCCATCTCTTGTACCAGAGATTTTCTTGCAGTTCAATGGCAAAACAGTGCCTACTG
ATGAAGCCCACGTTGCTGATGCTATCAATCTTGTCACTGATGAGATAGCAGGCAACGGTAAGGGCATATCTAACACTGAATTAACTCTTGTAGTGAAAAA
GAATGGTGTTCCTGATCTTACTCTTGTTGATCTCCCTGGAATAACAAGAGTTCCTGTTCATGGTCAACCTGAAAATATCTATGAGCAGATAGCATACATT
ATAATGAAGTATATTAGTCCTGATGAGAGTGTTATCCTTAATGTTTTGTCTGCGAGCGTTGATTTTTCCACATGTGAATCCATAAGGATGTCACAGAAAG
TCGACAAGAACGGTCAGAGGACTCTTGCTGTGGTTACAAAGGTTGATAAATCACCTGAAGGGTTACTTGAGAAGGTCACTAGAAATGATGTGAATATAGG
CCTTGGTTATGTGTGTGTTAGAAACCGTATTGGTAATGAGTCTTATGAGGATGCAAGGAAGGAAGAAGCTGCACTGTTTGCTACACATCAACTTTTGTCC
AAGATTGATAAATCTACAGTGGGTATTCAAGTTTTGGCTCAAAAACTGGTGCAAATTCAGGCTAATATTATTGCCAAGTGTTTGCCTGATATCGTCAGAA
AGATTGATGAGAAGTTGAGTGCGAGTATTTCAGAGCTGAATAGGATACCTAGGAGGTTGTTGTCAGTTGCTGAGGTCATGGCAGCTTTCATGGGCATTAT
TGGATCTTCCAAGGACTCTTTGAGGAAAATCCTTTTAAGAGGGGAAATTGATGAATACCGACATGAAAAAGATATGCACTGCACTGCTAGATTGGTTGAA
ATGCTCAATCAATTCTCAACAGAACTTCATAAGTGTTCTGACCATACAAAGAATTTTATGATAAATGAGATTGAGGTTTTGGAGGAAACAAAAGGGATTG
AATTGCCAAATTTCCTTCCTCACGCTGCCATCCTTGCCATCCTGCAGCAAAAAGTTGAAGAAATATCAGAACTACAAATAGGGTTTGTTGAAAAAGTGTG
GGCTTACATTCGAGGTGTGGTCATTTCAGTTTTGAATCATCACTCGGCGAACTATCACCAACTTCAGTTATTTATCGGACGAGCGGCCCATAAACTTGTA
GATAAAATGAAAGACCGGTCCATTGATTGGGTAACAGAGATTCTCCAGATGGAGAAGGAAACGGATTACACATGTAATCCAGAATATATGAAGGAATGGA
ACAAGCTCATCGCACAGCAGCAAGCAGTCATCGACAACATTACGAAATTTGGATCTTCCAGGGTGACGATTGATGGCTCGAGAGAGGTTGTAGTGGGAGA
TCTTAGGGGTCATAAACATGTTCTTCTTCAAGCTTTTGACTTGAAGATGAGGCTGATTGCTTACTGGAAGATTGTCTTGATGAGGTTGGTTGACAATATG
GCTCTGCATCTCCAGCTCAGCATCCGAAACTTGGTGAACAAGGAGATGGAAAAGGAGATTGTTAATGCGTTGTTGGGTACTGGTGGTGGTGTTGCAATAG
AAAGAATGTTAGAGGAACCTCCATCTGTTGCTAGCAAGCGTGAGAGGCTGAACACAAGTATCAAGTTGTTGAGGGAGTCTAAAGAGGTGCTGGCAAATAT
CAGAGACAAAATTGAATGTGGTGATCACTAG
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.013G119900.1 pacid=42811324 polypeptide=Potri.013G119900.1.p locus=Potri.013G119900 ID=Potri.013G119900.1.v4.1 annot-version=v4.1
MSGGKPSSKRKGVEKYEDASVNTEVELHVEHEAIFHDHVPIVSYYNDRITPLLDAVDRLRQLQVMKEGIQLPTIVVVGDQSSGKSSVLESLACINLPRGD
GICTRVPLIVRLKHHPSLVPEIFLQFNGKTVPTDEAHVADAINLVTDEIAGNGKGISNTELTLVVKKNGVPDLTLVDLPGITRVPVHGQPENIYEQIAYI
IMKYISPDESVILNVLSASVDFSTCESIRMSQKVDKNGQRTLAVVTKVDKSPEGLLEKVTRNDVNIGLGYVCVRNRIGNESYEDARKEEAALFATHQLLS
KIDKSTVGIQVLAQKLVQIQANIIAKCLPDIVRKIDEKLSASISELNRIPRRLLSVAEVMAAFMGIIGSSKDSLRKILLRGEIDEYRHEKDMHCTARLVE
MLNQFSTELHKCSDHTKNFMINEIEVLEETKGIELPNFLPHAAILAILQQKVEEISELQIGFVEKVWAYIRGVVISVLNHHSANYHQLQLFIGRAAHKLV
DKMKDRSIDWVTEILQMEKETDYTCNPEYMKEWNKLIAQQQAVIDNITKFGSSRVTIDGSREVVVGDLRGHKHVLLQAFDLKMRLIAYWKIVLMRLVDNM
ALHLQLSIRNLVNKEMEKEIVNALLGTGGGVAIERMLEEPPSVASKRERLNTSIKLLRESKEVLANIRDKIECGDH
|
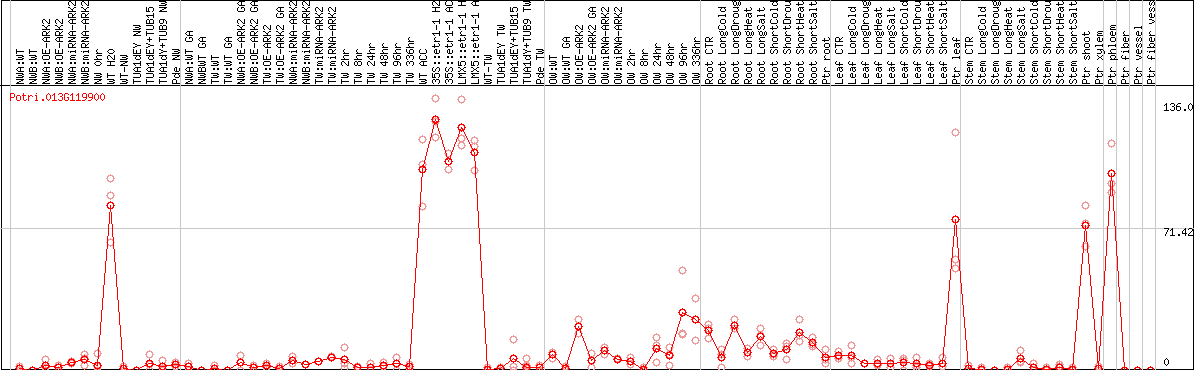
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.013G119900 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
