Potri.013G131300 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.013G131300.2 pacid=42810965 polypeptide=Potri.013G131350.1.p locus=Potri.013G131300 ID=Potri.013G131300.2.v4.1 annot-version=v4.1
ATGTCTCTCCGTCGCCCGCCCCCTCCGAACCCGGCCCGAACCGAATCCCAACCCTACAACATAATCCCCATCCAGAACCTCCTTGCCGACCACCCTTCCC
TCCGCTACCCCGAAGTTCGAGCAGCCGCCGCCGCCTTACGCACCGTCGGCAACCTCCGTAAACCACCCTACGCTCAATGGCACCCTTCAATGGACCTCCT
CGACTGGCTTGCTCTGCTCTTCGGCTTCCAAAAGGACAACGTTCGCAACCAGCGGGAGCACCTAGTCCTTCACCTTGCAAACGCTCAGATGCGGCTGACT
CCTCCGCCGGACAACATTGATACCCTAGACGCCGGTGTACTCCGTCGGTTCAGGCGGAAACTGCTGAAGAATTATACAAATTGGTGTGATTACTTGAATA
AGAAGTCGAATATCTGGATCTCTGACCGGTCGACGGATCTGAGAAGGGAGTTACTGTATGTCTCGTTGTATTTGTTAATTTGGGGGGAATCGGCGAATTT
ACGGTTTATGCCGGAGTGTATTTGCTTTATATTTCATAATATGTGTTTTGAATTGAATAGAGTTTTAGAGGATTATATTGATGAGAATACAGGGCAGCCG
GTGATGCCGTCAATTTCAGGGGAGAATGCGTTTTTGAATGGTGTTGTGAAGCCAATTTATGAGACGGTGAGAAGGGAAGTGGATAGGAGCTTTAACGGGG
CTGCGCCACATAGTGCTTGGCGGAATTATGATGATTTGAATGAGTATTTTTGGAGTAAGAGGTGTTTTGAGAGGTTGAAATGGCCGATTGATTTAGGGAG
TAATTTTTTTGTGACTTCGGGATCGAGGAAGAAGGTGGGGAAGACGGGGTTTGTGGAGCAACGGTCGTTTTGGAATATTGTAAGGAGTTTTGATAGGTTG
TGGGTGATGTTGATTTTGTTTTTGCAAGCGGGGATTATTGTTGCGTGGGAGGAGAAGGAGTATCCGTGGAAAGCGTTGAAGAGCAGGGATGTGCAGGTTA
GGGTGTTGACTGTTTTTTTTACTTGGAGTGGGTTGAGGTTCTTGCAGTCATTGCTTGATGTCGGGACACAGTATAATTTGGTTTCGAGGGAGACTTTGGG
GCTTGGGGTTAGGATGATTTTGAAAAGTGTGGTTGCAGTAGTGTGGATTATTGTGTTTGGTGCATTTTATGGGAGGATTTGGAGTCAGAGGAACAGTGAT
CTGAGGAGGTCGCCCAGGGATCTGAGTTGGTCATCGGAGGCTGATAAGAAGGTAGTGACTTTTCTTGAGGTTGCTTTGGTGTTTGTGGCGCCAGAGATAT
TGGCATTGGCTCTGTTTATTCTTCCATGGATTAGGAATTTTCTTGAGAATACGGATTGGAGGATATTTCGGATGATGACATGGTGGTTTCAGAGTAGTAG
TTTTATTGGTAGAGGGTTGAGGGAGGGGCTTGTGGATAATATTAAATATACTTTGTTTTGGGCTATGGTTTTAGCTACCAAATTTGCTTTCAGTTACTTT
ATGCAGATTAAACCCATGGTTAAACCATCAAAACAGATGCTGAAGCTGAAGGATGTGAATTATGAATGGCACGAGTTTTTTGACCACAGCAATAGGTTTT
CAGTTGGGTTGCTGTGGCTTCCTGTGGTGTTGATTTACTTGATGGACTTGCAGATTTGGTATGCCATTTATTCCTCTTTTGTTGGAGCAGGAGTGGGGTT
GTTTCAACATTTGGGTGAGATTCGAAACATCCAGCAATTAAGGTTGAGATTTCAGTTCTTTGCAAGTGCAATTCAGTTTAATCTGATGCCAGAGGAGCAG
CTGTTGAATGCAAGGGGTACGTTCAAGAGCAAGTTCAAAGATGCCATTCACAGGTTGAAGCTTAGGTATGGGTTTGGCCACCCTTACAAGAAGCTGGAGT
CTAACCAGGTAGAGGCAAACAAGTTTGCTTTGATATGGAATGAGATCATAATAATTTTCAGGGAGGAGGATATTATCTCTGACAAGGAGCTTGAGTTGAT
GGAGTTGCCACAGAATTCTTGGAATGTGAGGGTGATTCGCTGGCCAAGTTTTCTCCTGTGCAATGAGCTGCTGCTTGCTCTTAGCCAGGCCAAAGAGTTG
GTAGATGCTCCTGATAAGTGGCTCTGGTACAAGATATGCAAGAACGAGTATAGGCGCTGTGCGGTCATAGAAGCTTATGATAGTGTCAAGCACCTGTTAC
TTGAAATCATCAAGACCAACACAGAAGAGCACTCAATTATCACGGTTTTGTTTCAAGAAATTGATCACTCTCTGCAGATTGAGAAATTCACCAAGACTTT
CAAGATGACAGGTCTGCCTAATTTCCATGCCAAGTTGATAAAGCTTCTTGAGCTGTTGAACAAGCCCAAGCGGGATCTGAACCAGGTGGTAGATACTCTA
CAGGCTCTATATGAGATTGCTGTACGAGAATTTTTCAGAGAGAAGAGGAGCACTGAACAGTTGATGGAGGACGGGTTGGCTCCTCGTGACCCAGCTGCCA
TGGCTGGGCTTCTTTTTGGGAATGCAGTTCAGTTGCCTGATGCTAGCAATGAGACCTTTTATAGGCAGGCACGGCGTTTGCACATGATTCTTACCTCTAG
GGATTCGATGAACACTATCCCAGAAAATCTAGAGGCCAGGCGCAGAATTGCATTTTTCAGCAATTCCCTGTTCATGAGCATGCCCCACGCTCCCCAGGTT
GAGAAAATGATGGCTTTTAGTGTGCTGACCCCTTATTACAATGAGGAGGTGCTATACAGCAGAGAACAGCTTCGAACTGAAAATGAAGATGGGGTTTCCA
CACTGTACTACCTGCAAACTATTTATGCTGATGAGTGGAAAAACTTCATGCAGAGGATGCGCCGTGAAGGAATGGAAAAGGATGGTGAGATATGGACAAC
CAAGTTGAGAGATCTGAGGCTCTGGGCATCTTATAGAGGCCAGACGCTTGGCCGTACTGTGAGGGGAATGATGTATTATTACCGTGCTCTTAAGATGCTG
GCTTTTCTTGATTCTGCCTCGGAGATGGACATTAAAGAGGGTTCACGAGAACTTGGCTCGATGAGGCGAGACAATGGTTTGGATAGCTTTGACTCAGAAA
GTTCTTCTAAGAGCTTAAGCAGAAATAGCAGTTCAGTGAATTTGTTGTTTAAAGGTCATGAGTATGGGACTGCTTTGATGAAATACACATATGTGGTTGC
CTGCCAGATATACGGGGCACAAAAGGCAAAGAAGGATCCCCATGCTGAGGAAATCTTGTATCTGATGAAAAATAATGAGGCCCTTCGCGTTGCCTATGTT
GATGAAGTAAACACTGGGAGGGATGAGATGGAATATTATTCTGTACTTGTGAAGTATGATCAGCAGTTGGACAAGGAAGTGGAAATCTACAGGGTGAAGT
TGCCGGGTCCCTTGAAACTCGGTGAGGGAAAACCAGAGAATCAAAATCATGCCCTAATCTTCACTCGTGGGGATGCAGTGCAGACTATTGATATGAACCA
GGATAACTATTTTGAAGAGGCTCTCAAAATGCGGAATCTTTTGGAAGAATACAGGCACTACTATGGAGCTCGTAAACCTACTATCTTGGGAGTCAGGGAA
CACATTTTTACTGGTTCTGTATCATCTCTTGCATGGTTTATGTCTGCTCAGGAAACTAGTTTTGTCACCCTGGGTCAGCGTGTTTTGGCAAACCCTTTGA
AAATTCGAATGCATTATGGCCATCCAGATGTCTTTGACAGGTTTTGGTTCATGACTAGAGGTGGAATCAGCAAGGCTTCCAGAGTGATTAACATTAGTGA
GGACATATTTGCTGGCTTTAATTGCACCTTGAGAGGAGGCAATATTACTCACCACGAATACATCCAAGTTGGAAAAGGAAGGGATGTTGGGTTGAATCAA
ATATCCATGTTTGAAGCAAAAGTTGCCAGTGGAAATGGCGAGCAAACTCTTAGCAGAGATGTCTATAGATTGGGCCATAGGCTGGACTTCTTCCGTATGC
TGTCATTCTTTTATACTACGGTGGGATTTTTTTTGAACACTATGATGGTCATTCTGACTGTGTATGCATTTCTGTGGGGCCGTCTTTATCTGGCTCTTAG
CGGTGTTGAGGGTTCTGCTTTGGCCGACAACAGCAGTAACAATAAGGCACTTGGTGCTATTTTGAATCAGCAATTCATCATCCAACTTGGCCTCTTCACT
GCCCTTCCGATGATAGTGGAGAACTCTCTTGAGCACGGGTTTCTCGAAGCTATCTGGGATTTCTTGACAATGCAGCTCCAGCTCTCATCTGTTTTCTACA
CATTCTCTATGGGAACTCGGACGCACTACTTTGGCCGTACCATCCTCCATGGTGGCGCAAAATATCGGGCAACTGGGCGTGGTTTTGTTGTGCAGCACAA
GAGTTTTGCAGAGAATTACAGGCTCTATGCTCGTAGCCATTTTGTGAAGGCAATTGAGCTTGGACTGATACTTGTAGTTTATGCAGCATACAGCCCTGTA
GCTAAGGACACATTTGTTTATATAGCAATGACCATCTCTAGTTGGTTCCTGGTTGTGTCGTGGATAATGGCCCCGTTTGTCTTCAATCCATCTGGCTTTG
ATTGGTTGAAGACAGTATACGACTTTGATGATTTTATGAACTGGATTTGGTACCAAGGTGGTGTGTTTGCAAAATCTGAACAGAGCTGGGAAAGATGGTG
GTATGAGGAGCAGGATCATCTGAGGACAACTGGGCTTTGGGGAAAGTTACTGGATGTAATATTGGACCTTCGCTTCTTCTTTTTTCAATATGGGATCGTA
TACCAACTAGGTATTGCTGCTGGGAGTACTAGCATTGCTGTTTACTTGCTTTCTTGGATTTATGTAGTTGTCGCCTTTGGGTTTTTTTTGATGGTAGCAT
ATGCCCGGAACAAGTATGCTGCAAAAGAACACATCTACTATCGGATGGTCCAGTTTCTGATCATTGTGCTTGGCATCTTTGTGATTATAGCCCTGCTTCA
GTTCACATCTTTCAAATTTACTGATGTTTTCACAAGTTTGTTGGCTTTTATCCCCACTGGATGGGGCATTTTATTGATTGCCCAAGTACTTCGCCCCTTC
CTGCCCGCTATACTTTGGGAAGCAGTGGTTTCTGTGGCTCGGCTATATGATATATTGTTTGGGGTGATAGTTATGGTCCCTGTGGCATTTTTGTCATGGA
TGCCTGGGTTCCAATCAATGCAGACTAGGATCCTCTTCAACGAGGCATTCAGCAGGGGCCTCCGGATCTTCCAGCTTTTCACGGGAAAAAAATCGTAG
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.013G131300.2 pacid=42810965 polypeptide=Potri.013G131350.1.p locus=Potri.013G131300 ID=Potri.013G131300.2.v4.1 annot-version=v4.1
MSLRRPPPPNPARTESQPYNIIPIQNLLADHPSLRYPEVRAAAAALRTVGNLRKPPYAQWHPSMDLLDWLALLFGFQKDNVRNQREHLVLHLANAQMRLT
PPPDNIDTLDAGVLRRFRRKLLKNYTNWCDYLNKKSNIWISDRSTDLRRELLYVSLYLLIWGESANLRFMPECICFIFHNMCFELNRVLEDYIDENTGQP
VMPSISGENAFLNGVVKPIYETVRREVDRSFNGAAPHSAWRNYDDLNEYFWSKRCFERLKWPIDLGSNFFVTSGSRKKVGKTGFVEQRSFWNIVRSFDRL
WVMLILFLQAGIIVAWEEKEYPWKALKSRDVQVRVLTVFFTWSGLRFLQSLLDVGTQYNLVSRETLGLGVRMILKSVVAVVWIIVFGAFYGRIWSQRNSD
LRRSPRDLSWSSEADKKVVTFLEVALVFVAPEILALALFILPWIRNFLENTDWRIFRMMTWWFQSSSFIGRGLREGLVDNIKYTLFWAMVLATKFAFSYF
MQIKPMVKPSKQMLKLKDVNYEWHEFFDHSNRFSVGLLWLPVVLIYLMDLQIWYAIYSSFVGAGVGLFQHLGEIRNIQQLRLRFQFFASAIQFNLMPEEQ
LLNARGTFKSKFKDAIHRLKLRYGFGHPYKKLESNQVEANKFALIWNEIIIIFREEDIISDKELELMELPQNSWNVRVIRWPSFLLCNELLLALSQAKEL
VDAPDKWLWYKICKNEYRRCAVIEAYDSVKHLLLEIIKTNTEEHSIITVLFQEIDHSLQIEKFTKTFKMTGLPNFHAKLIKLLELLNKPKRDLNQVVDTL
QALYEIAVREFFREKRSTEQLMEDGLAPRDPAAMAGLLFGNAVQLPDASNETFYRQARRLHMILTSRDSMNTIPENLEARRRIAFFSNSLFMSMPHAPQV
EKMMAFSVLTPYYNEEVLYSREQLRTENEDGVSTLYYLQTIYADEWKNFMQRMRREGMEKDGEIWTTKLRDLRLWASYRGQTLGRTVRGMMYYYRALKML
AFLDSASEMDIKEGSRELGSMRRDNGLDSFDSESSSKSLSRNSSSVNLLFKGHEYGTALMKYTYVVACQIYGAQKAKKDPHAEEILYLMKNNEALRVAYV
DEVNTGRDEMEYYSVLVKYDQQLDKEVEIYRVKLPGPLKLGEGKPENQNHALIFTRGDAVQTIDMNQDNYFEEALKMRNLLEEYRHYYGARKPTILGVRE
HIFTGSVSSLAWFMSAQETSFVTLGQRVLANPLKIRMHYGHPDVFDRFWFMTRGGISKASRVINISEDIFAGFNCTLRGGNITHHEYIQVGKGRDVGLNQ
ISMFEAKVASGNGEQTLSRDVYRLGHRLDFFRMLSFFYTTVGFFLNTMMVILTVYAFLWGRLYLALSGVEGSALADNSSNNKALGAILNQQFIIQLGLFT
ALPMIVENSLEHGFLEAIWDFLTMQLQLSSVFYTFSMGTRTHYFGRTILHGGAKYRATGRGFVVQHKSFAENYRLYARSHFVKAIELGLILVVYAAYSPV
AKDTFVYIAMTISSWFLVVSWIMAPFVFNPSGFDWLKTVYDFDDFMNWIWYQGGVFAKSEQSWERWWYEEQDHLRTTGLWGKLLDVILDLRFFFFQYGIV
YQLGIAAGSTSIAVYLLSWIYVVVAFGFFLMVAYARNKYAAKEHIYYRMVQFLIIVLGIFVIIALLQFTSFKFTDVFTSLLAFIPTGWGILLIAQVLRPF
LPAILWEAVVSVARLYDILFGVIVMVPVAFLSWMPGFQSMQTRILFNEAFSRGLRIFQLFTGKKS
|
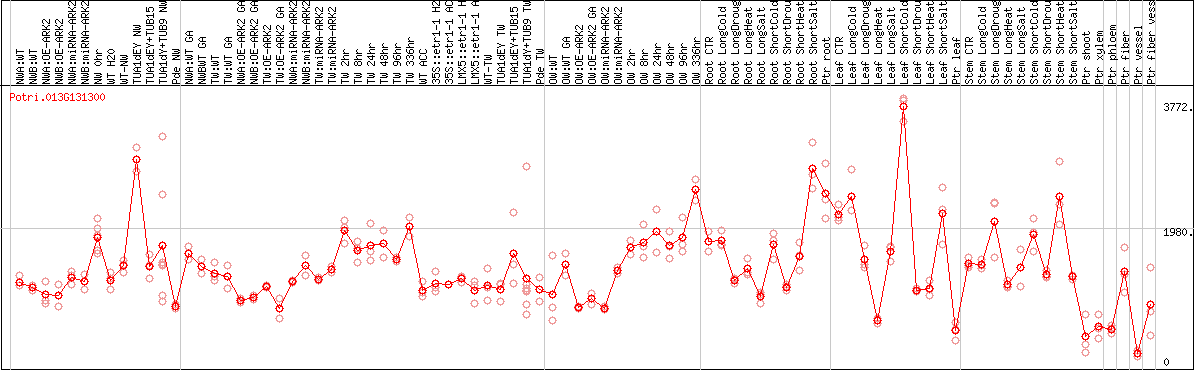
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.013G131300 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
