External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G03420 414 / 4e-146
Protein of unknown function (DUF789) (.1)
AT1G03610 409 / 3e-144
Protein of unknown function (DUF789) (.1)
AT4G28150 329 / 3e-113
Protein of unknown function (DUF789) (.1), Protein of unknown function (DUF789) (.2)
AT1G73210 269 / 4e-89
Protein of unknown function (DUF789) (.1), Protein of unknown function (DUF789) (.2)
AT1G17830 258 / 1e-84
Protein of unknown function (DUF789) (.1)
AT2G01260 219 / 8e-69
Protein of unknown function (DUF789) (.1), Protein of unknown function (DUF789) (.2), Protein of unknown function (DUF789) (.3)
AT1G15030 215 / 2e-67
Protein of unknown function (DUF789) (.1)
AT4G16100 209 / 5e-65
Protein of unknown function (DUF789) (.1)
AT5G49220 191 / 1e-57
Protein of unknown function (DUF789) (.1)
AT5G23380 150 / 4e-43
Protein of unknown function (DUF789) (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.019G103000
531 / 0
AT4G03420 427 / 2e-151
Protein of unknown function (DUF789) (.1)
Potri.017G155200
299 / 2e-100
AT1G17830 328 / 9e-112
Protein of unknown function (DUF789) (.1)
Potri.018G152500
294 / 7e-99
AT1G17830 321 / 3e-109
Protein of unknown function (DUF789) (.1)
Potri.010G000700
225 / 8e-71
AT5G49220 342 / 3e-115
Protein of unknown function (DUF789) (.1)
Potri.008G213300
218 / 3e-68
AT5G49220 338 / 2e-113
Protein of unknown function (DUF789) (.1)
Potri.010G120700
215 / 4e-67
AT2G01260 367 / 1e-125
Protein of unknown function (DUF789) (.1), Protein of unknown function (DUF789) (.2), Protein of unknown function (DUF789) (.3)
Potri.008G124000
211 / 1e-65
AT2G01260 384 / 1e-132
Protein of unknown function (DUF789) (.1), Protein of unknown function (DUF789) (.2), Protein of unknown function (DUF789) (.3)
Potri.004G068700
101 / 2e-23
AT1G73210 100 / 2e-22
Protein of unknown function (DUF789) (.1), Protein of unknown function (DUF789) (.2)
Potri.017G152601
93 / 1e-20
AT1G03610 95 / 9e-21
Protein of unknown function (DUF789) (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10018572
435 / 4e-154
AT4G03420 405 / 1e-142
Protein of unknown function (DUF789) (.1)
Lus10033707
406 / 8e-143
AT4G03420 359 / 2e-124
Protein of unknown function (DUF789) (.1)
Lus10031632
404 / 1e-142
AT4G03420 345 / 2e-119
Protein of unknown function (DUF789) (.1)
Lus10039803
303 / 3e-103
AT1G03610 303 / 2e-103
Protein of unknown function (DUF789) (.1)
Lus10038327
281 / 2e-93
AT1G73210 319 / 1e-108
Protein of unknown function (DUF789) (.1), Protein of unknown function (DUF789) (.2)
Lus10036189
251 / 3e-81
AT1G73210 298 / 2e-99
Protein of unknown function (DUF789) (.1), Protein of unknown function (DUF789) (.2)
Lus10016841
209 / 2e-64
AT4G16100 310 / 2e-102
Protein of unknown function (DUF789) (.1)
Lus10037716
206 / 2e-63
AT4G16100 311 / 4e-103
Protein of unknown function (DUF789) (.1)
Lus10037717
193 / 4e-56
AT4G16100 321 / 2e-103
Protein of unknown function (DUF789) (.1)
Lus10016842
162 / 6e-47
AT4G16100 272 / 8e-89
Protein of unknown function (DUF789) (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF05623
DUF789
Protein of unknown function (DUF789)
Representative CDS sequence
>Potri.013G144000.1 pacid=42810908 polypeptide=Potri.013G144000.1.p locus=Potri.013G144000 ID=Potri.013G144000.1.v4.1 annot-version=v4.1
ATGTTTCTTGAAAAAGGGTCAATGCAATCAAACCTTGATTGCTTCCTTCATTGCACAACCCCACGAGTCCAATCTCAATTCCTCGCGAAGAGTGAGACTA
GAAATCTAAACAGATTATGGCACCCATGGGAGAGAGACACAGTGGAGTATTTTACTTTGGGTGATCTATGGAATTTTTATGATGAATGGAGTGCTTATGG
GGCTGGAATTCCTATTGTTTTGAACAATGATGAAACTTTGGTTCAGTACTATGTGCCTTATCTTTCTGCAATCCAAATTTTCACAAGCAATTCTTCAGTC
AACAGTTTCAGGGAAGACACCGAGTCCGGTGATGGCGAAACGAGGGACTCCTTTAGCGATTCGTGGAGTAATGAGAGTGTGACTGACAAGGTATGTAGGT
CGGATGGATGCTCATCAGAGGAAGGAGGGTCAGAGCAAGACAATCCTTGGCCTATAAATGATAGATTGGGGCGACTATATTTTGAGTACTTTGAGAGATC
AACTCCTTATGGAAGAGTTCCTTTGATGGATAAGATTAATGGATTTGCTCGACGATTTCCGGGTTTAATGTCATTGAGAAGTGTAGATCTTTCACCAGCT
AGTTGGATGGCTGTTGCTTGGTATCCAATCTATCACATTCCAATGGGAAGAACCATAAAGGACTTGTCCACATGTTTCTTGACCTACCACACCCTCTCAT
CATCTTTTCAAGATATGGATGTAGAAGATGACATTGGGAGCCCAGAAAAGAAGCGAAAGGAAGGGGAAATCATCACTCTCCCTCCATTTGGTTTGGCCAC
TTACAAGATGCAAGGGAAGTTGTGGGTCTCAGGCAATGGCGGTAGGGACCAAGAGAGGCTAGTCTCACTTTTAAGTGTGGCAGATTCATGGCTAAAGCAA
CTAACGGTCCAGCACCATGACTTCGATTACTTCACGGGGATTAAGGGTGGCTAA
AA sequence
>Potri.013G144000.1 pacid=42810908 polypeptide=Potri.013G144000.1.p locus=Potri.013G144000 ID=Potri.013G144000.1.v4.1 annot-version=v4.1
MFLEKGSMQSNLDCFLHCTTPRVQSQFLAKSETRNLNRLWHPWERDTVEYFTLGDLWNFYDEWSAYGAGIPIVLNNDETLVQYYVPYLSAIQIFTSNSSV
NSFREDTESGDGETRDSFSDSWSNESVTDKVCRSDGCSSEEGGSEQDNPWPINDRLGRLYFEYFERSTPYGRVPLMDKINGFARRFPGLMSLRSVDLSPA
SWMAVAWYPIYHIPMGRTIKDLSTCFLTYHTLSSSFQDMDVEDDIGSPEKKRKEGEIITLPPFGLATYKMQGKLWVSGNGGRDQERLVSLLSVADSWLKQ
LTVQHHDFDYFTGIKGG
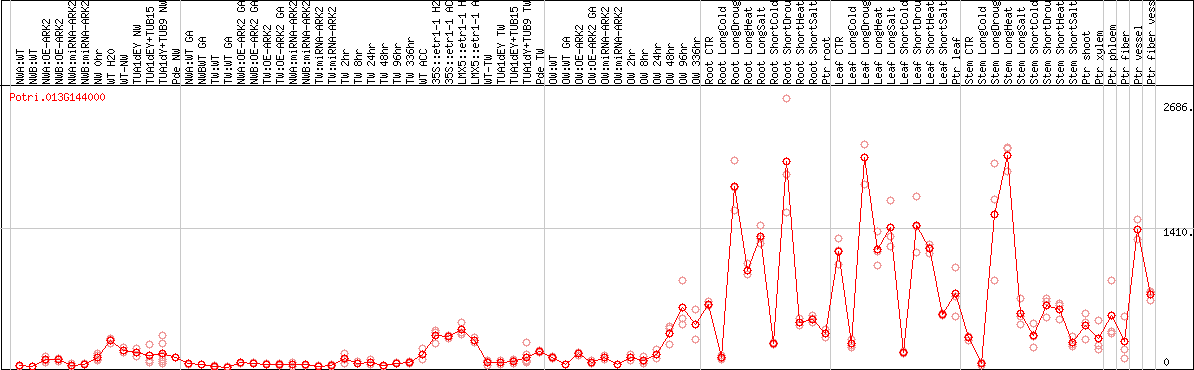
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.013G144000 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.