External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G11360 209 / 1e-67
Adenine nucleotide alpha hydrolases-like superfamily protein (.1.2.3.4)
AT5G54430 183 / 1e-57
ATPHOS32
Adenine nucleotide alpha hydrolases-like superfamily protein (.1)
AT4G27320 176 / 2e-54
ATPHOS34
Adenine nucleotide alpha hydrolases-like superfamily protein (.1)
AT3G21210 123 / 7e-32
zinc ion binding (.1)
AT3G17020 57 / 6e-10
Adenine nucleotide alpha hydrolases-like superfamily protein (.1)
AT3G53990 55 / 3e-09
Adenine nucleotide alpha hydrolases-like superfamily protein (.1.2)
AT3G03270 51 / 7e-08
Adenine nucleotide alpha hydrolases-like superfamily protein (.1.2)
AT1G09740 43 / 6e-05
Adenine nucleotide alpha hydrolases-like superfamily protein (.1)
AT4G31230 44 / 8e-05
Protein kinase protein with adenine nucleotide alpha hydrolases-like domain (.1)
AT2G24370 44 / 0.0001
Protein kinase protein with adenine nucleotide alpha hydrolases-like domain (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.019G119400
321 / 4e-111
AT1G11360 164 / 2e-49
Adenine nucleotide alpha hydrolases-like superfamily protein (.1.2.3.4)
Potri.011G039800
206 / 2e-66
AT1G11360 269 / 3e-91
Adenine nucleotide alpha hydrolases-like superfamily protein (.1.2.3.4)
Potri.011G125500
199 / 2e-63
AT5G54430 237 / 3e-78
Adenine nucleotide alpha hydrolases-like superfamily protein (.1)
Potri.001G409100
188 / 2e-59
AT5G54430 226 / 2e-74
Adenine nucleotide alpha hydrolases-like superfamily protein (.1)
Potri.010G144100
58 / 3e-10
AT3G17020 234 / 3e-80
Adenine nucleotide alpha hydrolases-like superfamily protein (.1)
Potri.013G009800
54 / 1e-08
AT3G62550 172 / 2e-55
Adenine nucleotide alpha hydrolases-like superfamily protein (.1)
Potri.006G092700
53 / 1e-08
AT3G53990 207 / 1e-69
Adenine nucleotide alpha hydrolases-like superfamily protein (.1.2)
Potri.014G122000
50 / 1e-07
AT3G62550 195 / 1e-64
Adenine nucleotide alpha hydrolases-like superfamily protein (.1)
Potri.017G144301
45 / 8e-06
AT3G03270 248 / 9e-86
Adenine nucleotide alpha hydrolases-like superfamily protein (.1.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10031594
224 / 3e-73
AT1G11360 257 / 2e-86
Adenine nucleotide alpha hydrolases-like superfamily protein (.1.2.3.4)
Lus10039730
224 / 1e-72
AT1G11360 288 / 3e-98
Adenine nucleotide alpha hydrolases-like superfamily protein (.1.2.3.4)
Lus10033749
220 / 2e-71
AT1G11360 245 / 1e-81
Adenine nucleotide alpha hydrolases-like superfamily protein (.1.2.3.4)
Lus10037867
178 / 2e-53
AT1G11360 244 / 1e-78
Adenine nucleotide alpha hydrolases-like superfamily protein (.1.2.3.4)
Lus10038561
152 / 2e-45
AT1G11360 224 / 7e-74
Adenine nucleotide alpha hydrolases-like superfamily protein (.1.2.3.4)
Lus10016900
67 / 2e-12
AT3G17020 229 / 1e-72
Adenine nucleotide alpha hydrolases-like superfamily protein (.1)
Lus10018515
62 / 3e-12
AT1G11360 127 / 9e-38
Adenine nucleotide alpha hydrolases-like superfamily protein (.1.2.3.4)
Lus10037761
61 / 3e-11
AT3G17020 224 / 4e-76
Adenine nucleotide alpha hydrolases-like superfamily protein (.1)
Lus10022602
51 / 9e-08
AT3G53990 225 / 2e-76
Adenine nucleotide alpha hydrolases-like superfamily protein (.1.2)
Lus10017207
51 / 1e-07
AT3G53990 217 / 2e-73
Adenine nucleotide alpha hydrolases-like superfamily protein (.1.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0039
HUP
PF00582
Usp
Universal stress protein family
Representative CDS sequence
>Potri.013G150200.1 pacid=42812605 polypeptide=Potri.013G150200.1.p locus=Potri.013G150200 ID=Potri.013G150200.1.v4.1 annot-version=v4.1
ATGAACTCAAAACAACCTTACAAATCTGAATCTGACCTCCCAGTCCCACCACTCCCCACCCTTCCCTGCCTCCACTCCCCTACCACCCCGAATTCCACCT
CCAACCGCCGTGTCGCCATAGCCGTGGACCTTTCCGACGAGAGTGCATATGCAGTCAAATGGGCAGTCGAGAACTACCTCCGACCAGGCGACGCCGTGAT
CCTCCTCCACGTCCGGCCAACTTCCGTCCTCTATGGCGCTGATTGGGGGTCAATTCAACTCCAAATCAATAACAACAACACGCCTTTTGAGCTAAGTGGC
TCAAACAGTCCTGATAATCGAGAAAGGCAGAAACTTGAGGATGATTTTGATAGTTTTACTAATAACAAAACCAATTTGTTGGCTAAACCTTTGTTGGAGG
CTAATGTTCCTTTCAAGATTCATGTAGTGAAAGATCATGACATGAAGGAGAGGCTCTGTTTGGAAGTTGAGAGATTGGGGTTAAGTGCAGTGATAATGGG
AAGTAGAGGTTTTGGTGCTACAAGAAAGAAGGGGATTAGTAAAGGAAGAAGTGTTGGTGGTGGGAGGTTAGGGAGTGTTAGTGATCATTGTGTGCAGCAT
TGTGTTTGCCCTGTTGTTGTTGTTAGGTGTAGTGATGATGGTAAAGAAGAAGAGAGTGTGAAAACAGGTGGTGTTGGTGACGGGGTTGAAGAGGGTCTCC
ATCCTGTGCCTGAGGAGGATCAAGAGGAGTGTGTTGATGATGAGCTGAAAGATGCTTAG
AA sequence
>Potri.013G150200.1 pacid=42812605 polypeptide=Potri.013G150200.1.p locus=Potri.013G150200 ID=Potri.013G150200.1.v4.1 annot-version=v4.1
MNSKQPYKSESDLPVPPLPTLPCLHSPTTPNSTSNRRVAIAVDLSDESAYAVKWAVENYLRPGDAVILLHVRPTSVLYGADWGSIQLQINNNNTPFELSG
SNSPDNRERQKLEDDFDSFTNNKTNLLAKPLLEANVPFKIHVVKDHDMKERLCLEVERLGLSAVIMGSRGFGATRKKGISKGRSVGGGRLGSVSDHCVQH
CVCPVVVVRCSDDGKEEESVKTGGVGDGVEEGLHPVPEEDQEECVDDELKDA
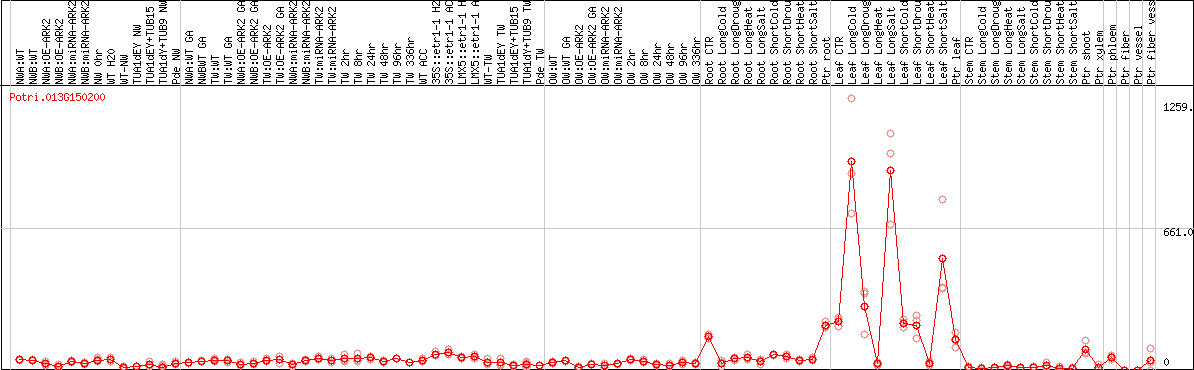
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.013G150200 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.