External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G20020 305 / 1e-103
ATPRMT6
ARABIDOPSIS THALIANA PROTEIN ARGININE METHYLTRANSFERASE 6, protein arginine methyltransferase 6 (.1.2)
AT2G19670 91 / 2e-21
ATPRMT1A
ARABIDOPSIS THALIANA PROTEIN ARGININE METHYLTRANSFERASE 1A, protein arginine methyltransferase 1A (.1)
AT4G29510 82 / 3e-18
ATPRMT1B, ATPRMT11
PROTEIN ARGININE METHYLTRANSFERASE 1B, ARABIDOPSIS THALIANA PROTEIN ARGININE METHYLTRANSFERASE 1B, ARABIDOPSIS THALIANA ARGININE METHYLTRANSFERASE 11, arginine methyltransferase 11 (.1)
AT3G12270 79 / 4e-17
ATPRMT3
ARABIDOPSIS THALIANA PROTEIN ARGININE METHYLTRANSFERASE 3, protein arginine methyltransferase 3 (.1)
AT1G04870 69 / 8e-14
PRMT10, ATPRMT10
protein arginine methyltransferase 10 (.1.2)
AT3G06930 63 / 1e-11
ATPRMT4B
ARABIDOPSIS THALIANA PROTEIN ARGININE METHYLTRANSFERASE 4B, protein arginine methyltransferase 4B (.1.2)
AT5G49020 60 / 2e-10
ATPRMT4A
ARABIDOPSIS THALIANA PROTEIN ARGININE METHYLTRANSFERASE 4A, protein arginine methyltransferase 4A (.1.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.007G000300
385 / 2e-135
AT3G20020 582 / 0.0
ARABIDOPSIS THALIANA PROTEIN ARGININE METHYLTRANSFERASE 6, protein arginine methyltransferase 6 (.1.2)
Potri.006G149400
82 / 2e-18
AT4G29510 597 / 0.0
PROTEIN ARGININE METHYLTRANSFERASE 1B, ARABIDOPSIS THALIANA PROTEIN ARGININE METHYLTRANSFERASE 1B, ARABIDOPSIS THALIANA ARGININE METHYLTRANSFERASE 11, arginine methyltransferase 11 (.1)
Potri.018G067200
78 / 7e-17
AT4G29510 597 / 0.0
PROTEIN ARGININE METHYLTRANSFERASE 1B, ARABIDOPSIS THALIANA PROTEIN ARGININE METHYLTRANSFERASE 1B, ARABIDOPSIS THALIANA ARGININE METHYLTRANSFERASE 11, arginine methyltransferase 11 (.1)
Potri.001G030200
71 / 4e-14
AT3G12270 674 / 0.0
ARABIDOPSIS THALIANA PROTEIN ARGININE METHYLTRANSFERASE 3, protein arginine methyltransferase 3 (.1)
Potri.018G071200
62 / 3e-11
AT1G04870 551 / 0.0
protein arginine methyltransferase 10 (.1.2)
Potri.010G014600
58 / 8e-10
AT5G49020 764 / 0.0
ARABIDOPSIS THALIANA PROTEIN ARGININE METHYLTRANSFERASE 4A, protein arginine methyltransferase 4A (.1.2)
Potri.008G222700
53 / 3e-08
AT5G49020 762 / 0.0
ARABIDOPSIS THALIANA PROTEIN ARGININE METHYLTRANSFERASE 4A, protein arginine methyltransferase 4A (.1.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10010162
335 / 5e-116
AT3G20020 614 / 0.0
ARABIDOPSIS THALIANA PROTEIN ARGININE METHYLTRANSFERASE 6, protein arginine methyltransferase 6 (.1.2)
Lus10017367
333 / 3e-114
AT3G20020 636 / 0.0
ARABIDOPSIS THALIANA PROTEIN ARGININE METHYLTRANSFERASE 6, protein arginine methyltransferase 6 (.1.2)
Lus10008577
87 / 1e-19
AT4G29510 553 / 0.0
PROTEIN ARGININE METHYLTRANSFERASE 1B, ARABIDOPSIS THALIANA PROTEIN ARGININE METHYLTRANSFERASE 1B, ARABIDOPSIS THALIANA ARGININE METHYLTRANSFERASE 11, arginine methyltransferase 11 (.1)
Lus10040546
78 / 1e-16
AT4G29510 617 / 0.0
PROTEIN ARGININE METHYLTRANSFERASE 1B, ARABIDOPSIS THALIANA PROTEIN ARGININE METHYLTRANSFERASE 1B, ARABIDOPSIS THALIANA ARGININE METHYLTRANSFERASE 11, arginine methyltransferase 11 (.1)
Lus10000982
78 / 1e-16
AT4G29510 619 / 0.0
PROTEIN ARGININE METHYLTRANSFERASE 1B, ARABIDOPSIS THALIANA PROTEIN ARGININE METHYLTRANSFERASE 1B, ARABIDOPSIS THALIANA ARGININE METHYLTRANSFERASE 11, arginine methyltransferase 11 (.1)
Lus10026158
77 / 2e-16
AT3G12270 613 / 0.0
ARABIDOPSIS THALIANA PROTEIN ARGININE METHYLTRANSFERASE 3, protein arginine methyltransferase 3 (.1)
Lus10008664
77 / 3e-16
AT3G12270 610 / 0.0
ARABIDOPSIS THALIANA PROTEIN ARGININE METHYLTRANSFERASE 3, protein arginine methyltransferase 3 (.1)
Lus10043412
63 / 2e-11
AT5G49020 770 / 0.0
ARABIDOPSIS THALIANA PROTEIN ARGININE METHYLTRANSFERASE 4A, protein arginine methyltransferase 4A (.1.2)
Lus10034172
63 / 2e-11
AT5G49020 743 / 0.0
ARABIDOPSIS THALIANA PROTEIN ARGININE METHYLTRANSFERASE 4A, protein arginine methyltransferase 4A (.1.2)
Lus10019439
55 / 1e-08
AT1G04870 551 / 0.0
protein arginine methyltransferase 10 (.1.2)
PFAM info
Representative CDS sequence
>Potri.014G004000.2 pacid=42763431 polypeptide=Potri.014G004000.2.p locus=Potri.014G004000 ID=Potri.014G004000.2.v4.1 annot-version=v4.1
ATGCTGGGAAGTGTCATCACTGCCAGAGATCACTGGCTGAAGCATGGAGGCCTTATTCTTCCATCAAATGCAACTTTGTACATGGCACCTGTCACACATC
CAGACAGATATGGTGAAAGCATTGATTTTTGGCAAAACGTTAATGGAATCAATATGTCTGCCATGTTGCCATTAGCAAAACAGTGTGCATTTGAAGAGCC
ATCTGTGGAATCAATATCAGGTGAAAATGTGTTGACATGGCCACATGTGGTTAAGCATGAGGATTTCTATACAATTGCAATTGATGAACTAGAATCTGTT
ACCACGAGATACAAATTCAGGTCAATGATGAGAGCTCCATTGCATGGCTTTGCGTTTTGGTTCAATGTTGAATTTGGTTGGCCTGCAGCATCTCCGATCA
TTCCTCAAGCATCAGTCTTGCCAACTGCGCCATCAAGCAATCCTTCAATGGACGGTAGTAAGAGGAAGAAAAGAACAAATCAAAATGAGGCACTTGTGCT
ATCCACTGCACCCGAGGACCCTCCAACACATTGGCAAAATACTTTGATATACTCTTACGACCCAATAGATGTGGAGCAAGATCAGCTCATTGAAGGATCA
GCTACAATAACACAAAGCAAAGAAAACCTTTGA
AA sequence
>Potri.014G004000.2 pacid=42763431 polypeptide=Potri.014G004000.2.p locus=Potri.014G004000 ID=Potri.014G004000.2.v4.1 annot-version=v4.1
MLGSVITARDHWLKHGGLILPSNATLYMAPVTHPDRYGESIDFWQNVNGINMSAMLPLAKQCAFEEPSVESISGENVLTWPHVVKHEDFYTIAIDELESV
TTRYKFRSMMRAPLHGFAFWFNVEFGWPAASPIIPQASVLPTAPSSNPSMDGSKRKKRTNQNEALVLSTAPEDPPTHWQNTLIYSYDPIDVEQDQLIEGS
ATITQSKENL
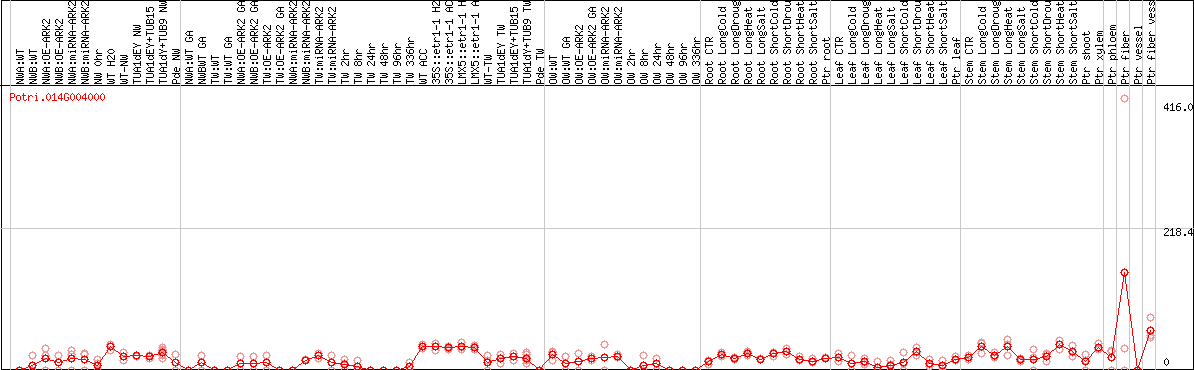
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.014G004000 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.