External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G57870 251 / 5e-87
SCE1A, SCE1, AHUS5, EMB1637, ATSCE1
SUMO CONJUGATING ENZYME 1A, EMBRYO DEFECTIVE 1637, sumo conjugation enzyme 1 (.1)
AT2G02760 96 / 1e-25
ATUBC2
ubiquitin-conjugating enzyme 2, ubiquiting-conjugating enzyme 2 (.1)
AT1G14400 94 / 8e-25
ATUBC1, UBC1
UBIQUITIN CONJUGATING ENZYME 1, ubiquitin carrier protein 1 (.1.2)
AT5G62540 92 / 2e-24
UBC3
ubiquitin-conjugating enzyme 3 (.1)
AT3G08690 79 / 3e-19
ATUBC11, UBC11
ubiquitin-conjugating enzyme 11 (.1.2)
AT1G64230 77 / 2e-18
UBC28
ubiquitin-conjugating enzyme 28 (.1.2.3.4.5)
AT4G27960 77 / 2e-18
UBC9
ubiquitin conjugating enzyme 9 (.1.2)
AT5G53300 77 / 3e-18
UBC10
ubiquitin-conjugating enzyme 10 (.1.2.3.4)
AT5G41700 76 / 4e-18
ATUBC8, UBC8
ARABIDOPSIS THALIANA UBIQUITIN CONJUGATING ENZYME 8, ubiquitin conjugating enzyme 8 (.1.2.3.4.5)
AT2G16740 72 / 1e-16
UBC29
ubiquitin-conjugating enzyme 29 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.014G024950
292 / 4e-103
AT3G57870 291 / 6e-103
SUMO CONJUGATING ENZYME 1A, EMBRYO DEFECTIVE 1637, sumo conjugation enzyme 1 (.1)
Potri.002G123600
281 / 1e-98
AT3G57870 283 / 1e-99
SUMO CONJUGATING ENZYME 1A, EMBRYO DEFECTIVE 1637, sumo conjugation enzyme 1 (.1)
Potri.005G141100
247 / 3e-85
AT3G57870 303 / 1e-107
SUMO CONJUGATING ENZYME 1A, EMBRYO DEFECTIVE 1637, sumo conjugation enzyme 1 (.1)
Potri.015G064000
100 / 4e-27
AT2G02760 284 / 3e-100
ubiquitin-conjugating enzyme 2, ubiquiting-conjugating enzyme 2 (.1)
Potri.013G064400
97 / 6e-26
AT2G02760 313 / 1e-111
ubiquitin-conjugating enzyme 2, ubiquiting-conjugating enzyme 2 (.1)
Potri.019G039200
96 / 1e-25
AT2G02760 285 / 2e-100
ubiquitin-conjugating enzyme 2, ubiquiting-conjugating enzyme 2 (.1)
Potri.001G094900
80 / 2e-19
AT1G64230 302 / 2e-107
ubiquitin-conjugating enzyme 28 (.1.2.3.4.5)
Potri.012G033000
79 / 3e-19
AT5G53300 302 / 1e-107
ubiquitin-conjugating enzyme 10 (.1.2.3.4)
Potri.015G023300
79 / 3e-19
AT5G53300 302 / 1e-107
ubiquitin-conjugating enzyme 10 (.1.2.3.4)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10019328
254 / 4e-88
AT3G57870 308 / 3e-109
SUMO CONJUGATING ENZYME 1A, EMBRYO DEFECTIVE 1637, sumo conjugation enzyme 1 (.1)
Lus10009376
254 / 6e-88
AT3G57870 306 / 7e-109
SUMO CONJUGATING ENZYME 1A, EMBRYO DEFECTIVE 1637, sumo conjugation enzyme 1 (.1)
Lus10019327
252 / 3e-87
AT3G57870 308 / 2e-109
SUMO CONJUGATING ENZYME 1A, EMBRYO DEFECTIVE 1637, sumo conjugation enzyme 1 (.1)
Lus10009377
251 / 1e-86
AT3G57870 306 / 1e-108
SUMO CONJUGATING ENZYME 1A, EMBRYO DEFECTIVE 1637, sumo conjugation enzyme 1 (.1)
Lus10037203
95 / 3e-25
AT2G02760 310 / 1e-110
ubiquitin-conjugating enzyme 2, ubiquiting-conjugating enzyme 2 (.1)
Lus10036727
95 / 3e-25
AT2G02760 310 / 1e-110
ubiquitin-conjugating enzyme 2, ubiquiting-conjugating enzyme 2 (.1)
Lus10030786
92 / 3e-24
AT2G02760 305 / 2e-108
ubiquitin-conjugating enzyme 2, ubiquiting-conjugating enzyme 2 (.1)
Lus10013263
91 / 2e-23
AT2G02760 304 / 6e-108
ubiquitin-conjugating enzyme 2, ubiquiting-conjugating enzyme 2 (.1)
Lus10039323
80 / 1e-19
AT1G64230 303 / 8e-108
ubiquitin-conjugating enzyme 28 (.1.2.3.4.5)
Lus10027570
80 / 1e-19
AT1G64230 303 / 8e-108
ubiquitin-conjugating enzyme 28 (.1.2.3.4.5)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0208
UBC
PF00179
UQ_con
Ubiquitin-conjugating enzyme
Representative CDS sequence
>Potri.014G024801.1 pacid=42764417 polypeptide=Potri.014G024801.1.p locus=Potri.014G024801 ID=Potri.014G024801.1.v4.1 annot-version=v4.1
ATGTCAGGAGGAGGCATAGCTCGTGGTCGTCTTGCTGAAGAGAGAAAGTCATGGCGTAAGAATCATCCCCACGGTTTTGTGGCTAAGCCTGATAATGCAC
AAGATGGTTCTCTTGATTTGATGGTGTGGAAGTGCATCATACCTGGCAAACCCGGGACAGATTGGGAGGGTGGCTTCTTCCCCCTCACGCTTCATTTTAG
CGAGGACTACCCAAGCAAACCTCCAAAGTGCAAGTTCCCCCCGGGTTTCTTCCATCCTAATGTCTACCCTTCTGGAACTGTGTGCTTATCTATCCTCAAT
GAGGACTATGGCTGGAGACCAGCCATAACTGTGAAGCAAATTTTAGTTGGCATTCAGGATTTGCTTGATCAACCAAATCCTGCTGATCCTGCGCAAACTG
ATGGCTACCAGCTTTTTGTCCAGGACCCGACTGAGTACAGGAGAAAGGTGCGCCAACAAGCCAAGCAATATCCACCTGCGCTCTGA
AA sequence
>Potri.014G024801.1 pacid=42764417 polypeptide=Potri.014G024801.1.p locus=Potri.014G024801 ID=Potri.014G024801.1.v4.1 annot-version=v4.1
MSGGGIARGRLAEERKSWRKNHPHGFVAKPDNAQDGSLDLMVWKCIIPGKPGTDWEGGFFPLTLHFSEDYPSKPPKCKFPPGFFHPNVYPSGTVCLSILN
EDYGWRPAITVKQILVGIQDLLDQPNPADPAQTDGYQLFVQDPTEYRRKVRQQAKQYPPAL
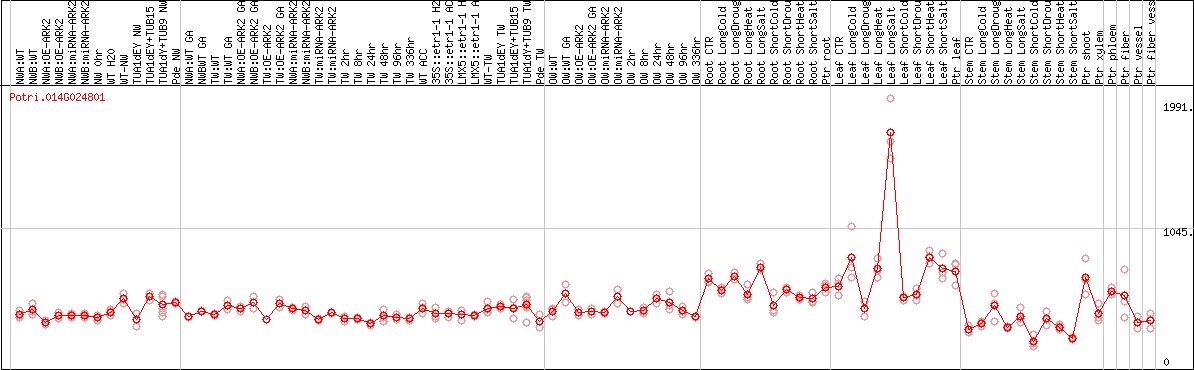
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.014G024801 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.