External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G47510 418 / 4e-147
AT5PTASE11, 5PTASE11
ARABIDOPSIS THALIANA INOSITOL POLYPHOSPHATE 5-PHOSPHATASE 11, inositol polyphosphate 5-phosphatase 11 (.1.2)
AT2G43900 115 / 3e-28
Endonuclease/exonuclease/phosphatase family protein (.1)
AT1G65580 108 / 6e-26
FRA3
FRAGILE FIBER3, Endonuclease/exonuclease/phosphatase family protein (.1)
AT2G31830 107 / 2e-25
endonuclease/exonuclease/phosphatase family protein (.1.2)
AT1G71710 105 / 5e-25
DNAse I-like superfamily protein (.1.2)
AT1G34120 102 / 8e-24
AT5PTASE1, ATIP5PI, AT5P1, IP5PI
MYO-INOSITOL POLYPHOSPHATE 5-PHOSPHATASE 1, inositol polyphosphate 5-phosphatase I (.1.2.3)
AT1G05630 100 / 8e-23
AT5PTASE13, 5PTASE13
Endonuclease/exonuclease/phosphatase family protein (.1.2)
AT4G18010 97 / 6e-22
AT5PTASE2, IP5PII
INOSITOL\(1,4,5\)P3 5-PHOSPHATASE II, myo-inositol polyphosphate 5-phosphatase 2 (.1.2)
AT5G65090 96 / 7e-22
DER4, MRH3, BST1
DEFORMED ROOT HAIRS 4, BRISTLED 1, DNAse I-like superfamily protein (.1.2)
AT2G37440 95 / 2e-21
DNAse I-like superfamily protein (.1.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.007G145100
113 / 2e-27
AT2G43900 1511 / 0.0
Endonuclease/exonuclease/phosphatase family protein (.1)
Potri.008G079300
111 / 6e-27
AT1G65580 1475 / 0.0
FRAGILE FIBER3, Endonuclease/exonuclease/phosphatase family protein (.1)
Potri.005G197800
103 / 4e-24
AT1G71710 658 / 0.0
DNAse I-like superfamily protein (.1.2)
Potri.003G088300
96 / 2e-21
AT4G18010 615 / 0.0
INOSITOL\(1,4,5\)P3 5-PHOSPHATASE II, myo-inositol polyphosphate 5-phosphatase 2 (.1.2)
Potri.001G145900
96 / 2e-21
AT4G18010 615 / 0.0
INOSITOL\(1,4,5\)P3 5-PHOSPHATASE II, myo-inositol polyphosphate 5-phosphatase 2 (.1.2)
Potri.005G078000
94 / 4e-21
AT5G65090 666 / 0.0
DEFORMED ROOT HAIRS 4, BRISTLED 1, DNAse I-like superfamily protein (.1.2)
Potri.007G090200
93 / 1e-20
AT5G65090 671 / 0.0
DEFORMED ROOT HAIRS 4, BRISTLED 1, DNAse I-like superfamily protein (.1.2)
Potri.010G177400
93 / 1e-20
AT1G65580 1490 / 0.0
FRAGILE FIBER3, Endonuclease/exonuclease/phosphatase family protein (.1)
Potri.017G074200
92 / 2e-20
AT2G37440 328 / 8e-108
DNAse I-like superfamily protein (.1.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10008330
109 / 4e-26
AT2G43900 1477 / 0.0
Endonuclease/exonuclease/phosphatase family protein (.1)
Lus10040107
104 / 2e-24
AT4G18010 638 / 0.0
INOSITOL\(1,4,5\)P3 5-PHOSPHATASE II, myo-inositol polyphosphate 5-phosphatase 2 (.1.2)
Lus10003010
103 / 4e-24
AT4G18010 655 / 0.0
INOSITOL\(1,4,5\)P3 5-PHOSPHATASE II, myo-inositol polyphosphate 5-phosphatase 2 (.1.2)
Lus10011041
102 / 6e-24
AT4G18010 655 / 0.0
INOSITOL\(1,4,5\)P3 5-PHOSPHATASE II, myo-inositol polyphosphate 5-phosphatase 2 (.1.2)
Lus10008934
98 / 1e-23
AT5G04980 395 / 3e-138
DNAse I-like superfamily protein (.1.2)
Lus10028884
98 / 2e-22
AT5G04980 526 / 0.0
DNAse I-like superfamily protein (.1.2)
Lus10028727
98 / 3e-22
AT1G71710 674 / 0.0
DNAse I-like superfamily protein (.1.2)
Lus10027073
97 / 6e-22
AT2G43900 1455 / 0.0
Endonuclease/exonuclease/phosphatase family protein (.1)
Lus10034857
97 / 6e-22
AT5G04980 530 / 0.0
DNAse I-like superfamily protein (.1.2)
Lus10033398
96 / 8e-22
AT5G04980 540 / 0.0
DNAse I-like superfamily protein (.1.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0530
DNase_I-like
PF03372
Exo_endo_phos
Endonuclease/Exonuclease/phosphatase family
Representative CDS sequence
>Potri.014G039800.1 pacid=42763192 polypeptide=Potri.014G039800.1.p locus=Potri.014G039800 ID=Potri.014G039800.1.v4.1 annot-version=v4.1
ATGGGAAATTGGAACACAACTTGTGCTTGTAAGAGATCTAAGTTGATGAGGAAGCAAATGGATTCCATGAATCATAATACTCATCATGAAGGAATAAGGA
CTGTAAAGGTGAACAAGGTCTGCGACTTCTCTGGAAACTCAGACCTTTGCATTTGCGCAGTGACCTGGAACATGAATGGAAAGGTGGTTTATGAAGACTT
GGTGGAGCTGGTTGGGAGCAACTCCAAGTCTGATCTACTGGTGGTAGGTTTGCAAGAGGTGCCTCGGAAAAACATCGCACGGTTATTGGAAACAGCACTT
GTGGATACTCATGTCCTGCTAGGAAAGGCAATCATGCAATCTCTTCAACTTTATGTATTTGGACCTCAGAACTCGGGGTTTTCCATCAAAGAATTGAAGG
TGGATAAGCATTCTGTTGGTGGTTGTGGAGGATTAATTAGGAGAAAGAAAGGAGCTGTGGCCATCTGCATGAGCTACAATGGCATCCGAATGGTATTCAT
CTCTTGCCATCTTTCTGCTCATGCTCGTAACGTGGAAGAGAGGAATTCACAATGCAGACATATATCACACAACCTTTTCTCTAAGTATATGAATCCTTAC
AGCAAGCCTGCTCAAGTCACGGTATGGCTGGGAGATCTTAACTACAGAATACAAGGGATAGAAACACATCCAGTCAGAAATCTTATACAGAAAAACCTTC
ACAGATTCCTCACCAGCAAAGATCAACTTCTACAAGAAGCTGAAAGAGGACAGGTGTTTGATGGATACTGCGAAGGGACATTGACCTTTAAACCAACATA
CAAGTACGATGTTGGAAGCAGCAATTACGATACAAGTTACAAGGTGCGAGTCCCATCATGGACAGATCGAATCTTGTTCAAGATTCAAGACATGGAGGAA
ATCAGGGCAAGTTTGCATTCTTACGAGTCAGTTGATGACATTCACAGTTCGGATCACAAGCCAGTGAAAGCTCACCTCTGCTTGAAAGTTAATGAACAGC
CATAA
AA sequence
>Potri.014G039800.1 pacid=42763192 polypeptide=Potri.014G039800.1.p locus=Potri.014G039800 ID=Potri.014G039800.1.v4.1 annot-version=v4.1
MGNWNTTCACKRSKLMRKQMDSMNHNTHHEGIRTVKVNKVCDFSGNSDLCICAVTWNMNGKVVYEDLVELVGSNSKSDLLVVGLQEVPRKNIARLLETAL
VDTHVLLGKAIMQSLQLYVFGPQNSGFSIKELKVDKHSVGGCGGLIRRKKGAVAICMSYNGIRMVFISCHLSAHARNVEERNSQCRHISHNLFSKYMNPY
SKPAQVTVWLGDLNYRIQGIETHPVRNLIQKNLHRFLTSKDQLLQEAERGQVFDGYCEGTLTFKPTYKYDVGSSNYDTSYKVRVPSWTDRILFKIQDMEE
IRASLHSYESVDDIHSSDHKPVKAHLCLKVNEQP
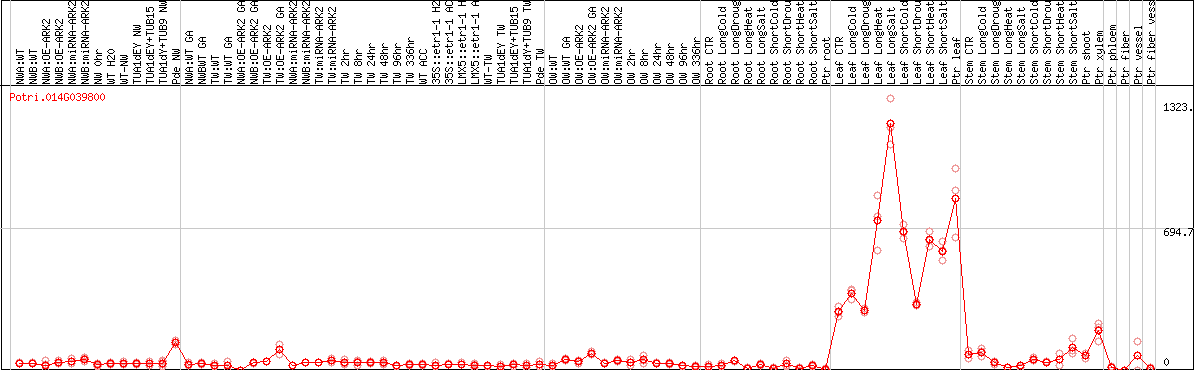
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.014G039800 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.