External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G26667 358 / 2e-127
PYR6
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1.2.3)
AT3G60180 307 / 9e-108
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1.2)
AT4G25280 242 / 2e-81
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
AT3G60961 204 / 5e-68
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
AT5G35170 103 / 2e-25
adenylate kinase family protein (.1.2)
AT5G47840 100 / 3e-25
AMK2
adenosine monophosphate kinase (.1)
AT2G37250 85 / 8e-20
ADK, ATPADK1
adenosine kinase (.1)
AT5G63400 83 / 3e-19
ADK1
adenylate kinase 1 (.1.2)
AT2G39270 81 / 3e-18
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
AT5G50370 78 / 2e-17
Adenylate kinase family protein (.1)
Paralogs
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10031611
369 / 1e-131
AT5G26667 343 / 1e-121
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1.2.3)
Lus10033733
357 / 5e-126
AT5G26667 336 / 2e-117
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1.2.3)
Lus10031714
244 / 1e-76
AT4G25270 672 / 0.0
organelle transcript processing 70, Tetratricopeptide repeat (TPR)-like superfamily protein (.1)
Lus10019509
226 / 3e-76
AT5G26667 217 / 5e-73
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1.2.3)
Lus10031135
224 / 2e-74
AT4G25280 263 / 3e-89
P-loop containing nucleoside triphosphate hydrolases superfamily protein (.1)
Lus10037995
96 / 9e-25
AT5G47840 315 / 9e-110
adenosine monophosphate kinase (.1)
Lus10009228
97 / 3e-24
AT5G47840 296 / 3e-101
adenosine monophosphate kinase (.1)
Lus10003504
95 / 2e-23
AT5G47840 300 / 1e-102
adenosine monophosphate kinase (.1)
Lus10009478
90 / 1e-22
AT5G47840 293 / 1e-101
adenosine monophosphate kinase (.1)
Lus10040358
91 / 8e-22
AT2G37250 405 / 1e-143
adenosine kinase (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0023
P-loop_NTPase
PF13238
AAA_18
AAA domain
Representative CDS sequence
>Potri.014G043300.7 pacid=42763463 polypeptide=Potri.014G043300.7.p locus=Potri.014G043300 ID=Potri.014G043300.7.v4.1 annot-version=v4.1
ATGGGGACTGCCGGTGATTCTGAGAAGAAGCCTACTGTTGTTTTTGTTTTGGGTGGCCCTGGAAGTGGAAAGGGTACCCAGTGTGCCAATATTGTCGAAC
ACTTTGGTTACACTCATCTTAGTGCTGGAGATCTTCTTCGAGCAGAAATTAAATCTGGTTCTGAAAATGGAACCATGATTCAGAACATGATTAAGGAGGG
CAAGATAGTGCCTTCAGAGGTAACAATAAAGCTCCTCCAAAAAGCAATGCAGGATAGTGGAAACGACAAATTTCTGATTGATGGCTTTCCCCGCAATGAG
GAAAACCGAGCTGCTTTTGAAGCTGTTACCAAAATTGAGCCGGCATTTGTCCTTTTCTTTGATTGTCCTGAAGAGGAGATGGAGAGGCGTATTTTGAGCA
GGAACCAGGGAAGAGAAGATGACAACATTGAGACAATAAGGAAGCGGTTTAAAGTTTTTCTAGAGTCTAGCCTTCCCGTGGTTGAGTATTATGAATCCAA
GGGGAAAGTTCAAAAGGTTGATGCTGCAAAGCCTATTGATGAGGTTTTTGAGGTAGTTAAAGCCATCTTTACCCCAAAAGACGAGAAGGTAAAGCAGCAT
TGCTGTGCTATCTTGTAG
AA sequence
>Potri.014G043300.7 pacid=42763463 polypeptide=Potri.014G043300.7.p locus=Potri.014G043300 ID=Potri.014G043300.7.v4.1 annot-version=v4.1
MGTAGDSEKKPTVVFVLGGPGSGKGTQCANIVEHFGYTHLSAGDLLRAEIKSGSENGTMIQNMIKEGKIVPSEVTIKLLQKAMQDSGNDKFLIDGFPRNE
ENRAAFEAVTKIEPAFVLFFDCPEEEMERRILSRNQGREDDNIETIRKRFKVFLESSLPVVEYYESKGKVQKVDAAKPIDEVFEVVKAIFTPKDEKVKQH
CCAIL
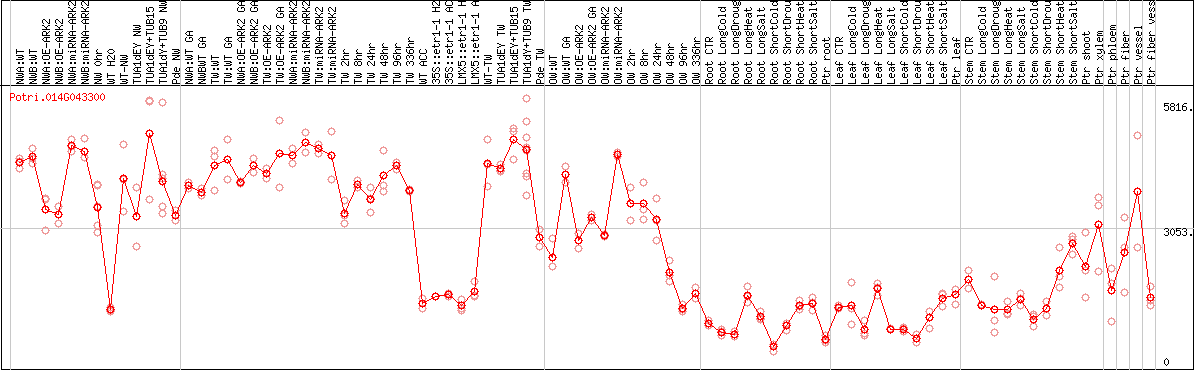
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.014G043300 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.