External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G60410 105 / 5e-26
Protein of unknown function (DUF1639) (.1), Protein of unknown function (DUF1639) (.2), Protein of unknown function (DUF1639) (.3)
AT4G17440 55 / 1e-08
Protein of unknown function (DUF1639) (.1), Protein of unknown function (DUF1639) (.2)
AT3G18295 53 / 5e-08
Protein of unknown function (DUF1639) (.1)
AT1G25370 53 / 8e-08
Protein of unknown function (DUF1639) (.1)
AT1G48770 43 / 7e-05
Protein of unknown function (DUF1639) (.1)
AT4G20300 43 / 0.0002
Protein of unknown function (DUF1639) (.1), Protein of unknown function (DUF1639) (.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.002G136200
238 / 1e-77
AT3G60410 109 / 6e-28
Protein of unknown function (DUF1639) (.1), Protein of unknown function (DUF1639) (.2), Protein of unknown function (DUF1639) (.3)
Potri.012G055200
63 / 1e-11
AT3G18295 109 / 3e-29
Protein of unknown function (DUF1639) (.1)
Potri.010G122700
63 / 3e-11
AT1G25370 110 / 1e-28
Protein of unknown function (DUF1639) (.1)
Potri.008G122600
60 / 4e-10
AT1G25370 100 / 6e-25
Protein of unknown function (DUF1639) (.1)
Potri.001G073400
44 / 6e-05
AT4G20300 226 / 3e-71
Protein of unknown function (DUF1639) (.1), Protein of unknown function (DUF1639) (.2)
Potri.003G157500
44 / 6e-05
AT4G20300 201 / 2e-61
Protein of unknown function (DUF1639) (.1), Protein of unknown function (DUF1639) (.2)
Potri.014G044700
42 / 0.0001
ND /
Potri.006G233100
42 / 0.0004
AT1G55340 130 / 9e-37
Protein of unknown function (DUF1639) (.1), Protein of unknown function (DUF1639) (.2)
Potri.018G059200
41 / 0.0008
AT1G55340 133 / 3e-38
Protein of unknown function (DUF1639) (.1), Protein of unknown function (DUF1639) (.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10010984
61 / 2e-10
AT3G60410 89 / 3e-20
Protein of unknown function (DUF1639) (.1), Protein of unknown function (DUF1639) (.2), Protein of unknown function (DUF1639) (.3)
Lus10004378
58 / 3e-09
AT4G17440 96 / 4e-23
Protein of unknown function (DUF1639) (.1), Protein of unknown function (DUF1639) (.2)
Lus10038942
50 / 3e-08
AT1G25370 72 / 7e-17
Protein of unknown function (DUF1639) (.1)
Lus10027232
49 / 4e-07
AT1G68340 75 / 3e-17
Protein of unknown function (DUF1639) (.1)
Lus10035144
47 / 4e-06
AT1G25370 87 / 2e-20
Protein of unknown function (DUF1639) (.1)
Lus10040176
41 / 0.0006
AT3G60410 76 / 1e-16
Protein of unknown function (DUF1639) (.1), Protein of unknown function (DUF1639) (.2), Protein of unknown function (DUF1639) (.3)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF07797
DUF1639
Protein of unknown function (DUF1639)
Representative CDS sequence
>Potri.014G044800.1 pacid=42764101 polypeptide=Potri.014G044800.1.p locus=Potri.014G044800 ID=Potri.014G044800.1.v4.1 annot-version=v4.1
ATGGTGTTTCGCACTGAACCGGAAACTGAGCCAAACACAAACAACAACACTATCATGGCTTCTTCTACTACTACTACTAGTACTGTGAAACCACACCAGC
CACTCCATAACTTCCCTTTGCAAGACCTAAAATGGTCAATGAACCATTCCAATAACGTTACCAACCACCACCACCGCTTTCGCACTCACAAATCTCCTCA
CCGCGATGCTGCTTCTGCGGCGGATTCGGAGGGAGACGGCGGGGTCAAGATTGATAAACTATTGAAGAAAAAATCTGGTGATGTTGAGAATTCGGAGAGG
AAATCGAAGATCTTTATTCGGTTGAGGACTAACAAAAACAGCAGCGGAAGCGGAAGCAGCAGCGGAAGCAGCAAGGGCGTGGTTGGTGATGTTGCTGCGG
GTGCAGTGGATCAGGGCTCTGCTGCTGTAGTGGAGGACGTGGAGGAACTGATGCCGAAGACGTGGAATCTGAGGCCGAGGAGAGCTGTTAATAATATTAA
TAATGATAGTAATAATAATAATAACAAGGGTTTGAATGGGAATGGAGGTGCTTTGAAGATTTGTGGGGGTGCGGTGCCGGAGATCAAACCTCAGGTGCCG
GGTGGTAACCGGACTGAATTGACTCGGTCCAACCGGAATGGTAATGACGCTAATAATTATGATAATGATAATGACAATAATAATAATAACAGGAAGGAGA
GGGGGAAGGAAAAGGAAAAGGAGAAGAAGGTGAGGTTTTCGATTCCTCTTACGAAAGTGGAAATTGAAGAGGATATTTATTCCTTGACTGGGTCAAAGCC
GGCGAGAAGGCCTAAGAAGAGAGCAAAGCATGTGCAGAAACAACTTGATTGCCTGTTTCCTGGTATGTGGCTGGATTCGATCACCCCGGATTGCTATAAG
GTTCATGAAGCTCCTTCGAAGTTGAGAATGTAG
AA sequence
>Potri.014G044800.1 pacid=42764101 polypeptide=Potri.014G044800.1.p locus=Potri.014G044800 ID=Potri.014G044800.1.v4.1 annot-version=v4.1
MVFRTEPETEPNTNNNTIMASSTTTTSTVKPHQPLHNFPLQDLKWSMNHSNNVTNHHHRFRTHKSPHRDAASAADSEGDGGVKIDKLLKKKSGDVENSER
KSKIFIRLRTNKNSSGSGSSSGSSKGVVGDVAAGAVDQGSAAVVEDVEELMPKTWNLRPRRAVNNINNDSNNNNNKGLNGNGGALKICGGAVPEIKPQVP
GGNRTELTRSNRNGNDANNYDNDNDNNNNNRKERGKEKEKEKKVRFSIPLTKVEIEEDIYSLTGSKPARRPKKRAKHVQKQLDCLFPGMWLDSITPDCYK
VHEAPSKLRM
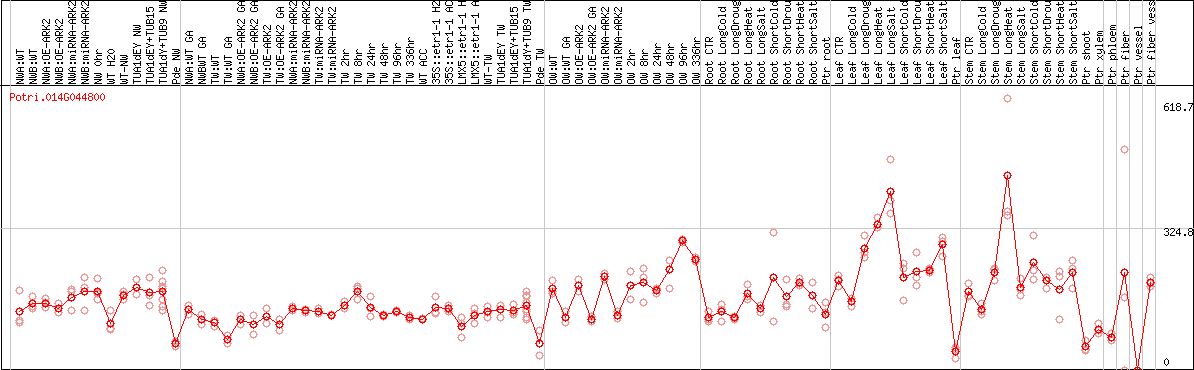
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.014G044800 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.