Potri.014G052600 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.014G052600.2 pacid=42762363 polypeptide=Potri.014G052600.2.p locus=Potri.014G052600 ID=Potri.014G052600.2.v4.1 annot-version=v4.1
ATGTCCTACAATCAATCAAGGAGTGGCAGCGATAAGAGCGAGTTACAATATAGGAAATCAGGGCGATCCATCAGCTCTAATCAGCTACGGACTTCCTCTC
AATCCTACGGTAAGGGCGGTGGTGGCGGCGGCGGCGGCCCCCCAGTCCCTTCACCGTCTTCTTCTTCTCTTTCCTCCAATCGCAGCAGTTTCAATAAGAA
GTCTAGTTATGTTCCACAAGGAGGAGGGCAATCTAGTAGGGTAAATGTGGCACCTGTTGTGAATTCTTCCGATTCCGGTAATAATGCTGCTTCTACAATC
CGTAATGTACAGAATGGTGCCGCGGCGCAACCCCCATTACATGGAACATCTGATGCTCCGCCGCCTGCAAGCAGTGTTACCAAGCCGACAGAAACATCAG
CTACTCAGAGGAGTGCCCGAGCTGTTCCAAAGGCTCCAACTTCTCAACCTGCCACCATAAGTTCGGAGAGTGGAGCACCCACTACACCTGCCAAGGCCCC
AGTAGATGCATCTAAGGCGTTTGCGTTTCAATTTGGGTCCATTAGTCCTGGCTTTATGAACGGGATGCAGGTTCCTGCTCGAACAAGCTCAGCTCCCCCA
AATTTGGATGAGCAGAAACGTGACCAGGCACGCCAAGATACATTTAGGCCTGCTCCTTCATTACCGACTCCTGCTCCGAAGCAGCAATTTCAGAAGAAGG
AAGTGAGCGCAACAGAACAAACTATCTCTGGGGGGGTTCATCCGCTTCCCAAGGCCAAAAAGGAGACACAAGTCTCACCTGCACCATCTGCGAGCCACTC
ACAGAAGCATTCTGTTCTTCCGGTTACTATGACTTCTATGCAAATGCAATATCTGCAGCCACAGGTCTCTGTGCAGTTTGGTGGCCGAGGCCCACAAATT
CAGTCCCAGGGTGTCCCACCAACTTCTCTTCAGATGCCAATTCCAGTGCCATTGCAGATGGGCAGTGCTCCTCAAGTGCAACAGCCAGTGTTTATTCAAG
GTATCCAACATCATCCTATGCAGCCTCAGGGAATGATGCGTCAAGGGCAAAACTTGAGTTTTACGACCACAATGGGTCCTCAGATGCCTCCTCAGTTAGG
CAGCTTAGGAATGAACATTGCCTCACAATATTCTCAACAACAGGGTGGTAAATTTGGTGGTCAGCGTAAAACATCTGTCAAGATTACTGATCCAAAGACA
CATGAAGAGCTGAGGCTTGATAAACGGACAGATCCATATCCAGACACTGGACCATCAGGTCTAAGGTCTCATCTCAATGCTCCCCAGTCCCAGCCAATTC
CATCATTTACACCTTCACGTCCAATTAACTATTATCCTAGTTCTTACAATACAAATAATTTATTTTTTCAAACTCCAAGCTCTCTACCATTAACTGGTGG
CCAAATAGCACCAAATTCTCAGCCACCACCTAGGTTTAATTATCCAGTTAGCCAGGGGCCCCAAAATGTTCCATATACCAATGCATCTGCTCTTAATTCT
CTTCCAGCTAGTAAGTCTGGGATTGCAATTCACGGTGTTGCCGAACTGCATAAATCAGAACATGCCAGTGATGCACCTAATGCAATCTCCTCAACTCCAT
CTGGAGTAGTACAAGTGACCATTAAACCACCTGTTGGTTCTATTGGAGAGAAAGTTGTGGAGCCATCTTTGCCAAAGATCTCTCCTGTCGAAAAGGGCGG
ATCCCACAAATCCTCAAGGTCATCTGGGGAAGCTAGCCCATCTCCTTCTCAAAGAGATTCAGAGACTTCATCAGAAAGCTCTTTACGGCAGGCAAAACCT
GTTGGTGAATCGCTGGTCAAGTCACCTCCAGTGGCAGCAAAACAGCTTGCAGAGGTTGCTGTTGATGGTGCAGCATCTACTTTGCCAGCTCAGTCTGTGG
AGGCAATACCAGGTGTATCTAATGCTGAAGACCAAAAAAAGGAAGCTCCGAGTATTCAGAAGAAACCAGGCAAGAAAGGAAATATTGAGCCTCAGCATCA
GATTGGTGGACAGACTACCTTGTCCACAAGCTTGTCTTCTCGCACTGTGGAACTTGGTGTATTTTATGGCAGTGGAGTTTCTGAAACTGCAGAAACTAAT
ACAGCTCCTTCCCCATCACCAGCAAACAGTGAAGCTTTGACAAAATCAATCAAGGAACCAGTGTCAACAATTTCTGCTTTGAATCCTGATGTTTCTGAAA
TGAAGGTTGAGAATGCTGGAGATGGTTTTAACACTGTTTCAGCTCTGGGTCTTGTTGCTGGAGTTGCTAAAACCCCACACACCACTCCACAAGCTATGCT
GGATGGTTCCTCCTCACAGGAAGAACTACAATGTGAAATTCCAACCGCAGAAGAGAAAGGACAGAAATCACTGTCTGAATGTCTTAAACAAGATTACAGC
ATGTCTCCAGCACCAGTTAATTCAAAATTTGCAGACATTGTCAAGCAAGATAAAGAGGTATCTGATTTGACAGGGACATCTGTTGGCAATGAGGTCCCAG
CCTCAGAAACTGGACAGGAGGGTCTGGTGGAACCTGTGACTCGCCACGCAGCAAATGACAGGGTATCAGATAGTGTGGATGTCTCTGCATCCAGAAATTT
GGATTCTGCCGATGATAGAAAGCCTTCAGATGCATCTTTGAGGCACGGTGATGGCATAGGTAATAAAGAAGCCTCTGTTACAAAATCAAGTGTATCAGGT
CAGCAGGAATCTCTTCCAGTACCTGATCTTTCTGAGGCAACTGCAAAACACAAAGGGCAATGTGCGGAAAACCCCGGCAGTGGAACAGTCCCCCATGCAA
TATCTAGTTCCAAAGAAAAACCCACTGAGCCGACTTTATCAAAGAGTACCTCTGGTAAATTTAAGAAGAAGAGAAGAGAATTTCTTCTGAAAGCAGATCT
TGCTGGGACTACTTCTGATCTTTATGGGGCATATAAAGGCCCTGAGGAAAAGAAAGAAAATGTCATATCTTCAGAAGTCACAGAAAGCACCAGCCCTATT
CTGAATCAGACACCTGCTGATGCCCTGCAGGTTGACTCTGTAGCAAGCGAGAAAAATAAAGCCGAGCCAGATGATTGGGAAGATGCTGCTGACATGTCTA
CACCGAAACTGGATAGTGATGGGGAACTATCATGTGGAGGATTGGGACAACATGATTCAGATGGGAATGCAAATACAGCCAAAAAATATTCCAGAGATTT
CCTCCTTAAATTTTCTGAGCAATTTTCTAATCTTCCTGAAGGTTTTGTAATTACATCTGATATAGCAGAGGCTTTGAGTGTCAACGTTTCTCATCCTGCT
GATCTTGATTCTTACCCCAGTCCTGCAAGAGTTATGGACAGGTCTAATAGTGGATCTCGAATAGGACGTGGGAGTGGCATGGTTGATGATGGCCGATGGA
GTAAACAGCCTGGTCCATTTGGTCCGGGAAGGGATTTGCACTTGGATATGGGTTATGGACCTAATGCCAGTTTTCGACCTGTTGCAGGAGGCAACCATGG
TGTTCTGAGGAACCCGCGAGCACAGAGTCCTGGACAGTATGCAGGAGGGATTCTATCTGGACCCGTACAATCAACGGGTCTGCAGGGAGGGATGCAAAGA
GGTGGCTCAGATGCTGACAAATGGCAGCGTTCTGTTAGTTCTGTGTATAAAGGATTGATTCCTTCTCCTCATACTCCATTACAGACGATGCACAAAGCTG
AGAGAAAGTATGAAGTGGGTAAAGTGGCAGATGAAGAAGCAGCCAAGCAAAGACAGTTGAAAGGCATATTGAACAAGTTAACTCCTCAGAATTTCGAGAA
GCTTTTTGAGCAAGTAAAAGCTGTTAACATTGACAATGCTGTGACACTTAACGGCGTTATCTCACAGATCTTTGACAAAGCTTTAATGGAGCCCACATTC
TGTGAAATGTATGCTAATTTCTGCTTTCATCTAGCAGCAGAGTTGCCTGAACTCATTGAAGATGATGAAAAGGTGACTTTCAAGAGGTTACTTCTGAACA
AGTGCCAGGAGGAATTTGAGAGAGGGGAGCGAGAGCAAGAGGAAGCTAATAAAGCTGATGAAGAGGGTGAGATTAAAAAGTCTGATGAGGAAAGAGAAGA
ACAGAGAATCAAGGCACGGAGACGAATGCTTGGTAACATAAGACTAATTGGTGAGCTCTATAAAAAAAGAATGTTGACTGAAAGAATAATGCATGAGTGC
ATCAAAAAGTTGCTGGGCCAATATCAGAATCCTGACGAGGAGGATGTTGAGTCTCTTTGTAAACTAATGAGCACTATTGGAGAGATGATTGACCATCCCA
AAGCTAAGGTGCATATGGATGCATATTTTGACATGATGGCAAAATTATCAAACAACATGAAACTCTCATCTAGGGTTAGATTTATGTTGAAGGATGCAAT
TGATTTGAGGAAAAACAAATGGCAGCAGAGGAGGAAGGTTGAAGGGCCGAAAAAGATCGAGGAAGTGCACAGAGATGCTGCTCAAGAACGACAGCTGCAA
ACTAGTAGGCTGGCTCGCAATCCTGGCATGAATTCTTCTCCAAGAAGGGGCCCTATGGATTTTGGTCCGAGAGGGTCAACAATGTTATCTTCTCCAAATG
CTCATATGGGTGGTTTCCGAGGGTTTCCTTCTCAGGTTCGTGGGCATGGCAACCAGGATGTTCGGCATGAGGACAGACAATCTTACGAGGCTAGGACTGT
GTCTGTTCCCTTACCTCAAAGACCTCTTGGTGATGATTCTATTACTCTGGGTCCACAAGGTGGTCTTGCAAGAGGAATGTCCATTAGAGGAACACCTGCT
ATTACAGTTGCTCCTGTATCTGAGATTTCTCCAAGCCCAAGTGACTCGAGAAGAATGGCAGCTGGTTTGAATGGTGTTAGTGCCATTTTAGAACGATCAA
ATTATAGCCCAAGGGAAGATCTCATTCCAAGATATTCTCCTGACAGATTTGCAGTTCCACCTACTCATGATCAAATGAGTGGTCAGGAACGGAATATGAA
TTATGTTAACAGGGACCTGAGGAATCTAGATCATGGTTTTGATAGGCCGTTGGGATCATCATCACTAACAAATACACAAGGGCCATCTTTTGCCCAGAGT
ATTCCAACAGGAAAGATGTGGCCTGAAGAACAACTGCGAGAAATGTCCATGGTCACAATAAAAGAATTTTACAGTGCCAGAGATGAGAAAGAAGTTGCTC
TATGCATAAAAGATTTGAATTCCCCAAGTTTCCACCCTTCAATGATCTCTCTCTGGGTCACAGACTCATTTGAAAGGAAGGACATGGATAGAGATCTTTT
GGCTAAGCTTCTTGCCAGTCTTACAAGATCCCAGGATTGCATTTTGGATTCAAATCAGCTCGTCAAAGGGTTTGAATCAGTTCTGACTACCTTGGAGGAT
GCTGTAACTGATGCCCCCAAAGCACCAGAATTTCTTGGACGCATCTTGGGCAGAGTTGTTGTAGAAAATGTTGTTCCACTAAAAGAGATTGGGCGGTTAT
TGCATGAAGGCGGAGAGGAGCCGGGTAGCCTTCTTAAGTTTGGGCTTGCAGGGGACGTTCTTGGAAGTGTTTTGGAGATGATCAAAGCAGAGAACGGACA
AGGTGTCTTAAATGAGATTCGCAACGCCTCCAATTTGCGTTTCGAAGATTTCCGGCCTCCACATCCTAACAGATCAAGGATACTAGAGAAGTTTATTTAG
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.014G052600.2 pacid=42762363 polypeptide=Potri.014G052600.2.p locus=Potri.014G052600 ID=Potri.014G052600.2.v4.1 annot-version=v4.1
MSYNQSRSGSDKSELQYRKSGRSISSNQLRTSSQSYGKGGGGGGGGPPVPSPSSSSLSSNRSSFNKKSSYVPQGGGQSSRVNVAPVVNSSDSGNNAASTI
RNVQNGAAAQPPLHGTSDAPPPASSVTKPTETSATQRSARAVPKAPTSQPATISSESGAPTTPAKAPVDASKAFAFQFGSISPGFMNGMQVPARTSSAPP
NLDEQKRDQARQDTFRPAPSLPTPAPKQQFQKKEVSATEQTISGGVHPLPKAKKETQVSPAPSASHSQKHSVLPVTMTSMQMQYLQPQVSVQFGGRGPQI
QSQGVPPTSLQMPIPVPLQMGSAPQVQQPVFIQGIQHHPMQPQGMMRQGQNLSFTTTMGPQMPPQLGSLGMNIASQYSQQQGGKFGGQRKTSVKITDPKT
HEELRLDKRTDPYPDTGPSGLRSHLNAPQSQPIPSFTPSRPINYYPSSYNTNNLFFQTPSSLPLTGGQIAPNSQPPPRFNYPVSQGPQNVPYTNASALNS
LPASKSGIAIHGVAELHKSEHASDAPNAISSTPSGVVQVTIKPPVGSIGEKVVEPSLPKISPVEKGGSHKSSRSSGEASPSPSQRDSETSSESSLRQAKP
VGESLVKSPPVAAKQLAEVAVDGAASTLPAQSVEAIPGVSNAEDQKKEAPSIQKKPGKKGNIEPQHQIGGQTTLSTSLSSRTVELGVFYGSGVSETAETN
TAPSPSPANSEALTKSIKEPVSTISALNPDVSEMKVENAGDGFNTVSALGLVAGVAKTPHTTPQAMLDGSSSQEELQCEIPTAEEKGQKSLSECLKQDYS
MSPAPVNSKFADIVKQDKEVSDLTGTSVGNEVPASETGQEGLVEPVTRHAANDRVSDSVDVSASRNLDSADDRKPSDASLRHGDGIGNKEASVTKSSVSG
QQESLPVPDLSEATAKHKGQCAENPGSGTVPHAISSSKEKPTEPTLSKSTSGKFKKKRREFLLKADLAGTTSDLYGAYKGPEEKKENVISSEVTESTSPI
LNQTPADALQVDSVASEKNKAEPDDWEDAADMSTPKLDSDGELSCGGLGQHDSDGNANTAKKYSRDFLLKFSEQFSNLPEGFVITSDIAEALSVNVSHPA
DLDSYPSPARVMDRSNSGSRIGRGSGMVDDGRWSKQPGPFGPGRDLHLDMGYGPNASFRPVAGGNHGVLRNPRAQSPGQYAGGILSGPVQSTGLQGGMQR
GGSDADKWQRSVSSVYKGLIPSPHTPLQTMHKAERKYEVGKVADEEAAKQRQLKGILNKLTPQNFEKLFEQVKAVNIDNAVTLNGVISQIFDKALMEPTF
CEMYANFCFHLAAELPELIEDDEKVTFKRLLLNKCQEEFERGEREQEEANKADEEGEIKKSDEEREEQRIKARRRMLGNIRLIGELYKKRMLTERIMHEC
IKKLLGQYQNPDEEDVESLCKLMSTIGEMIDHPKAKVHMDAYFDMMAKLSNNMKLSSRVRFMLKDAIDLRKNKWQQRRKVEGPKKIEEVHRDAAQERQLQ
TSRLARNPGMNSSPRRGPMDFGPRGSTMLSSPNAHMGGFRGFPSQVRGHGNQDVRHEDRQSYEARTVSVPLPQRPLGDDSITLGPQGGLARGMSIRGTPA
ITVAPVSEISPSPSDSRRMAAGLNGVSAILERSNYSPREDLIPRYSPDRFAVPPTHDQMSGQERNMNYVNRDLRNLDHGFDRPLGSSSLTNTQGPSFAQS
IPTGKMWPEEQLREMSMVTIKEFYSARDEKEVALCIKDLNSPSFHPSMISLWVTDSFERKDMDRDLLAKLLASLTRSQDCILDSNQLVKGFESVLTTLED
AVTDAPKAPEFLGRILGRVVVENVVPLKEIGRLLHEGGEEPGSLLKFGLAGDVLGSVLEMIKAENGQGVLNEIRNASNLRFEDFRPPHPNRSRILEKFI
|
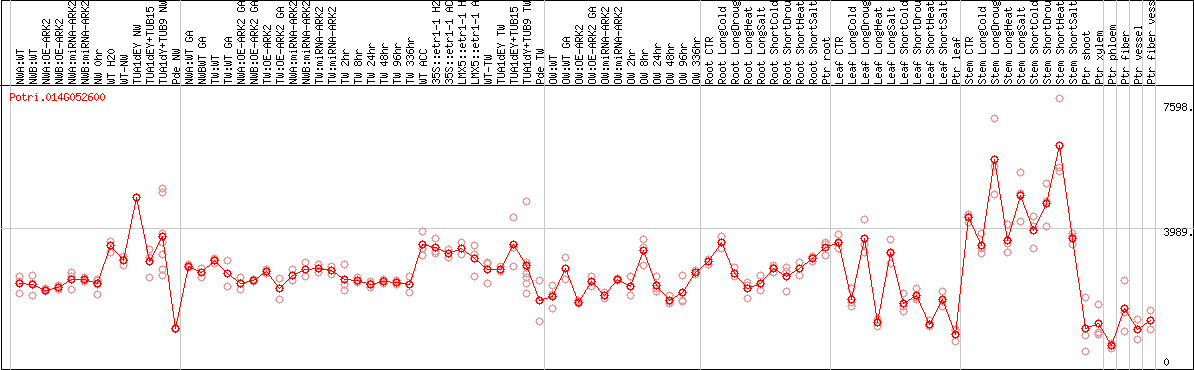
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.014G052600 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
