External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G45130 269 / 4e-91
ATSPX3
ARABIDOPSIS THALIANA SPX DOMAIN GENE 3, SPX domain gene 3 (.1)
AT5G20150 222 / 3e-72
ATSPX1
ARABIDOPSIS THALIANA SPX DOMAIN GENE 1, SPX domain gene 1 (.1)
AT5G15330 202 / 8e-64
ATSPX4
ARABIDOPSIS THALIANA SPX DOMAIN GENE 4, SPX domain gene 4 (.1)
AT2G26660 198 / 1e-62
ATSPX2
ARABIDOPSIS THALIANA SPX DOMAIN GENE 2, SPX domain gene 2 (.1)
AT4G11810 59 / 1e-09
Major Facilitator Superfamily with SPX (SYG1/Pho81/XPR1) domain-containing protein (.1)
AT4G22990 52 / 1e-07
Major Facilitator Superfamily with SPX (SYG1/Pho81/XPR1) domain-containing protein (.1), Major Facilitator Superfamily with SPX (SYG1/Pho81/XPR1) domain-containing protein (.2)
AT1G63010 45 / 4e-05
Major Facilitator Superfamily with SPX (SYG1/Pho81/XPR1) domain-containing protein
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.002G143900
431 / 6e-155
AT2G45130 276 / 1e-93
ARABIDOPSIS THALIANA SPX DOMAIN GENE 3, SPX domain gene 3 (.1)
Potri.018G131500
210 / 4e-67
AT5G20150 356 / 1e-124
ARABIDOPSIS THALIANA SPX DOMAIN GENE 1, SPX domain gene 1 (.1)
Potri.006G069500
206 / 2e-65
AT5G20150 359 / 8e-126
ARABIDOPSIS THALIANA SPX DOMAIN GENE 1, SPX domain gene 1 (.1)
Potri.018G028200
196 / 6e-62
AT2G26660 321 / 2e-110
ARABIDOPSIS THALIANA SPX DOMAIN GENE 2, SPX domain gene 2 (.1)
Potri.001G135950
189 / 3e-61
AT2G45130 121 / 7e-35
ARABIDOPSIS THALIANA SPX DOMAIN GENE 3, SPX domain gene 3 (.1)
Potri.017G086600
192 / 6e-60
AT5G15330 335 / 4e-115
ARABIDOPSIS THALIANA SPX DOMAIN GENE 4, SPX domain gene 4 (.1)
Potri.006G253400
184 / 5e-57
AT5G20150 308 / 1e-105
ARABIDOPSIS THALIANA SPX DOMAIN GENE 1, SPX domain gene 1 (.1)
Potri.001G193200
133 / 2e-38
AT5G20150 158 / 2e-48
ARABIDOPSIS THALIANA SPX DOMAIN GENE 1, SPX domain gene 1 (.1)
Potri.003G120600
56 / 1e-08
AT4G22990 1112 / 0.0
Major Facilitator Superfamily with SPX (SYG1/Pho81/XPR1) domain-containing protein (.1), Major Facilitator Superfamily with SPX (SYG1/Pho81/XPR1) domain-containing protein (.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10043272
213 / 2e-68
AT5G20150 343 / 8e-120
ARABIDOPSIS THALIANA SPX DOMAIN GENE 1, SPX domain gene 1 (.1)
Lus10030095
210 / 2e-67
AT5G20150 356 / 8e-125
ARABIDOPSIS THALIANA SPX DOMAIN GENE 1, SPX domain gene 1 (.1)
Lus10019415
201 / 9e-62
AT5G20150 326 / 2e-110
ARABIDOPSIS THALIANA SPX DOMAIN GENE 1, SPX domain gene 1 (.1)
Lus10002018
180 / 2e-55
AT2G26660 308 / 2e-105
ARABIDOPSIS THALIANA SPX DOMAIN GENE 2, SPX domain gene 2 (.1)
Lus10002908
176 / 1e-53
AT2G26660 309 / 1e-105
ARABIDOPSIS THALIANA SPX DOMAIN GENE 2, SPX domain gene 2 (.1)
Lus10009314
61 / 2e-10
AT4G22990 941 / 0.0
Major Facilitator Superfamily with SPX (SYG1/Pho81/XPR1) domain-containing protein (.1), Major Facilitator Superfamily with SPX (SYG1/Pho81/XPR1) domain-containing protein (.2)
Lus10015852
53 / 1e-07
AT4G22990 926 / 0.0
Major Facilitator Superfamily with SPX (SYG1/Pho81/XPR1) domain-containing protein (.1), Major Facilitator Superfamily with SPX (SYG1/Pho81/XPR1) domain-containing protein (.2)
Lus10006340
53 / 1e-07
AT4G22990 1128 / 0.0
Major Facilitator Superfamily with SPX (SYG1/Pho81/XPR1) domain-containing protein (.1), Major Facilitator Superfamily with SPX (SYG1/Pho81/XPR1) domain-containing protein (.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF03105
SPX
SPX domain
Representative CDS sequence
>Potri.014G061400.1 pacid=42763726 polypeptide=Potri.014G061400.1.p locus=Potri.014G061400 ID=Potri.014G061400.1.v4.1 annot-version=v4.1
ATGAAGTTTGGCAAGAGATTGAAGCAACAAGTTCAAGAAACCTTGCCAGATTGGCGTGACAAGTTTTTGTCTTACAAGGAATTGAAGAAGCTTGTTCGTC
TGATCTCATCGGCTCCACCTTTCTCGTATGGATCAGTTGAGTATGGAAAGGCAGAGGCTGAGTTTGTGCGTCTGCTGAACAGTGAAATTGACAAGTTTAA
TACTTTTTTCATGGAACAAGAAGAGGATTTCATTATTCGGCATGAGGAGCTAAAGCAGAGGATTCAGAAAGTGATTGATACTTGGGGGCCAAGTGGAAGT
CAGCCTTCAGAGGCAGAGTACAAAGAGCAGATGAGAAAGATTAGAAAAAACAGTGTCAATTTCCACGGTGAAATGGTGCTTCTGGAGAATTACAGCAACA
TCAATTACACAGGATTGGCCAAGATATTGAAGAAGTATGACAAGAGAACAGGTGGGTTATTGCGCCTTCCATTCATTCAAAAGGTGTTGGAGCAACCCTT
TTTTATTACCGATCTAGTATCAAAGCTTGTCAAGCAATGCGAATACATGATAGATACAGTATTCCCTGTAGAGGAAGAAGAGAGAGTGAAAGAAGGAAGA
GAAGCAATAACGGTTGCGGGAGAAGGGATTTTCAGGAACACAATTGCAGCTCTAATGACCATGCAAGAAATTAGAAGAGGCAGCTCTACTTACAGCCATT
TCTCTCTGCCTCCTCTGAACTTGCCAGATTCTGATCTTATTCAGTCATTCCAGCTCAATTCTCCTATATCTATTGTCTAA
AA sequence
>Potri.014G061400.1 pacid=42763726 polypeptide=Potri.014G061400.1.p locus=Potri.014G061400 ID=Potri.014G061400.1.v4.1 annot-version=v4.1
MKFGKRLKQQVQETLPDWRDKFLSYKELKKLVRLISSAPPFSYGSVEYGKAEAEFVRLLNSEIDKFNTFFMEQEEDFIIRHEELKQRIQKVIDTWGPSGS
QPSEAEYKEQMRKIRKNSVNFHGEMVLLENYSNINYTGLAKILKKYDKRTGGLLRLPFIQKVLEQPFFITDLVSKLVKQCEYMIDTVFPVEEEERVKEGR
EAITVAGEGIFRNTIAALMTMQEIRRGSSTYSHFSLPPLNLPDSDLIQSFQLNSPISIV
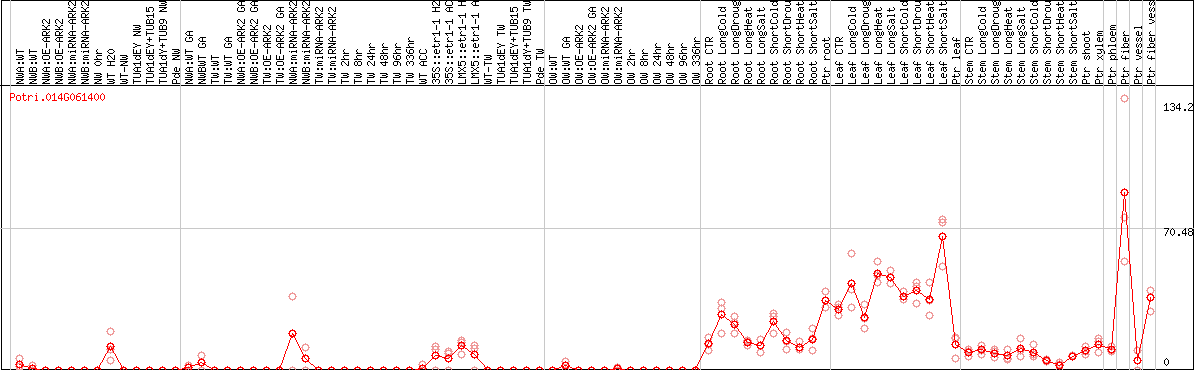
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.014G061400 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.