Potri.014G066000 [POPLAR]

| External link |
|
||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | |||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| ||||||||||||||||||||||||||||||
|
Paralogs
|
|
||||||||||||||||||||||||||||||
|
Flax homologues |
|
||||||||||||||||||||||||||||||
| PFAM info |
| ||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.014G066000.5 pacid=42763470 polypeptide=Potri.014G066000.5.p locus=Potri.014G066000 ID=Potri.014G066000.5.v4.1 annot-version=v4.1
ATGTCCGTACCAAACCCCAATAATAATTTTTCAAAGCACCAAAATATCCCTAACGCGCTCATGACTCAGAACCACCTTATCGGCTCACTCACTTCTCACA
TCTCTCTCTACCACTCCCAATCCAACCCCCCCTCCTCTCCAAACCCTAACCCTAGATCGTCAATCTTAAAATGGTTTAAATCTCTCTCCGTTCATCAACG
CCAATCCCACTTAACTACCGTTGATTTCAAATTCACTCAAATCCTCCTTCAAATGCTAGCCAAATTACACTCTCACGGTCACTGCCGTTTCATTATCCTC
CCCGATCTCCTCTCACGCGACCTCCCTAGCCTCTGCTTTAAGAAATCACGTGGGCTGCTGTCCAGAGTCGCTGAGTCTAATGAGTCGGAACGACTGATTT
TTGAGTCGACTCGGTTGTTTAGTTCTAGAGAAGGGGAGAAAGTGGACGATTGTAGGAGTGGAGCTGAGGGTCTGGATTCGGTTACGGTTAGCGAGGATTT
AATTGAAAATGTTGAAAAGTTTGTGGAGTTGATGGATGACATTTCGAATGGGGGGTTTTTGAGAGGGGAAGAGAGCGAGTTAGGGACGGATTGGGTGGAA
TTGGAGTGGTTAAAAGTGAGAGGTTATTATTGTATCGAGGCGTTTTTAGCGAACAAGCTAGAAGTGGCGTTGAGGCTGGCGTGGTTGAATTGTGGGAATG
GGAAGAAAAGAGGGGTGAAGTTGAAAGAGAAATTGAGTGCCGCAGGTGTGGCAGCTAATGTGTTTTGGAGGAGGAAAGGGTGTGTTGATTGGTGGAGGAA
TTTGGATGCGGAAGTAAGGAGGAAGGTTTTAAATTTTGCGCTTGGGAAGGCGGCAAAATCATTGACTCGTGAAATTCTGAAGGATGTGAGTGGTGTATCG
GGGGATGAGCTGTCACTGTTTAGAGCAGGAGTGCAGCGACCTTGGAGGGACTTGCATGCTGAATCACGGCAAAGGATTTTCCTAAAACTTCCAGCTGATG
CGGAATTTGGTTTGGCCCCAAAGCCTTCTTTTTCTGGGAAAGATGCTTCTTTTGCAAATATATTTAATAGTTTATTCGTGCTTCAGGATATTGTTAGTTT
GGTGTTGCCAGATCAGGGTAGTGAGTATGATACATCACATATATTTTTTAGCATGCTGGGTTCTCTTGGCACCCTTTCTGATTGTATATTAAGAAAACTA
CGAGGACTTGTGATGGTTATTTCACTTGATTGCACAAGGCTTGAACTTCTCGGAGAGGGAACATCAAACTCTTCAGCCAATAAACCTAGCGAAAAGCTCG
GTGCAGGTAGTCGTAGAAAAAAGGGGAAGACTCAAAATATGAAAAAGCTGATGAACCCCACACCAGTGAAAAGTGTTGACGAGTCTTCCTTTAAGAAACT
TGCCGAGGATATCAAATGTGCGCCAGCCTGTATTAAGAAGACAGAGTTGATGGAGTCCAATGAGATGCCTGGCATTCCTCATGAGAATGAAAATCACAGA
GACATATCATCTCCAACAGTTGAAATGGAACACACACAAGGATTGGTTCATGAAAAAAAACGAACTGCTGGAAGGAAGAACAGGAAAGGAAGAAACAAAA
AGAAAAAATCTAGCTTTAGCAATCCAGTTGAAGTAAGAAAACCTGAAATTGCCGTTTCAGAGGCTCCCTCCTTTTCTGTTTGCTCATCAGACGAGGAAGC
AAAGCTTTGTAGGTTATCTGACAATTTGACTACACAGAAAGCATCAAATGATAGTTTAATTGATCCTTCAATTAATGAGCCCACCAGGAAGGAGATAGAT
GCCCTTGGCATTCCAGAGGATCATGCTGTTGGTTGCACTGAAGGCATCTCTGATGCAGGTCTGGAGCATTACCGATCTTCAAATGGTTTTGTTGACAATA
AAAGTATGCCATCCAGAAGAGAGACACGTTGTGGGGTAGGCCAGAATATAATATATCAAGTTGCAACCACAAAGGAGCTAATAACTGTTTCTAGCAATGA
AGGCACCAGCTTTTTGAACAAAAAAACTGAGGTTAAATTAGATGTTGGCAACAAATTAGTCAGAACCCATGAAGTGAAAGAGGTACCCACCTTAAATCGG
GGAGAAGAGAGTGAAAATTTTCATGAAAGTGGATCCAAAGGCCTTTCAGATTGCCTTTCATATGAGTGGCCTAGTTTAGGCCCCGTCTACTTTCCATCTA
TAAATTCACATCTTCCACCTGCTACATATAGATTGCATCTGGATGTTGGTCACAACTGGCATAATCACATTCATCAACCTTTTTTACCTACGGTGCATCA
GGCTAGAAATTCTCCTATTGAAGGTGGATCCAATAGGATGTTGTCTCAACCATTGCCAATGAGTTTAGATTGGCCTCCAATGGTTAGGAGCAACTGTGGA
TTAGCTCCGACTATGACATGCAATTATGATTCAGGATTTATCTCGAGGCGGCAATCTACTTTTCAGAAAAGTTACACCGCTAAGAACATGCAGTACATCT
CAAAAACCTTTGATGACGAGAGGAGGTGTTCTGGGGATGCAATTGACTTTACTGAAGCAACAAGTTCACAGGAACTAATGGATGAATATGAAAACCATTG
GATATCAGAGGAAGAATATGAGGTGCATGCAGTTTCTGGGATAGACTATAATCAGCACTTCGGTGGCGGTGTGATGTACTGGGATCCTTCTGATCATCCA
GGTACTGGTTTCTCTCGACCACCTTCCCTTAGTTCTGATGATAGCGGATGGCCTTGGCATGAAGCAGAGCTGAATAGGGCAGTTGATGATATGGTTGCCT
TCTCATCTTCTTATAGTACAACTGGATTGACTTCCCCAACTGCTGCTTCATTCTGTTCTGCTTTTGATCCTTTGGTCCCTGGACACCAGGCACTTGGTTA
TGTTATGTCAGGAAATGAAGTACCTGGTAAAGCTATGCTTTCATCAACTGTGACAGATGCTGCTGCAGAGGAGGATGTCTCTGGATCTTTGGCCAGTTTA
TCTAGTGATGTTGAAGGCAAGGCAGGAGATTCACTTCCTTACCCAATTTTGCGGCCTATTATCATTCCAAATATGTCAAGGGAAAGATCAAGATCTGATT
TCAAGCGTAGTCTTGATCACAAAAGCCCTTGTGTTCCTCCTACTAGACGGGAACATCCCCGGATAAAGCGGCCACCATCACCTGTAGTGCTTTGTGTTCC
ACGGGCTCCCCGTCCTCCACCACCCTCTCCTGTGAGTGACTCTAGAAAGCACAGGGGTTTTCCAACTGTTAGGTCTGGTAGCTCCAGCCCAAGACAGTGG
GGTGTGAGAGGTTGGTACCATGATGGAACTAATTTGGAGGAAGCTTGTGGACGTATGGATGGTGCTGAAGTTGTTTGGCCTTCTTGGAGAAATAAAAAAC
TTTCAACCCATCCAATGGTGCAGCCACTCCCTGGGGCTTTGCTGCAAGATCGTCTAATTGCAATGTCCCACCTAGCCCGTGATCAGGATCATCCAGATGT
TTTGTTTCCTCTTCAACGAGCTGAGATACAGAATTGCCCTACACGAAAGGCTTCTCTTTGCTTAGTACAGAGCCTTCTTCATGATGAAATTGACTCTTTT
TGTAAGCAGGTTGCTGCAGCAAATATGGCCAGGAAACCTTTCATTAATTGGGCTGTCAAGCGGGTTACTAGGTCTCTCCAAGTTCTATGGCCTAGGTCCA
GGATAAATATCTTTGGATCAAGTGCAACTGGTTTGGCCCTTCCAACAAGTGATGTGGATCTTGTGGTTTGCCTCCCTCCAGTGAGGAACTTGGAACCTAT
CAAGGAAGCTGGGATTTTAGAGGGTCGTAATGGTATTAAAGAAACTTGTCTTCAGCATGCAGCTCGGTATCTTGCCAATCAGGAGTGGGTAAAAAATGAT
TCACTGAAGACTGTGGAGAATACAGCTATACCTGTTATTATGCTTGTAGTGGAGGTTCCCACTGATCTGATCACCTCTACTGCTTCTAATGTACAATCAC
CAAAGGAGGAGCCAATTCATTTGACTGTTGAACATGACATCCAAGTCCAGTCCAATATGGTTGTTTTAGAGGACTCAATTTCACCTAAGTGCACACAACT
TAATTGTGATAGCAAAAGGGATGTGAAATCAATTCGTCTTGATATCAGTTTCAAGTCTCCATCTCATACAGGGCTCCAAACAACGCAACTGGTTAAAGAT
CTCACGGAACAATTTCCAGCTGCAACACCTCTTGCTTTGGTACTGAAGCAGTTCTTGGCAGATCGTAGCCTTGACCAGTCCTATTCTGGAGGTTTAAGTT
CCTATTGCTTGGTGTTGCTAATTATACGTTTTCTACAGCATGAGCATCATCTTGGCCGGCCTATCAATCAAAATGTCGGAAGCCTTCTAATGGATCTTCT
TTACTTCTTCGGGAATGTGTTTGATCCTCGCCAAATGCGTATCTCAGTGCAGGGAAGTGGAGTATATATAAATAGGGAAAGAGGTTATAGTATTGATCCA
ATACACATTGATGACCCTCTTTTCCCAACAAATAATGTGGGGAGAAATTGCTTTCGAATACATCAATGCATCAAGGCTTTTTCAGAAGCTTATTCTGTTT
TGGAGAAGGAGCTGGCATGCCTTCCTGATGAAGGTGATACATGTTCAAGGCCAGCACACAGATTACTTCCAAAAATTATCCCAAGTATTGATATTACGGG
GAGTTTAATTATCTGA
|
||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.014G066000.5 pacid=42763470 polypeptide=Potri.014G066000.5.p locus=Potri.014G066000 ID=Potri.014G066000.5.v4.1 annot-version=v4.1
MSVPNPNNNFSKHQNIPNALMTQNHLIGSLTSHISLYHSQSNPPSSPNPNPRSSILKWFKSLSVHQRQSHLTTVDFKFTQILLQMLAKLHSHGHCRFIIL
PDLLSRDLPSLCFKKSRGLLSRVAESNESERLIFESTRLFSSREGEKVDDCRSGAEGLDSVTVSEDLIENVEKFVELMDDISNGGFLRGEESELGTDWVE
LEWLKVRGYYCIEAFLANKLEVALRLAWLNCGNGKKRGVKLKEKLSAAGVAANVFWRRKGCVDWWRNLDAEVRRKVLNFALGKAAKSLTREILKDVSGVS
GDELSLFRAGVQRPWRDLHAESRQRIFLKLPADAEFGLAPKPSFSGKDASFANIFNSLFVLQDIVSLVLPDQGSEYDTSHIFFSMLGSLGTLSDCILRKL
RGLVMVISLDCTRLELLGEGTSNSSANKPSEKLGAGSRRKKGKTQNMKKLMNPTPVKSVDESSFKKLAEDIKCAPACIKKTELMESNEMPGIPHENENHR
DISSPTVEMEHTQGLVHEKKRTAGRKNRKGRNKKKKSSFSNPVEVRKPEIAVSEAPSFSVCSSDEEAKLCRLSDNLTTQKASNDSLIDPSINEPTRKEID
ALGIPEDHAVGCTEGISDAGLEHYRSSNGFVDNKSMPSRRETRCGVGQNIIYQVATTKELITVSSNEGTSFLNKKTEVKLDVGNKLVRTHEVKEVPTLNR
GEESENFHESGSKGLSDCLSYEWPSLGPVYFPSINSHLPPATYRLHLDVGHNWHNHIHQPFLPTVHQARNSPIEGGSNRMLSQPLPMSLDWPPMVRSNCG
LAPTMTCNYDSGFISRRQSTFQKSYTAKNMQYISKTFDDERRCSGDAIDFTEATSSQELMDEYENHWISEEEYEVHAVSGIDYNQHFGGGVMYWDPSDHP
GTGFSRPPSLSSDDSGWPWHEAELNRAVDDMVAFSSSYSTTGLTSPTAASFCSAFDPLVPGHQALGYVMSGNEVPGKAMLSSTVTDAAAEEDVSGSLASL
SSDVEGKAGDSLPYPILRPIIIPNMSRERSRSDFKRSLDHKSPCVPPTRREHPRIKRPPSPVVLCVPRAPRPPPPSPVSDSRKHRGFPTVRSGSSSPRQW
GVRGWYHDGTNLEEACGRMDGAEVVWPSWRNKKLSTHPMVQPLPGALLQDRLIAMSHLARDQDHPDVLFPLQRAEIQNCPTRKASLCLVQSLLHDEIDSF
CKQVAAANMARKPFINWAVKRVTRSLQVLWPRSRINIFGSSATGLALPTSDVDLVVCLPPVRNLEPIKEAGILEGRNGIKETCLQHAARYLANQEWVKND
SLKTVENTAIPVIMLVVEVPTDLITSTASNVQSPKEEPIHLTVEHDIQVQSNMVVLEDSISPKCTQLNCDSKRDVKSIRLDISFKSPSHTGLQTTQLVKD
LTEQFPAATPLALVLKQFLADRSLDQSYSGGLSSYCLVLLIIRFLQHEHHLGRPINQNVGSLLMDLLYFFGNVFDPRQMRISVQGSGVYINRERGYSIDP
IHIDDPLFPTNNVGRNCFRIHQCIKAFSEAYSVLEKELACLPDEGDTCSRPAHRLLPKIIPSIDITGSLII
|
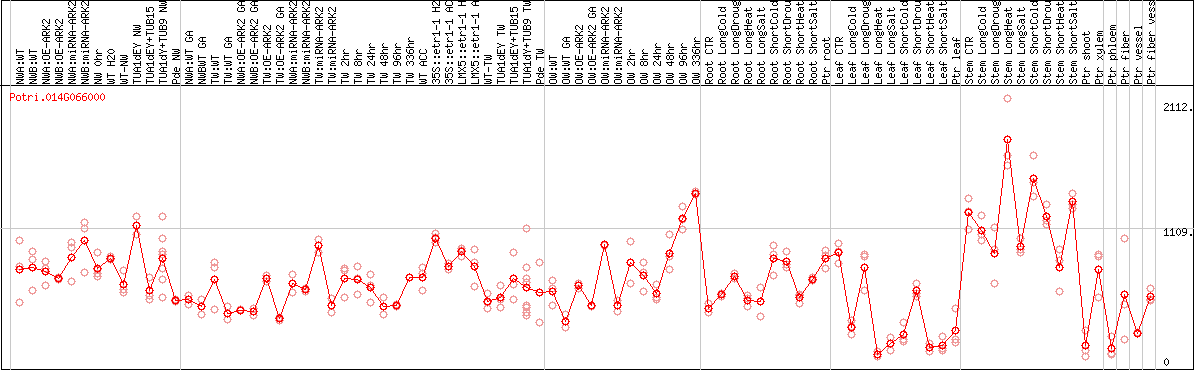
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.014G066000 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
