External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G45260 462 / 2e-163
Plant protein of unknown function (DUF641) (.1)
AT4G34080 281 / 1e-94
Plant protein of unknown function (DUF641) (.1)
AT4G33320 279 / 3e-93
Plant protein of unknown function (DUF641) (.1)
AT1G53380 189 / 2e-56
Plant protein of unknown function (DUF641) (.1), Plant protein of unknown function (DUF641) (.2), Plant protein of unknown function (DUF641) (.3)
AT3G14870 189 / 3e-56
Plant protein of unknown function (DUF641) (.1), Plant protein of unknown function (DUF641) (.2), Plant protein of unknown function (DUF641) (.3)
AT4G36100 169 / 3e-51
Sec1/munc18-like (SM) proteins superfamily (.1)
AT1G29300 176 / 4e-51
UNE1
unfertilized embryo sac 1, Plant protein of unknown function (DUF641) (.1)
AT3G60680 160 / 3e-45
Plant protein of unknown function (DUF641) (.1)
AT5G58960 118 / 7e-30
GIL1
GRAVITROPIC IN THE LIGHT, Plant protein of unknown function (DUF641) (.1), Plant protein of unknown function (DUF641) (.2), Plant protein of unknown function (DUF641) (.3)
AT2G30380 103 / 2e-24
Plant protein of unknown function (DUF641) (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.002G145900
553 / 0
AT2G45260 692 / 0.0
Plant protein of unknown function (DUF641) (.1)
Potri.001G387700
216 / 2e-66
AT1G53380 524 / 0.0
Plant protein of unknown function (DUF641) (.1), Plant protein of unknown function (DUF641) (.2), Plant protein of unknown function (DUF641) (.3)
Potri.011G108000
208 / 2e-63
AT1G53380 531 / 0.0
Plant protein of unknown function (DUF641) (.1), Plant protein of unknown function (DUF641) (.2), Plant protein of unknown function (DUF641) (.3)
Potri.013G155400
195 / 1e-58
AT2G45260 300 / 4e-98
Plant protein of unknown function (DUF641) (.1)
Potri.014G066600
182 / 1e-53
AT3G60680 522 / 0.0
Plant protein of unknown function (DUF641) (.1)
Potri.011G071000
180 / 6e-53
AT1G29300 483 / 3e-169
unfertilized embryo sac 1, Plant protein of unknown function (DUF641) (.1)
Potri.004G061600
175 / 4e-51
AT1G29300 485 / 6e-170
unfertilized embryo sac 1, Plant protein of unknown function (DUF641) (.1)
Potri.002G145000
171 / 5e-49
AT3G60680 494 / 4e-172
Plant protein of unknown function (DUF641) (.1)
Potri.019G128300
146 / 2e-40
AT2G45260 241 / 2e-75
Plant protein of unknown function (DUF641) (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10001989
490 / 2e-174
AT2G45260 671 / 0.0
Plant protein of unknown function (DUF641) (.1)
Lus10030291
490 / 2e-174
AT2G45260 668 / 0.0
Plant protein of unknown function (DUF641) (.1)
Lus10009923
182 / 1e-53
AT2G45260 275 / 2e-88
Plant protein of unknown function (DUF641) (.1)
Lus10024190
181 / 2e-53
AT2G45260 270 / 1e-86
Plant protein of unknown function (DUF641) (.1)
Lus10029259
171 / 3e-49
AT1G29300 419 / 6e-144
unfertilized embryo sac 1, Plant protein of unknown function (DUF641) (.1)
Lus10007312
170 / 7e-49
AT1G29300 433 / 2e-149
unfertilized embryo sac 1, Plant protein of unknown function (DUF641) (.1)
Lus10008587
159 / 7e-45
AT2G45260 266 / 1e-84
Plant protein of unknown function (DUF641) (.1)
Lus10009284
157 / 1e-43
AT3G60680 541 / 0.0
Plant protein of unknown function (DUF641) (.1)
Lus10005424
155 / 1e-43
AT1G29300 365 / 3e-123
unfertilized embryo sac 1, Plant protein of unknown function (DUF641) (.1)
Lus10016444
130 / 1e-33
AT5G58960 635 / 0.0
GRAVITROPIC IN THE LIGHT, Plant protein of unknown function (DUF641) (.1), Plant protein of unknown function (DUF641) (.2), Plant protein of unknown function (DUF641) (.3)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF04859
DUF641
Plant protein of unknown function (DUF641)
Representative CDS sequence
>Potri.014G067600.1 pacid=42763817 polypeptide=Potri.014G067600.1.p locus=Potri.014G067600 ID=Potri.014G067600.1.v4.1 annot-version=v4.1
ATGCAACCCACTGGGTTGAAAGATAATCAACCCCGTGAGAACAAATGTCAGAAGGTCCACCCTCAACCCATGGAAGATTCTGCAAATCAAAATCCAGAAG
CTGGGGAAGCCTTGATATCCAAAATATTTACCAACATCTCCTCTCTGAAGTCAGCATACATTCAGCTCCAAGCTGCTCATACTCCCTATGATCCTGATAA
AATACAAGCTTTTGACAAAGCTGTAATTTCTGAGCTGAAAAATCTATCCGAGCTAAATTATATCTACAGGGAAAATAACCCCAAACCAATATGTGTTTCT
CCTCAGGACTCTCGGTTAGCTGCAGAGATCCAAGAACAACTGAGCCTGCTCAAAACATACGAGAATAAAGATTCTGAGATTCTTCAGTCTGAGCAGATGA
TTGAGGAAGCAAACCAGAAACGAGCAAAACTGGAAAAGAATCTTAAGCTCAGGGGTTTCTGGTGGGATCTTGATGCCGCAGCTAACTCCATTGAATCCAA
CGTTGTTTATGCAAAGAGAGCTCACAAACAGTATGCATTTGAGTCTCATATATCTCAAAGAATGTTCATTGGGTTTCATCACGAGAACTTCTCAATTAAA
GCAGACGGCGGGGCAGTTTCAAAGGAGAGTTTCTTTCATCAATTTCTTGCTACGAGGGAAATGGATCCTTTAGACATGCTATGTCAGAACCCAAATTCTG
TTTTTGGGAAATTTTGCACGAGCAAGTACCTGGTGGTGGTTCACCCAAAGGGAGGGCATCCAAGAACGCCCTTCTACCAGGCCTTCTTGAAACTGGCCAA
GTCGATCTGGCTTTCTCACAGGCTTGCTTATTCCTTTGACCCAAATGTCAAGGTCTTCCAAGTTATGAGAGGAAGTGAGTTCTCAGAGCCTAGGGTTGGC
CTAATGGTTATGCCTGGTTTTTGGATAGGAGGCAGTGTGATTCAGAGCCCTGTTTATCTCTCAGGTGTGAAGGTTGCTGAATGA
AA sequence
>Potri.014G067600.1 pacid=42763817 polypeptide=Potri.014G067600.1.p locus=Potri.014G067600 ID=Potri.014G067600.1.v4.1 annot-version=v4.1
MQPTGLKDNQPRENKCQKVHPQPMEDSANQNPEAGEALISKIFTNISSLKSAYIQLQAAHTPYDPDKIQAFDKAVISELKNLSELNYIYRENNPKPICVS
PQDSRLAAEIQEQLSLLKTYENKDSEILQSEQMIEEANQKRAKLEKNLKLRGFWWDLDAAANSIESNVVYAKRAHKQYAFESHISQRMFIGFHHENFSIK
ADGGAVSKESFFHQFLATREMDPLDMLCQNPNSVFGKFCTSKYLVVVHPKGGHPRTPFYQAFLKLAKSIWLSHRLAYSFDPNVKVFQVMRGSEFSEPRVG
LMVMPGFWIGGSVIQSPVYLSGVKVAE
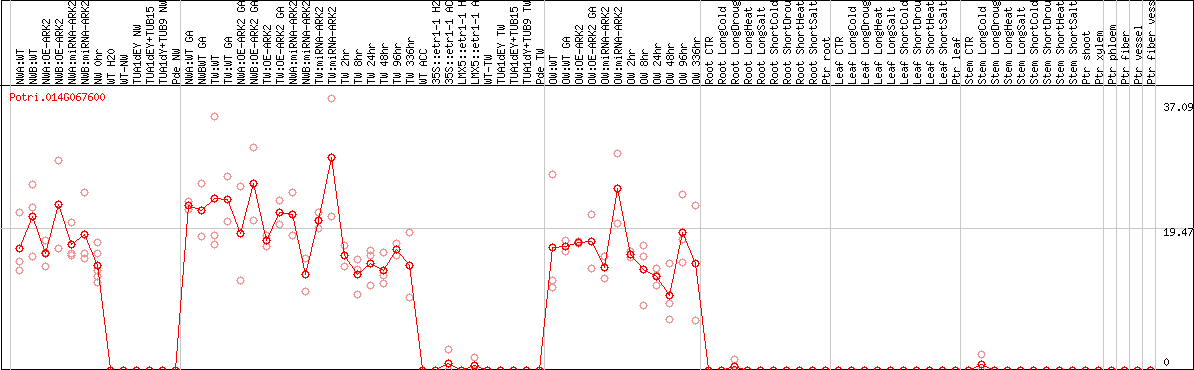
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.014G067600 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.