External link
Symbol
PBF1.2
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G60820 363 / 5e-129
PBF1
N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.1), N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.2), N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.3)
AT1G21720 82 / 4e-19
PBC1
proteasome beta subunit C1 (.1)
AT1G77440 80 / 2e-18
PBC2
20S proteasome beta subunit C2 (.1.2)
AT4G31300 61 / 5e-11
PBA1
N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.1), N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.2), N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.3)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.002G148300
431 / 4e-156
AT3G60820 387 / 2e-138
N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.1), N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.2), N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.3)
Potri.002G080800
86 / 9e-21
AT1G21720 383 / 1e-137
proteasome beta subunit C1 (.1)
Potri.005G180500
86 / 2e-20
AT1G21720 391 / 9e-141
proteasome beta subunit C1 (.1)
Potri.006G077900
66 / 8e-13
AT4G31300 416 / 2e-149
N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.1), N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.2), N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.3)
Potri.008G155500
51 / 8e-08
AT3G22630 364 / 6e-130
20S proteasome beta subunit D1 (.1)
Potri.010G084800
50 / 3e-07
AT3G22630 366 / 6e-131
20S proteasome beta subunit D1 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10015867
371 / 4e-132
AT3G60820 379 / 2e-135
N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.1), N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.2), N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.3)
Lus10009294
309 / 6e-100
AT1G48050 835 / 0.0
ARABIDOPSIS THALIANA KU80 HOMOLOG, Ku80 family protein (.1)
Lus10000062
220 / 9e-74
AT3G60820 232 / 6e-79
N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.1), N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.2), N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.3)
Lus10004024
171 / 2e-53
AT3G60820 190 / 4e-61
N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.1), N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.2), N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.3)
Lus10030271
85 / 1e-21
AT3G60820 82 / 3e-21
N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.1), N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.2), N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.3)
Lus10030272
83 / 7e-21
AT3G60820 81 / 2e-20
N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.1), N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.2), N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.3)
Lus10020599
70 / 1e-14
AT1G21720 355 / 7e-127
proteasome beta subunit C1 (.1)
Lus10004885
69 / 3e-14
AT1G21720 356 / 2e-127
proteasome beta subunit C1 (.1)
Lus10020180
59 / 3e-10
AT4G31300 429 / 7e-155
N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.1), N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.2), N-terminal nucleophile aminohydrolases (Ntn hydrolases) superfamily protein (.3)
Lus10041194
54 / 8e-09
AT3G22630 363 / 9e-130
20S proteasome beta subunit D1 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0052
NTN
PF00227
Proteasome
Proteasome subunit
Representative CDS sequence
>Potri.014G069800.2 pacid=42763610 polypeptide=Potri.014G069800.2.p locus=Potri.014G069800 ID=Potri.014G069800.2.v4.1 annot-version=v4.1
ATGACTAAGCAACAAGCAAGATTTGAACCCTACGATTTTAATGGAGGGACTTGTGTTGCGATTGCTGGAGCTGATTACTGTGTTGTCGCTGCTGATACTC
GAATGTCTACCGGATACAATATCCTCACTCGCGATTACTCCAAAATCTGTAAACTAGCAGACAAATGTTTGATGGCCTCTTCAGGGTTTCAAGCTGATGT
CAAAGCCCTACAAAAGGTGTTGGGTGCCAAGCACCTGATCTATCAACATCAACACAACAAGCAGATGAGCTGCCCTGCCATGGCTCGACTGCTCTCCAAC
ACTCTCTACTACAAACGTTTCTTCCCTTATTATACCTTCAATGTTTTGGTTGGCTTTGACGAGGAAGGAAAGGGCTGTGTTTACACTTATGATGCTGTTG
GTTCCTATGAGAAGGTTGGGTATAGTGCCCAAGGTTCTGGTGCTAAACTCATCATGCCTGTGCTGGATAACCAATTGAAGTCCCCCAGCCCGCTTTTATT
ACCTGCTCTGGATGCTGTAACTCCACTTACTGAGGCAGAAGCGATTGATGTGGTCAAAGATGTTTTTGCATCTGCAACTGAGAGGGATATATACACTGGA
GACAAACTTGAAATTGTCATCGTAAATGCTGATGGTATTCGTCGTGAATACGCGGAACTCAGAAAAGATTGA
AA sequence
>Potri.014G069800.2 pacid=42763610 polypeptide=Potri.014G069800.2.p locus=Potri.014G069800 ID=Potri.014G069800.2.v4.1 annot-version=v4.1
MTKQQARFEPYDFNGGTCVAIAGADYCVVAADTRMSTGYNILTRDYSKICKLADKCLMASSGFQADVKALQKVLGAKHLIYQHQHNKQMSCPAMARLLSN
TLYYKRFFPYYTFNVLVGFDEEGKGCVYTYDAVGSYEKVGYSAQGSGAKLIMPVLDNQLKSPSPLLLPALDAVTPLTEAEAIDVVKDVFASATERDIYTG
DKLEIVIVNADGIRREYAELRKD
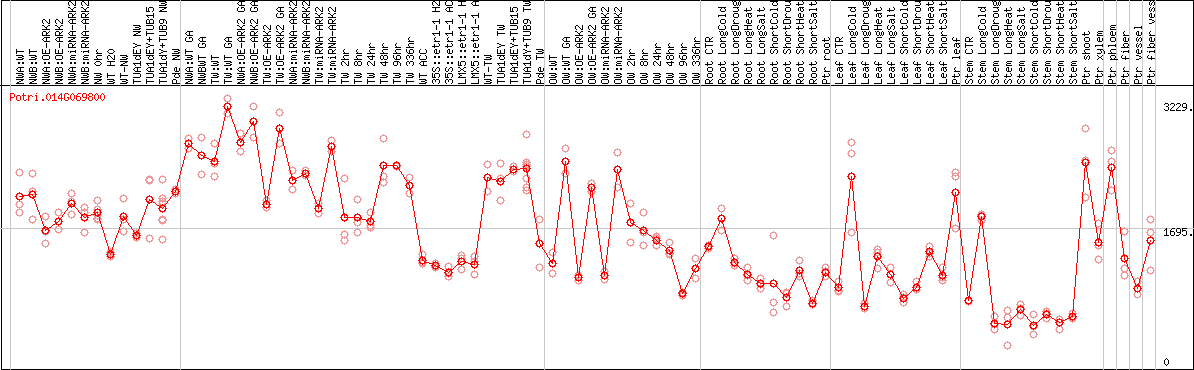
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.014G069800 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.