External link
Symbol
LBD18.1
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G45420 236 / 3e-78
AS2
LBD18, ASL20
LOB domain-containing protein 18 (.1)
AT4G00220 219 / 6e-72
AS2
JLO, LBD30, ASL19
LOB DOMAIN-CONTAINING PROTEIN 30, JAGGED LATERAL ORGANS, Lateral organ boundaries (LOB) domain family protein (.1)
AT4G00210 176 / 8e-55
AS2
LBD31
LOB domain-containing protein 31 (.1)
AT3G03760 175 / 6e-54
AS2
LBD20
LOB domain-containing protein 20 (.1)
AT2G45410 171 / 3e-53
AS2
LBD19
LOB domain-containing protein 19 (.1)
AT3G58190 168 / 4e-52
AS2
LBD29, ASL16
ASYMMETRIC LEAVES 2-LIKE 16, lateral organ boundaries-domain 29 (.1)
AT2G42430 169 / 7e-52
AS2
ASL18, LBD16
ASYMMETRIC LEAVES2-LIKE 18, lateral organ boundaries-domain 16 (.1)
AT2G42440 161 / 5e-49
AS2
Lateral organ boundaries (LOB) domain family protein (.1)
AT5G06080 153 / 1e-46
AS2
LBD33
LOB domain-containing protein 33 (.1)
AT2G31310 150 / 2e-45
AS2
LBD14
LOB domain-containing protein 14 (.1)
Paralogs
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10009298
177 / 3e-55
AT2G45410 193 / 2e-62
LOB domain-containing protein 19 (.1)
Lus10000613
171 / 4e-53
AT3G58190 215 / 1e-70
ASYMMETRIC LEAVES 2-LIKE 16, lateral organ boundaries-domain 29 (.1)
Lus10033873
171 / 5e-53
AT3G58190 220 / 1e-72
ASYMMETRIC LEAVES 2-LIKE 16, lateral organ boundaries-domain 29 (.1)
Lus10024877
166 / 2e-50
AT2G42430 211 / 6e-68
ASYMMETRIC LEAVES2-LIKE 18, lateral organ boundaries-domain 16 (.1)
Lus10000707
162 / 4e-49
AT2G42430 222 / 2e-72
ASYMMETRIC LEAVES2-LIKE 18, lateral organ boundaries-domain 16 (.1)
Lus10033872
159 / 3e-48
AT2G42430 212 / 6e-69
ASYMMETRIC LEAVES2-LIKE 18, lateral organ boundaries-domain 16 (.1)
Lus10014757
159 / 4e-48
AT2G42430 208 / 2e-67
ASYMMETRIC LEAVES2-LIKE 18, lateral organ boundaries-domain 16 (.1)
Lus10004707
142 / 4e-42
AT5G06080 154 / 1e-47
LOB domain-containing protein 33 (.1)
Lus10022264
128 / 1e-36
AT5G63090 223 / 9e-75
Lateral organ boundaries (LOB) domain family protein (.1), Lateral organ boundaries (LOB) domain family protein (.2), Lateral organ boundaries (LOB) domain family protein (.3), Lateral organ boundaries (LOB) domain family protein (.4)
Lus10027676
128 / 1e-36
AT5G63090 233 / 8e-79
Lateral organ boundaries (LOB) domain family protein (.1), Lateral organ boundaries (LOB) domain family protein (.2), Lateral organ boundaries (LOB) domain family protein (.3), Lateral organ boundaries (LOB) domain family protein (.4)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF03195
LOB
Lateral organ boundaries (LOB) domain
Representative CDS sequence
>Potri.014G070400.1 pacid=42763216 polypeptide=Potri.014G070400.1.p locus=Potri.014G070400 ID=Potri.014G070400.1.v4.1 annot-version=v4.1
ATGAGTACCGCAACTACAAACCCTATAAATACTGGTGCTGGTGGTGGAAGTAGTGGCGGTGGAGGTGGTGGTGGCGGCGGTGGTGGCGGTGGTGGTGGGC
CATGCGGTGCTTGTAAATTCTTGAGGAGGAAGTGTGTGCCAGGGTGTATATTTGCACCTTATTTTGACTCAGAGCAAGGTGCAGCACATTTTGCAGCGGT
GCATAAGGTTTTTGGTGCAAGTAACGTGTCAAAGCTTTTATTACACATTCCTGTCCATAAAAGACTTGATGCTGTGGTCACAATTTGTTATGAGGCTCAA
GCTCGCTTAAGAGATCCAGTCTATGGCTGTGTTGCTCACATCTTTGCTCTTCAACAACAGGTAGTAAATTTACAAGCAGAACTCTCATACTTGCAAGCCC
ATCTAGCAGCATTGGAGGTTCCGTCGCCACCTCCTCCGCCCCCGCCAACACTAGTGACCCCGCCTCCACTCTCAATAGCAGACCTACCATCAGCCTCTTC
AATTCCCGCCGCGTACGATTTATCATGTCTTTTTGATCCAATGATGCAACCCTCATGGTCCAACATGCAGCCGCAGCGGCAAATGGATATCCGTCAGTTT
GGAGGCAGCGGTGGCTCATCAGGAACTGGTGGCGGTGATCTCCAAGCATTGGCGCGTGAACTCCTTCATAGACGAGGATCTCCACCACCCGGGTTCATGT
CATGTAGTGACACATTAGCATCACCATCTATCTCTAAATGA
AA sequence
>Potri.014G070400.1 pacid=42763216 polypeptide=Potri.014G070400.1.p locus=Potri.014G070400 ID=Potri.014G070400.1.v4.1 annot-version=v4.1
MSTATTNPINTGAGGGSSGGGGGGGGGGGGGGGPCGACKFLRRKCVPGCIFAPYFDSEQGAAHFAAVHKVFGASNVSKLLLHIPVHKRLDAVVTICYEAQ
ARLRDPVYGCVAHIFALQQQVVNLQAELSYLQAHLAALEVPSPPPPPPPTLVTPPPLSIADLPSASSIPAAYDLSCLFDPMMQPSWSNMQPQRQMDIRQF
GGSGGSSGTGGGDLQALARELLHRRGSPPPGFMSCSDTLASPSISK
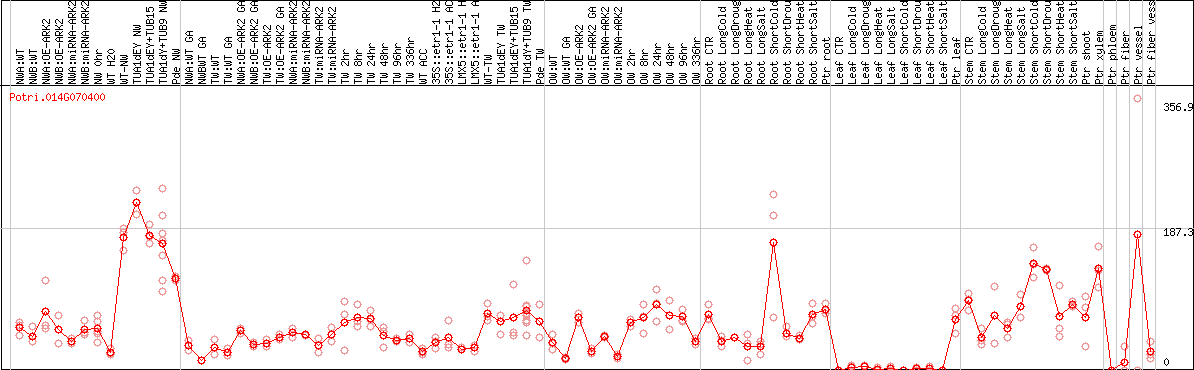
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.014G070400 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.