Potri.014G098600 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.014G098600.5 pacid=42762928 polypeptide=Potri.014G098600.5.p locus=Potri.014G098600 ID=Potri.014G098600.5.v4.1 annot-version=v4.1
ATGGCACCCAGGAACAATCGTGGAAAGGCTAGAGGAGAGAAGAGGAAGAAGGACGAGAAGGTTCTTCCTGTGGTTACTGATATCACCATTAACCTTCCTG
ACGAAACTCATGTTGTTTTGAAGGGAATCTCAACAGACAGAATCATAGATGTTCGTCGACTTTTATCGGTGAACACTGAAACTTGTTACATCACAAATTT
CTCGTTGTCGCATGAGGTTAGAGGAGCGCGGTTAAAAGACACGGTCGACGTGTCAGCACTCAAGCCTTGCGTTCTCACTTTAACCAATGAGGACCTCGAT
GAGGAGCTTGCAGTGGCGCACGTTAGACGACTCCTGGACATCGTGGCATGCACAACGTGCTTCGGTCCATCGGCTTGTGCGCATGATAAGATCAAGTCCG
ATATCGGAAAGAATGCGCCGGCTGCGCAGGATAATAAGACTTCTAAGAAAACCACCGCTAAATCTCAATCCTCCTCCACCACCACCACCACCACCACGAC
GAATAAACAATCATCGTCACCAAAATCAGCATCAAAAGATGTTCCGGTGGATGCAGAAGAAGAAATGAGCCACTCGTGCCCCAAGCTTGGGAGCTTTTAC
GAGTTCTTCTCTCTCTCGCATCTCACTCCTCCTCTTCAATTTATAAGGAAGGTAACGAAAAGACGGATTGATGAGATCTCCGTGGATGATCATCTTTTCT
CTCTTGACGTGAAGCTATGCAATGGGAAGCTGGTTCAAGTTGAAGCTTGCAAAAAGGGATTTTATGGTGTTGGAAAGCAGAGGATTTTATGTCACAATCT
TGTTGATTTGCTAAGGCAGCTTAGTCGAGCTTTTGATAATGCTTACGATGAGCTCATGAAAGCATTTGCAGAGCGTAACAAGTTCGGGAATCTTCCATAT
GGTTTTAGAGCAAACACATGGCTTATCCCTCCCGTAGCAGCACAGTTGCCATCAGTTTGTCCTCCTCTTCCTGTTGAGGATGAAACATGGGGAGGAAATG
GAGGTGGTCTTGGAAGAGATGGGAAAAAGGATTACATACCTTGGGCCGACGAGTTTTTGTTTGTCGCATCCATGCCTTGCAAGACAGCTGAGGAGAGGCA
GATTCGGGACAGGAAAGCCTTCCTGCTTCACAGCCTATTTGTCGATGTTGCCCTTTTCAGAGCCATTAAAGCTGTGCAACATGTAAAATTAAAGCCAAAT
TTATTGGGTTCAGTTGCAAACAGTAATATTCCTTACACAGAGAGAGTTGGGGATTTAAGCATCAAAGTTATGAAAGATGCTACCAATGCAAGCTCTAAGG
TAGACACTAAAATTGATGGGATTCAAGCAACTGGAACTGATAAGAAAAATTCAGTAGAGAGGAACCTTCTGAAAGGGATCACAGCTGATGAGAACACTGC
TGCTCATGACATTGCAACACTTGGTACTGTTAATGTAAGATATTGTGGTTTCATTGCTATTGTAAAAGCTGAGGCGAGGGAGGAAAAGAAAGCCAGTCCT
CCATCCAAAAGTATTGACCTTGAACAGCCTGAAGGTGGTGCAAATGCCCTTAACATCAACAGCTTGAGACTACTACTTCACAAACCAACACCTTCAGAAC
ACACTAAGCGGACACCAAATTTGCAAACTTTAGAGTGTGAAGAGCTCAGTGCTTCCGAAGCATTTGTAGAGAGATTACTGGAAGAAAGTCTTACCAGGCT
TGAAGAAGAGGTACCGAAGCAGGATCATTTGGTGAGATGGGAACTTGGAGCCTGCTGGATACAGCATTTGCAAGACCAGAAAAATACTGAAAAAGATAAG
AAACCATCCACTGAGAAGGGCAAGAAGCCTTCCACAGAAACAGAAATGAAGGTTGAGGGACTTGGGACCCCTCTTAAGTCCCTTAAGAACAAGAAAAAAT
CAGATGAAAGCAATGTAAAAATGCAGCCGGAGAATTCAAGACCTGCCTCTGATGGTTTAAGTGGGGCAGTTGAAGATGCTACACTGGCATCTGTGGAATC
TCACCTTGAAACTGAGGCTAAAGATAATGAGCTTGCACTGCAGCAGTTGTTGTCCGATGCAGCCTTTGCTCGACTAAAAGAATCAGACACTGGACTTCAT
TGCAAGTCTCTGCAACAGCTAATTGACTTGTCTCAGAAATATTACACTGAAGTTGCTCTTCCTAAACTGGTAGCAGATTTTGGTTCTTTGGAGCTTTCAC
CCGTTGATGGTCGGACTCTAACTGATTTTATGCATACTAGAGGTCTTCAGATGTGCTCTCTTGGGCGTGTTGTAAAGCTTTCTGAAAAGCTGTTGCATGT
ACAGTCACTCTGCATTCATGAGATGATAGTACGAGCTTTTAAGCATATACTTCAGGCTGTGATTGCTGCTGTTGTGGACCAAGAGAAAATGGCTGTATCA
ATAGCTGCTGCATTAAATCTGATGCTCGGGATTCCTGAAACCAGAGATTCAATCAAGTCTTGCCATGTCCACCCTCTTGTATGGAGATGGCTGGAGGTAT
TTTTGAAGAAGCGGTATGAATGGGATCTTAGCAGCTTAAACTTCAAAGATGTGAGGAAATTTGCAATCTTGCGAGGGTTGTGCCACAAGGTGGGCATTGA
GTTGGTTCCGAGGGATTTTGATATGGATTCTCCACATCCATTCAGAAAATCAGATGTTGTCAGTCTGGTTCCACTACATAAGCAAGCAGCATGCTCCTCT
GCAGATGGAAGGCAACTCCTGGAATCATCAAAAACAGCCTTGGACAAGGGAAAACTTGAAGATGCAGTCACCTATGGGACAAAGGCTCTTGCAAAGCTAG
TGGCAGTTTGTGGTCCCTACCATCGAATGACTGCAGGAGCATACAGCCTTCTTGCTGTAGTATTATATCACACTGGAGACTTCAATCAGGCCACTATATA
TCAGCAAAAAGCGTTGGATATTAATGAGAGGGAACTAGGACTTGATCATCCAGACACAATGAAAAGCTATGGGGATCTTGCCGTGTTCTATTACAGACTT
CAACATACAGAGCTGGCTCTCAAGTATGTAAAGCGTGCTTTGTATCTCTTACATCTCACATGCGGTCCATCACACCCAAACACTGCTGCAACATACATCA
ATGTGGCTATGATGGAGGAAGGTCTGGGGAATGTGCATGTCGCCCTCAGATATCTTCATAAAGCTCTAAAATGCAACCAAAAGTTACTTGGACCAGACCA
TATTCAGACTGCTGCAAGCTACCATGCAATAGCAATTGCACTCTCATTGATGGAAGCATATCCTTTGAGTGTTCAACATGAACAAACAACCTTGCAAATT
CTCCGAGCAAAGCTGGGTCCAGATGATTTACGCACTCAGGATGCTGCTGCTTGGCTTGAGTATTTTGAATCCAAGGCTTTTGAACAGCAAGAAGCTGTGC
GAAATGGTACTAAGAAGCCTGATGCATCAATTGCTAGCAAAGGCCACCTTAGTGTATCAGATTTGCTGGACTATATTAATCCAAGTCGTGATGCTAAGGT
GAGAGATGTTGTCGCTGGAAAGAGAAAGAGCTATATCACAAAGGTAAAGGATAAAACCCAGCCAAATGTCAGTACGGCAAGTTCCGACGAATCTACAAAA
GATACTCTTAAAGATGCTTCAGATGTGAAGATACCAGTGCCTGAAGATGATGCTTCCCAAGAGACCAGCTCTGCCCAAGTTCAACTCCAGACACCTGCTG
TGGAAGAGAATGTGGAAAAAAAACCAAGCATCTGGACTGAAGCCTTGCTTGAAACTCATGCTGAAGGAGATGATGGATGGCAACCAGTTCAAAGGCCAAG
ATCAGCTGGCTTGTATGGGCGACGTCTAAAGCAGCGGCGGGGGATTGTTGGAAAAGTTTATAGTTATCACAAAAAGATTGTGGATGCTAACATGGATTAT
GCTCCTGTCAAGAATGCTCACCAGAATAGTAAGTATTACCTTTTAAAGAAACGAGCACCATCTCATGGAAGTTATGGAGACCACCAAACCACCAATCTCC
CTCCAAGTGCCAAATTTGGGAGGAGAATGGTTAAAGCTGTAACCTACAGGGTCAAGTCTGTTCCTTCATCTTATAAAACTTCTACAACAGAGAATCCCAG
GATTGGCAACAAGGCTCTTACTTCATCAGAATCGGCTCCAGTTTCTGCCCCTAATGACATTCGTCCATCAAAGAATTCAATAGTGAGTCTTGGAAAATCT
CTTTCATACAAGGAAGTAGCATTGGCACCACCAGGTACCATTGCTAAGTTGCAGGCGTGGTTTCCCCAAAGTGATAATTCAGATAACCAGGAAATTGGTG
ATGGAAAACTTGAAGAGACAAATGAAGCAAAAGCAATTGCTGGTTCAGTAGTAATGGGTGTGGAAGAAAGAAGTGGAGAGAAGGATGAGAATTCTGAATC
AGATGACACGGATGATTTGAAGAAGGAAATAGTTGGTGTTCATAAAATGGAAGAACAACACTCAACTCACGTGTTGGAAGAAAATTCCTCCCTGATGGTG
TCTCAGAGTGTGCAAGGACATGAATCTGGTGATATTGAGGTTCATGAAATTATACAGAATGGAATGTTGATTGACCAGATCCCAAATTCAATTGATTCCC
TTCCAAAGGAACCTCATGAGAAGGATTCATCTAGTGAATTTGATCCACAAGTTGATCTCAACTCCACTTTGCCTGGAGCAGAGGACTTGAAGGACAAACC
ATTGATTTTGAATTCAGGCGATGCTCAGGGATTACCAAATAAGAAATTGTCTGCTTCTGCAGCTCCTTTCAACCCATCAACATCCATTGGACGTGCTCCA
CCAGTTGCCATCAACATCCCTCTTCCTTCTGCTCCTGGTGCTGTTCCAGCTGTTGCACCTTGGCCTGTAAACATGACTCTTCACCCTGGACCTGCAACTG
TCATACGCCCCATTAATCCAATGTCCTCCCCACACCATCCATACCCATACCCCTCACAACCACCAACCCCAAACATGATACAGCCATTGCCCTTTATGTA
TCCTCCTTACAGTCAAGCAGTACCAACTAGTACATTTCCTGTTACTAGCAGCGCCTTCCATCCTAATCATTTCTCCTGGCAGTGCAATGCGAGCCCCAAT
GTGTCAGAGTTTATTCCTACCACAGTATGGCCTGGCTGCCTTGCAGTGGAGTTCTCTGTCCTGCCACCTGTTGTTGAGCCAATTGCTGATCCTCTATTGG
AGCCTAAAGCACAATTTGAAAATTCTGAAAGCCCAAGTCCACCTCCAATTCTGTCAGTGGACAGTGACAACATAGGAGAAACAAATGACGAAGCAAATCT
TCAAGCATCAGACAGAAATGACAATGTGAAGGAATTGACTGGGGCTGGGTTGGAAAATATAAAGGAAAATGGTCATTCAAATCCATCCGAAGCTGAAATT
TATAGGAATGATTCGAGTCAGGAAAAAGGCTCACAAGAAAATGTCACTAGCAGTATTGACCAGCAGATTAATGAGGAGAAAACCTTCAGCATTTTGTTAA
GGGGAAAGAGAAACCGGAAGCAGACTCTGAGAATGCCAATGAGCTTGCTTAGTCGACCATATGGCTCACAGTCTTTTAAAGTTATATATAACAGAGTGGT
AAGAGGCAGTGAATCTCCCAAATCTACAAGCTTTGCTGCAGGTGAAGGTTGCACAACTAGTGCTACATGA
|
|||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.014G098600.5 pacid=42762928 polypeptide=Potri.014G098600.5.p locus=Potri.014G098600 ID=Potri.014G098600.5.v4.1 annot-version=v4.1
MAPRNNRGKARGEKRKKDEKVLPVVTDITINLPDETHVVLKGISTDRIIDVRRLLSVNTETCYITNFSLSHEVRGARLKDTVDVSALKPCVLTLTNEDLD
EELAVAHVRRLLDIVACTTCFGPSACAHDKIKSDIGKNAPAAQDNKTSKKTTAKSQSSSTTTTTTTTNKQSSSPKSASKDVPVDAEEEMSHSCPKLGSFY
EFFSLSHLTPPLQFIRKVTKRRIDEISVDDHLFSLDVKLCNGKLVQVEACKKGFYGVGKQRILCHNLVDLLRQLSRAFDNAYDELMKAFAERNKFGNLPY
GFRANTWLIPPVAAQLPSVCPPLPVEDETWGGNGGGLGRDGKKDYIPWADEFLFVASMPCKTAEERQIRDRKAFLLHSLFVDVALFRAIKAVQHVKLKPN
LLGSVANSNIPYTERVGDLSIKVMKDATNASSKVDTKIDGIQATGTDKKNSVERNLLKGITADENTAAHDIATLGTVNVRYCGFIAIVKAEAREEKKASP
PSKSIDLEQPEGGANALNINSLRLLLHKPTPSEHTKRTPNLQTLECEELSASEAFVERLLEESLTRLEEEVPKQDHLVRWELGACWIQHLQDQKNTEKDK
KPSTEKGKKPSTETEMKVEGLGTPLKSLKNKKKSDESNVKMQPENSRPASDGLSGAVEDATLASVESHLETEAKDNELALQQLLSDAAFARLKESDTGLH
CKSLQQLIDLSQKYYTEVALPKLVADFGSLELSPVDGRTLTDFMHTRGLQMCSLGRVVKLSEKLLHVQSLCIHEMIVRAFKHILQAVIAAVVDQEKMAVS
IAAALNLMLGIPETRDSIKSCHVHPLVWRWLEVFLKKRYEWDLSSLNFKDVRKFAILRGLCHKVGIELVPRDFDMDSPHPFRKSDVVSLVPLHKQAACSS
ADGRQLLESSKTALDKGKLEDAVTYGTKALAKLVAVCGPYHRMTAGAYSLLAVVLYHTGDFNQATIYQQKALDINERELGLDHPDTMKSYGDLAVFYYRL
QHTELALKYVKRALYLLHLTCGPSHPNTAATYINVAMMEEGLGNVHVALRYLHKALKCNQKLLGPDHIQTAASYHAIAIALSLMEAYPLSVQHEQTTLQI
LRAKLGPDDLRTQDAAAWLEYFESKAFEQQEAVRNGTKKPDASIASKGHLSVSDLLDYINPSRDAKVRDVVAGKRKSYITKVKDKTQPNVSTASSDESTK
DTLKDASDVKIPVPEDDASQETSSAQVQLQTPAVEENVEKKPSIWTEALLETHAEGDDGWQPVQRPRSAGLYGRRLKQRRGIVGKVYSYHKKIVDANMDY
APVKNAHQNSKYYLLKKRAPSHGSYGDHQTTNLPPSAKFGRRMVKAVTYRVKSVPSSYKTSTTENPRIGNKALTSSESAPVSAPNDIRPSKNSIVSLGKS
LSYKEVALAPPGTIAKLQAWFPQSDNSDNQEIGDGKLEETNEAKAIAGSVVMGVEERSGEKDENSESDDTDDLKKEIVGVHKMEEQHSTHVLEENSSLMV
SQSVQGHESGDIEVHEIIQNGMLIDQIPNSIDSLPKEPHEKDSSSEFDPQVDLNSTLPGAEDLKDKPLILNSGDAQGLPNKKLSASAAPFNPSTSIGRAP
PVAINIPLPSAPGAVPAVAPWPVNMTLHPGPATVIRPINPMSSPHHPYPYPSQPPTPNMIQPLPFMYPPYSQAVPTSTFPVTSSAFHPNHFSWQCNASPN
VSEFIPTTVWPGCLAVEFSVLPPVVEPIADPLLEPKAQFENSESPSPPPILSVDSDNIGETNDEANLQASDRNDNVKELTGAGLENIKENGHSNPSEAEI
YRNDSSQEKGSQENVTSSIDQQINEEKTFSILLRGKRNRKQTLRMPMSLLSRPYGSQSFKVIYNRVVRGSESPKSTSFAAGEGCTTSAT
|
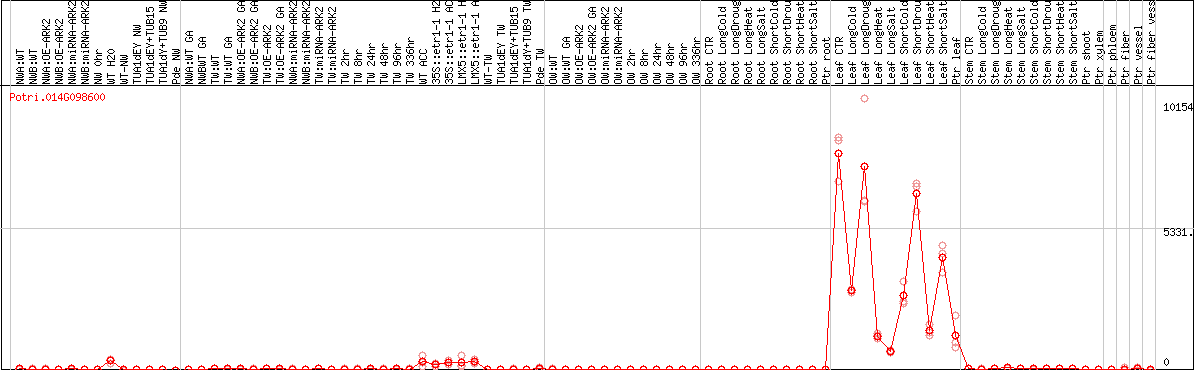
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.014G098600 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
