External link
Symbol
DAG1.2
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G61850 233 / 1e-75
DOF
DAG1, AtDof3,7
dof affecting germination 1, Dof-type zinc finger DNA-binding family protein (.1.2.3.4)
AT2G46590 195 / 2e-60
DOF
DAG2, AtDof2,5
DOF AFFECTING GERMINATION 2, Dof-type zinc finger DNA-binding family protein (.1.2)
AT4G24060 148 / 1e-42
DOF
AtDof4,6
Dof-type zinc finger DNA-binding family protein (.1)
AT4G00940 138 / 3e-39
DOF
AtDof4,1
Dof-type zinc finger DNA-binding family protein (.1)
AT1G64620 129 / 4e-35
DOF
AtDof1,8
Dof-type zinc finger DNA-binding family protein (.1)
AT3G55370 123 / 9e-33
DOF
OBP3, AtDof3. 6
OBF-binding protein 3 (.1.2.3)
AT5G60200 120 / 1e-32
DOF
TMO6, AtDof5,3
TARGET OF MONOPTEROS 6 (.1)
AT3G45610 114 / 2e-30
DOF
DOF6, AtDof3,2
DOF transcription factor 6, Dof-type zinc finger DNA-binding family protein (.1)
AT2G37590 114 / 1e-29
DOF
AtDof2. 4, ATDOF2.4
DNA binding with one finger 2.4 (.1)
AT1G51700 109 / 2e-29
DOF
ADOF1, AtDof1. 7
DOF zinc finger protein 1 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.002G174300
413 / 2e-147
AT3G61850 229 / 8e-74
dof affecting germination 1, Dof-type zinc finger DNA-binding family protein (.1.2.3.4)
Potri.003G144500
278 / 4e-94
AT4G24060 201 / 1e-62
Dof-type zinc finger DNA-binding family protein (.1)
Potri.001G086400
273 / 3e-92
AT4G24060 176 / 5e-53
Dof-type zinc finger DNA-binding family protein (.1)
Potri.012G018700
226 / 1e-73
AT3G61850 141 / 9e-40
dof affecting germination 1, Dof-type zinc finger DNA-binding family protein (.1.2.3.4)
Potri.015G009300
197 / 2e-62
AT4G24060 154 / 1e-44
Dof-type zinc finger DNA-binding family protein (.1)
Potri.012G081300
124 / 8e-34
AT5G62940 194 / 3e-59
HIGH CAMBIAL ACTIVITY2, DNA BINDING WITH ONE FINGER 5.6, Dof-type zinc finger DNA-binding family protein (.1)
Potri.015G077100
118 / 1e-31
AT5G62940 191 / 4e-58
HIGH CAMBIAL ACTIVITY2, DNA BINDING WITH ONE FINGER 5.6, Dof-type zinc finger DNA-binding family protein (.1)
Potri.010G205400
116 / 1e-30
AT2G37590 137 / 7e-38
DNA binding with one finger 2.4 (.1)
Potri.005G134200
115 / 3e-30
AT5G60200 124 / 2e-33
TARGET OF MONOPTEROS 6 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10012619
172 / 2e-51
AT3G61850 201 / 2e-62
dof affecting germination 1, Dof-type zinc finger DNA-binding family protein (.1.2.3.4)
Lus10010113
167 / 9e-50
AT3G61850 194 / 5e-60
dof affecting germination 1, Dof-type zinc finger DNA-binding family protein (.1.2.3.4)
Lus10003207
150 / 1e-43
AT4G24060 159 / 9e-46
Dof-type zinc finger DNA-binding family protein (.1)
Lus10017313
144 / 4e-41
AT4G24060 171 / 8e-51
Dof-type zinc finger DNA-binding family protein (.1)
Lus10003208
140 / 4e-39
AT4G24060 167 / 3e-48
Dof-type zinc finger DNA-binding family protein (.1)
Lus10023032
133 / 6e-37
AT1G64620 171 / 1e-50
Dof-type zinc finger DNA-binding family protein (.1)
Lus10033218
131 / 3e-36
AT1G64620 169 / 7e-50
Dof-type zinc finger DNA-binding family protein (.1)
Lus10016566
115 / 6e-30
AT3G55370 160 / 2e-46
OBF-binding protein 3 (.1.2.3)
Lus10042114
107 / 1e-29
AT5G62940 149 / 7e-45
HIGH CAMBIAL ACTIVITY2, DNA BINDING WITH ONE FINGER 5.6, Dof-type zinc finger DNA-binding family protein (.1)
Lus10034652
111 / 5e-29
AT5G60850 162 / 4e-48
OBF binding protein 4 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF02701
zf-Dof
Dof domain, zinc finger
Representative CDS sequence
>Potri.014G100900.8 pacid=42763160 polypeptide=Potri.014G100900.8.p locus=Potri.014G100900 ID=Potri.014G100900.8.v4.1 annot-version=v4.1
ATGGACGAGATGGCATCTAATTCATGTGGAAGGCCTGGTGTAGAGAGAAAACCAAGGCCTCAAGAGCAATTGAACTGTCCAAGATGTAACTCAACCAATA
CCAAGTTTTGTTACTATAACAACTACAGTCTCACACAACCAAGATACTTTTGCAAGACTTGCAGAAGGTACTGGACTGAAGGAGGATCTCTTAGAAATGT
TCCTGTTGGAGGAGGTTCAAGAAAGAACAAGAGATCTACATCATCGTCAGTGGCAACATCTTCGTCTAAGCTCCCTGATCTCAACCCACCAAGCCTCTCA
CACTTCTCTTCTCAAAACCCTAAGAGTACCCATGAAGGTCAAGACCTTAATCTGGCATTCCCAGCTATGCAGGATAGTCAAGGTATATCTCATTATACTG
AAGTGCCCAAAACTGAAAATAGGAACAACAATCAACATAACTCTTCTTCTTCTCCCTATACTTCTTCTCCAATTTCAGCTATGGAGCTACTAAGGAGTGG
ATTTGCTTCTAGGGGTTTGAATACTTTTATCCCAACACCGATGCCGGATTCAAATACACTTTATTCTTCTGGTGGATTCCCTTTGCAAGAACTGAGACCA
ACTCTGAGCTTTCCTGCTGATGGGCTTGGAAGTAGATATGGAATTCAAGAAAATAGTGGGAGGCTTTTATTTCCATTTGGTGAGCTGAAACAGCTTTCAA
GCACAACGTCTGAAGTTGATCAAAACAAGGGACAGGGGGGTTCAAGTGGATACTGGAATGGAATGTTTGGCGGAGGATCATGGTAA
AA sequence
>Potri.014G100900.8 pacid=42763160 polypeptide=Potri.014G100900.8.p locus=Potri.014G100900 ID=Potri.014G100900.8.v4.1 annot-version=v4.1
MDEMASNSCGRPGVERKPRPQEQLNCPRCNSTNTKFCYYNNYSLTQPRYFCKTCRRYWTEGGSLRNVPVGGGSRKNKRSTSSSVATSSSKLPDLNPPSLS
HFSSQNPKSTHEGQDLNLAFPAMQDSQGISHYTEVPKTENRNNNQHNSSSSPYTSSPISAMELLRSGFASRGLNTFIPTPMPDSNTLYSSGGFPLQELRP
TLSFPADGLGSRYGIQENSGRLLFPFGELKQLSSTTSEVDQNKGQGGSSGYWNGMFGGGSW
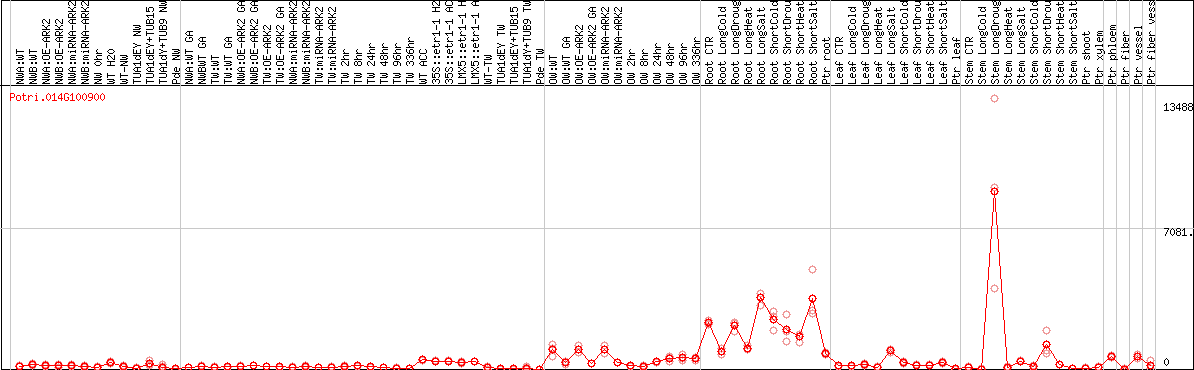
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.014G100900 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.