Potri.014G121701 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.014G121701.1 pacid=42762936 polypeptide=Potri.014G121701.1.p locus=Potri.014G121701 ID=Potri.014G121701.1.v4.1 annot-version=v4.1
ATGGCACCGTCGCGTAAACCGAGCAACGGTAGATCACCAATAGTAAATCCACAGCGCCAAATAACTGCCTTCTTCTCCAAGACGACGACTCCTTCTCCAT
CTCCTTCTCCCACGCTCTCTAAAAAACAAATCCCTAAATCCCATACTAAACCTAACCCTAACCCTAGCTCTCGAACCCAAAGCCCTAGTTCTAGCCCAAC
TACTCCTTCGCCAGTCCAATCTAAGCCTAAAAAACCCCTACTAGTTATTGGACAAACTCCTTCACCGTCTCCTTCAAAAGTTGGAGTGTATGGCAAAGAG
GCGGTGGAGAGGAGGGTTAGGGTTTACTGGCCCCTGGATAAGAGCTGGTACGAGGGTCTTGTCAAATCTTACGATGACGAGTCGAAGAAGCATTTGATAC
AATATGATGATTCTGAAGAGGAATTGTTGGATTTGAACAATGAGAAGATTGAGTGGGTTGAGCCATGTGTTAAGAAATTTAAGAGGTTGCGCAGGGGTTC
TTTAGGTTTTAGGAAGATTGTGCTTGAAGATGATGAGATGGAGAATGTCGAGGCCGATAATGGCGGTGCTGGTGGTGGTAGTGGTGGTGACGACTCGAGT
GATGAAGATTGGGGAAAGAATGCGGAGAAGGATGTTAGCGAGGAGGAGGACGTTGATTTGATGGATGAGGAAGAAGCGGATGATGGCAAGAAAGGAAAGA
GAGGTGGGAAGGATTCGAGGAAAAGGAAAGCGAGTGGAGAAGGTGGAAAGTTGGATTTGGGAAAGAAGGGAAAGAGTGGTGGGGATGCGAGTACGGGAGG
AGTCAAGGTTTCAGTTGTGGAGCCTGTAAAGAACAAGGAGAATGGAGTTTTTAATGGCTTTGAAAATGCCTTGATGACCTATGCATCAGAAAGATTTAGC
ACGCGTGAAGCTGAAAAGTTTCCATTTCTTGGACGAGAACGCAGGGATGCAAAGAGAAGGCGTCCTGGAGATGTAGGTTATGATCCCAGGACTCTATACT
TGCCCGCTGAGTTTGCAAAGAGTTTAACAGGTGGCCAGAGACAATGGTGGGAGTTTAAGTCAAAGCACATGGATAAGGTTTTGTTTTTCAAGATGGGCAA
ATTTTATGAGCTCTTTGAAATGGATGCCCATGTAGGAGCTAAAGAACTAGATTTGCAATACATGAAGGGAGAACAACCTCACTGTGGCTTTCCAGAGAAG
AACTTCTCTTTGAATGTGGAGAAATTGGCTAGAAAGGGTTATCGAGTTCTTGTTGTAGAGCAGACAGAAACACCTGAACAGTTAGAGCTTCGTCGCAAAG
AGAAAGGTTCTAAAGACAAGGTTGTAAAACGTGAAATATGTGCGGTGATTACAAAAGGAACACTAACTGAGGGGGAATTTCTTTCTGCAAATCCTGATGC
TTCTTATTTAATGGCACTGACTGAAAGTAGCCAGAGTTTGGCAAACCAAGGTCTTGAGCGGATTTTTGGTGTTTGTGTTGTTGATGTCACTACTAGCAGA
ATCATTCTTGGACAGTTTGGAGATGATGCTGAATGCAGTTCATTGTGCTGTCTATTATCTGAGCTGAGGCCGGTAGAAATTGTAAAACCTGCTAAAATGC
TTAGCTCCGAAACTGAGAGAGTAATGGTGAGACATACTAGAAATCCCTTGGTCAATGAGTTGGCTCCCCTCTCAGAATTCTGGGATGCGGAGAGAACTGT
TCAGGAAGTCAAAACTATCTACAAGCATATTGGTGATCTTTCAGCTTCTGGACCCTTAAATAAAACAGATTTGGATACAACTAACCTTAATGTTGGAGAA
TATAGGCCAAGCTGCCTGCCCAGCATTTTGTTAGAGTTTGTCAATAAAGGAGAAAATGGAAGCCTTGCTCTCTCAGCTCTTGGAGGCGCTCTCTATTATC
TAAAACAGGCATTTTTGGATGAGACATTGCTTAGATTTGCAAAATTTGAATCCCTTCCTTGCTCTGATTTCTGTGAGGTTGCTAAGAAACCATACATGAT
TCTCGATGCTGCTGCCCTGGAGAACCTTGAGATTTTTGAGAACAGCAGAAACGGAGACACTTCAGGGACGCTGTATGCGCAATTGAATCATTGTGTGACA
GCATTTGGAAAGAGGTTGCTGAAAACATGGCTAGCAAGACCATTATACCATTTGGAATCAATTAAAGATCGCCAGGATGCTGTAGCAGGTCTGCGGGGGG
TAAATCAACCTATGATGCTTGAATTTCAAAAGGTGTTGTCCAGGCTTCCAGACATTGAGCGGCTGCTTGCACGCATATTTTCTACCAGCGAAGCCAATGG
AAGAAATGCAAATAAAGTAGTTTTATATGAGGATGCAGCAAAGAAGCAGCTTCAGGAGTTTATATCCGCTCTACGTGGCTGTGAATTAGTGGCCCAAGCA
TGTTCTTCGCTTGCTGTCATTTTGGAAAATGTGGAGTCCGGACAGTTGCATCATTTATTAACACCTGGTAAAGGTCTTCCTGACATCCTTCCAATTCTTA
AACATTTTAAGAGTGCCTTTGATTGGGTAGAAGCCAACAATTCTGGACGTATAATACCTCATGAAGGTGTCGATGTAGAGTTTGACTCTGCTTGTGAAAA
AGTTAAAGAGGTTGAATCTAGTTTGGCTAGACACCTGAAGGAACAACAGAAATTACTAGGGGACAAATCAATTACTTATGTCACAGTTGGAAAAGAGGCA
TACCTGTTAGAAGTTCCTGAACATCTGCGTGGTAGTATTCCTCAAGATTATGAGCTGCGTTCATCCAAAAAGGGCTTCTACAGGTATTGGACTCCAAGTA
TTAAGAAGTTCTTAGGAGAGCTCTCACAGGCTGAATCTGAAAAGGAGTCGGCGTTGAAGAGCATTTTGCAGAGGTTAATTGTGTGTTTTTGCAAGTACCA
TGACAAGTGGAGACAGTTAGTTTCTGCAACTGCAGAACTGGATGTTTTGATCAGTCTGGCAATTGCTAGTGACTTTTATGAAGGACCTGCATGTTGTCCA
ACCATCGTAGGTTCATCATTGTCAAGTCAAGTGCCATGCCTTTCTGCAAAAAAATTAGGACACCCTGTTCTCAGGAGCGATTCTTTGGGAAAGGGTGCAT
TTGTCCCAAATGACATTAGTATTGGGGGTTCTGGTCGTGCCAGCTTTATCCTTCTCACTGGTCCTAACATGGGTGGAAAATCTACTCTTCTTCGGCAAGT
TTGTTTGGCTGTGATTCTGGCCCAGATTGGAGCTGATGTCCCTGCTGAAAGCTTTGAGTTGTCTCCTGTTGACCGCATCTTTGTCAGAATGGGTGCAAAA
GATCATATTATGGCAGGCCAGAGTACATTTTTGACAGAGCTCTCAGAAACTGCATTGATGCTGTCATCGGCAACCTGTAATTCATTGGTGGCGTTGGATG
AACTTGGACGTGGCACATCAACTTCAGATGGACAAGCAATAGCCGAATCAGTTCTTGAACATTTTGTCCACAAGGTACAGTGTCGAGGAATGTTTTCAAC
TCACTATCACCGATTAGCTGTAGATTATCAAAAAGATTCAAAGGTCTCACTGTACCATATGTCATGTCAAGTTGGAAATGGAGTTGGAGTGGAAGAAGTT
ACATTTCTTTATAGGTTGAGACCTGGTGCCTGCCCTAAGAGTTATGGTGTCAATGTTGCAAGACTGGCTGGACTTCCAGACTCGATACTTCATAATGCTG
CTGCCAAGTCTAGGGAATTTGAGGCTGTATATGGTCGACACAGAAAGGGATCTGAAGGAAAGTTGGCCATTCAAAGTTGTGACAAGATGGCAGTATTAAT
CCGAAGTTTGATCAATGCCACAACAAGCTTGAGTGGCCATAAGTCTGCTGGGATCGATATCAGCTCAGTGACCAAACTTCAGGATAAAGCTAGGATATTT
TTGCAGTAA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.014G121701.1 pacid=42762936 polypeptide=Potri.014G121701.1.p locus=Potri.014G121701 ID=Potri.014G121701.1.v4.1 annot-version=v4.1
MAPSRKPSNGRSPIVNPQRQITAFFSKTTTPSPSPSPTLSKKQIPKSHTKPNPNPSSRTQSPSSSPTTPSPVQSKPKKPLLVIGQTPSPSPSKVGVYGKE
AVERRVRVYWPLDKSWYEGLVKSYDDESKKHLIQYDDSEEELLDLNNEKIEWVEPCVKKFKRLRRGSLGFRKIVLEDDEMENVEADNGGAGGGSGGDDSS
DEDWGKNAEKDVSEEEDVDLMDEEEADDGKKGKRGGKDSRKRKASGEGGKLDLGKKGKSGGDASTGGVKVSVVEPVKNKENGVFNGFENALMTYASERFS
TREAEKFPFLGRERRDAKRRRPGDVGYDPRTLYLPAEFAKSLTGGQRQWWEFKSKHMDKVLFFKMGKFYELFEMDAHVGAKELDLQYMKGEQPHCGFPEK
NFSLNVEKLARKGYRVLVVEQTETPEQLELRRKEKGSKDKVVKREICAVITKGTLTEGEFLSANPDASYLMALTESSQSLANQGLERIFGVCVVDVTTSR
IILGQFGDDAECSSLCCLLSELRPVEIVKPAKMLSSETERVMVRHTRNPLVNELAPLSEFWDAERTVQEVKTIYKHIGDLSASGPLNKTDLDTTNLNVGE
YRPSCLPSILLEFVNKGENGSLALSALGGALYYLKQAFLDETLLRFAKFESLPCSDFCEVAKKPYMILDAAALENLEIFENSRNGDTSGTLYAQLNHCVT
AFGKRLLKTWLARPLYHLESIKDRQDAVAGLRGVNQPMMLEFQKVLSRLPDIERLLARIFSTSEANGRNANKVVLYEDAAKKQLQEFISALRGCELVAQA
CSSLAVILENVESGQLHHLLTPGKGLPDILPILKHFKSAFDWVEANNSGRIIPHEGVDVEFDSACEKVKEVESSLARHLKEQQKLLGDKSITYVTVGKEA
YLLEVPEHLRGSIPQDYELRSSKKGFYRYWTPSIKKFLGELSQAESEKESALKSILQRLIVCFCKYHDKWRQLVSATAELDVLISLAIASDFYEGPACCP
TIVGSSLSSQVPCLSAKKLGHPVLRSDSLGKGAFVPNDISIGGSGRASFILLTGPNMGGKSTLLRQVCLAVILAQIGADVPAESFELSPVDRIFVRMGAK
DHIMAGQSTFLTELSETALMLSSATCNSLVALDELGRGTSTSDGQAIAESVLEHFVHKVQCRGMFSTHYHRLAVDYQKDSKVSLYHMSCQVGNGVGVEEV
TFLYRLRPGACPKSYGVNVARLAGLPDSILHNAAAKSREFEAVYGRHRKGSEGKLAIQSCDKMAVLIRSLINATTSLSGHKSAGIDISSVTKLQDKARIF
LQ
|
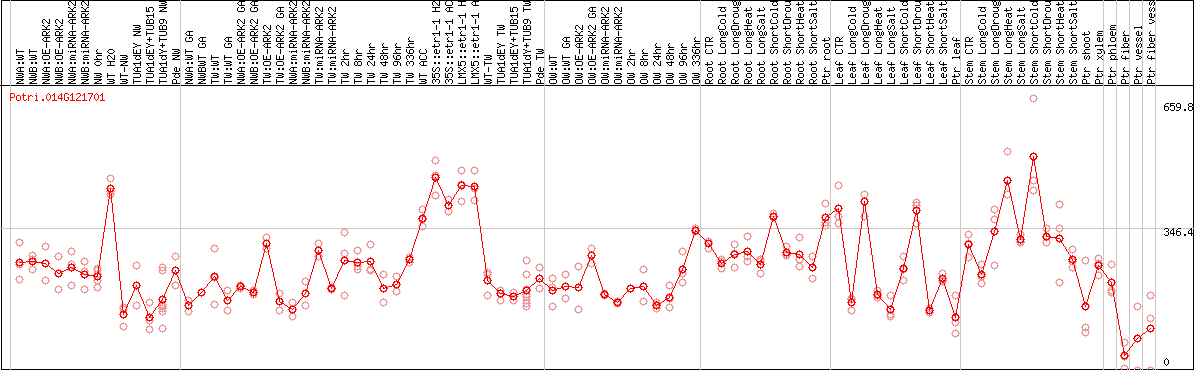
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.014G121701 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
