External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G03115 422 / 2e-149
Mitochondrial substrate carrier family protein (.1)
AT3G54110 171 / 5e-51
ATUCP1, UCP1, UCP2, UCP, ATPUMP1
ARABIDOPSIS THALIANA UNCOUPLING PROTEIN 1, ARABIDOPSIS THALIANA PLANT UNCOUPLING MITOCHONDRIAL PROTEIN 1, plant uncoupling mitochondrial protein 1 (.1)
AT1G14140 159 / 2e-46
Mitochondrial substrate carrier family protein (.1)
AT4G24570 158 / 6e-46
DIC2
dicarboxylate carrier 2 (.1)
AT5G58970 156 / 2e-45
ATUCP2
uncoupling protein 2 (.1.2)
AT5G09470 130 / 2e-35
DIC3
dicarboxylate carrier 3 (.1)
AT2G22500 129 / 5e-35
UCP5, ATPUMP5, DIC1
DICARBOXYLATE CARRIER 1, PLANT UNCOUPLING MITOCHONDRIAL PROTEIN 5, uncoupling protein 5 (.1)
AT5G19760 116 / 2e-30
Mitochondrial substrate carrier family protein (.1)
AT5G66380 106 / 1e-26
ATFOLT1
folate transporter 1 (.1)
AT3G53940 103 / 6e-25
Mitochondrial substrate carrier family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.006G095500
172 / 1e-51
AT3G54110 513 / 0.0
ARABIDOPSIS THALIANA UNCOUPLING PROTEIN 1, ARABIDOPSIS THALIANA PLANT UNCOUPLING MITOCHONDRIAL PROTEIN 1, plant uncoupling mitochondrial protein 1 (.1)
Potri.016G110100
166 / 5e-49
AT3G54110 483 / 5e-174
ARABIDOPSIS THALIANA UNCOUPLING PROTEIN 1, ARABIDOPSIS THALIANA PLANT UNCOUPLING MITOCHONDRIAL PROTEIN 1, plant uncoupling mitochondrial protein 1 (.1)
Potri.017G045100
164 / 4e-48
AT2G22500 451 / 6e-161
DICARBOXYLATE CARRIER 1, PLANT UNCOUPLING MITOCHONDRIAL PROTEIN 5, uncoupling protein 5 (.1)
Potri.017G047800
163 / 7e-48
AT4G24570 435 / 2e-154
dicarboxylate carrier 2 (.1)
Potri.017G045150
162 / 2e-47
AT2G22500 457 / 3e-163
DICARBOXYLATE CARRIER 1, PLANT UNCOUPLING MITOCHONDRIAL PROTEIN 5, uncoupling protein 5 (.1)
Potri.001G247800
161 / 2e-47
AT5G58970 456 / 3e-163
uncoupling protein 2 (.1.2)
Potri.007G113000
161 / 4e-47
AT4G24570 404 / 4e-142
dicarboxylate carrier 2 (.1)
Potri.010G165900
154 / 2e-44
AT1G14140 447 / 1e-159
Mitochondrial substrate carrier family protein (.1)
Potri.002G104400
145 / 7e-41
AT2G22500 427 / 1e-151
DICARBOXYLATE CARRIER 1, PLANT UNCOUPLING MITOCHONDRIAL PROTEIN 5, uncoupling protein 5 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10024769
452 / 2e-161
AT4G03115 401 / 3e-141
Mitochondrial substrate carrier family protein (.1)
Lus10035538
169 / 4e-50
AT3G54110 512 / 0.0
ARABIDOPSIS THALIANA UNCOUPLING PROTEIN 1, ARABIDOPSIS THALIANA PLANT UNCOUPLING MITOCHONDRIAL PROTEIN 1, plant uncoupling mitochondrial protein 1 (.1)
Lus10027755
163 / 6e-48
AT3G54110 515 / 0.0
ARABIDOPSIS THALIANA UNCOUPLING PROTEIN 1, ARABIDOPSIS THALIANA PLANT UNCOUPLING MITOCHONDRIAL PROTEIN 1, plant uncoupling mitochondrial protein 1 (.1)
Lus10021123
162 / 9e-48
AT3G54110 521 / 0.0
ARABIDOPSIS THALIANA UNCOUPLING PROTEIN 1, ARABIDOPSIS THALIANA PLANT UNCOUPLING MITOCHONDRIAL PROTEIN 1, plant uncoupling mitochondrial protein 1 (.1)
Lus10040709
159 / 2e-46
AT5G58970 512 / 0.0
uncoupling protein 2 (.1.2)
Lus10009777
157 / 5e-46
AT4G03115 140 / 4e-40
Mitochondrial substrate carrier family protein (.1)
Lus10017187
157 / 1e-45
AT3G54110 501 / 0.0
ARABIDOPSIS THALIANA UNCOUPLING PROTEIN 1, ARABIDOPSIS THALIANA PLANT UNCOUPLING MITOCHONDRIAL PROTEIN 1, plant uncoupling mitochondrial protein 1 (.1)
Lus10030443
152 / 5e-44
AT1G14140 421 / 3e-149
Mitochondrial substrate carrier family protein (.1)
Lus10016445
147 / 2e-41
AT5G58970 493 / 1e-177
uncoupling protein 2 (.1.2)
Lus10007883
116 / 3e-29
AT5G19760 536 / 0.0
Mitochondrial substrate carrier family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF00153
Mito_carr
Mitochondrial carrier protein
Representative CDS sequence
>Potri.014G135530.1 pacid=42764596 polypeptide=Potri.014G135530.1.p locus=Potri.014G135530 ID=Potri.014G135530.1.v4.1 annot-version=v4.1
ATGGATGGCAGCCATCCCCATTCGAGTCCCAGGAATTCAGATTCTTTTGTTGTTAAAGTGGCAGAGGAGAAGCAGAAATGGCCAGTCTCGTCTTCCCAAA
TGGTATCTCACTTCGCTACTAGTGGACTCTCCGTTGCTGTTGCCACCGCCATCACGCACCCCTTAGATGTGCTCAAGGTTCGACTACAAATGCAACTTGT
TGGCCGGAGAGGCCCCTTGACTGGAATGGGACAAGTGGCTGTTCAAGTATTGAAAAAGGAAGGGCCAAAGGCTTTGTATCTGGGGTTGATGCCTGCATTG
ATTAGGTCAGTTCTTTATGGAGGTCTACGTTTAGGCCTCTATGAACCCTCGAAGTATGCCTGTAATTTGGCTTTTGGGTCCACCAATATCTTACTTAAGA
TTGCATCAGGAGCATTTTCTGGTGCAGTTGCAACTGCACTTACTAATCCAGTCGAAGTTCTGAAGGTCCGGTTGCAGATGAATTCAAATCAGAGACAAGG
AGGACCAATGGCGGAAATGCGTACAATTGTTTCTGAAGAGGGAATAAGAGCTCTTTGGAAGGGGGTTGGCCCTGCTATGGTGAGAGCTGCTGCTTTGACT
GCATCACAACTGGCAACATATGATGAAACGAAGCAGGTTCTGATTAGGTGGACACCTCTGGATGAAGGATTTCATCTACATCTACTCTCGAGTACAGTTG
CTGGAACAGTGAGTACCCTTGTAACTGCACCTATGGACATGATTAAAACCCGCCTCATGCTGCAACGAGAATCTAAAACAGTTGGAAACTATAAAAATGG
ATTTCATTGTGCATATCAGGTTATGCTTAAAGAAGGCCCTAGGGCACTTTACAAGGGGGGCTTTGCAATATTTGCAAGATTGGGTCCGCAAACTACAATT
ACATTTATACTATGTGAGGAGCTGCGCAAGCTTGCCGGATTGGATGCCATCTAG
AA sequence
>Potri.014G135530.1 pacid=42764596 polypeptide=Potri.014G135530.1.p locus=Potri.014G135530 ID=Potri.014G135530.1.v4.1 annot-version=v4.1
MDGSHPHSSPRNSDSFVVKVAEEKQKWPVSSSQMVSHFATSGLSVAVATAITHPLDVLKVRLQMQLVGRRGPLTGMGQVAVQVLKKEGPKALYLGLMPAL
IRSVLYGGLRLGLYEPSKYACNLAFGSTNILLKIASGAFSGAVATALTNPVEVLKVRLQMNSNQRQGGPMAEMRTIVSEEGIRALWKGVGPAMVRAAALT
ASQLATYDETKQVLIRWTPLDEGFHLHLLSSTVAGTVSTLVTAPMDMIKTRLMLQRESKTVGNYKNGFHCAYQVMLKEGPRALYKGGFAIFARLGPQTTI
TFILCEELRKLAGLDAI
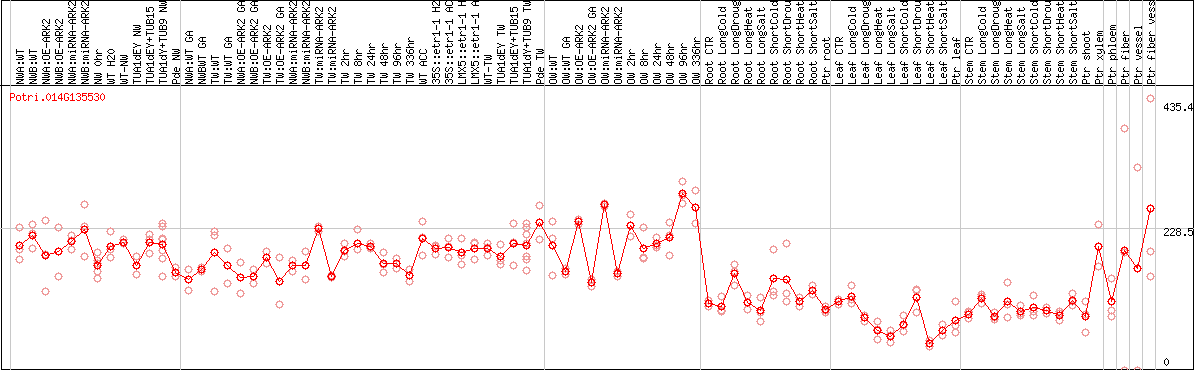
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.014G135530 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.