External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G26060 157 / 3e-48
EMB1345
embryo defective 1345, Transducin/WD40 repeat-like superfamily protein (.1.2)
AT4G32990 99 / 1e-25
Transducin/WD40 repeat-like superfamily protein (.1)
AT5G13480 56 / 8e-10
FY
Transducin/WD40 repeat-like superfamily protein (.1.2)
AT3G49660 54 / 5e-09
AtWDR5a
human WDR5 \(WD40 repeat\) homolog a, Transducin/WD40 repeat-like superfamily protein (.1)
AT5G67320 53 / 8e-09
HOS15
high expression of osmotically responsive genes 15, WD-40 repeat family protein (.1)
AT5G52820 53 / 8e-09
WD-40 repeat family protein / notchless protein, putative (.1)
AT3G44530 51 / 3e-08
HIRA
homolog of histone chaperone HIRA (.1.2)
AT2G43770 51 / 4e-08
Transducin/WD40 repeat-like superfamily protein (.1)
AT2G41500 51 / 5e-08
EMB2776, LIS
LACHESIS, WD-40 repeat family protein / small nuclear ribonucleoprotein Prp4p-related (.1)
AT5G25150 49 / 1e-07
TAF5
TBP-associated factor 5 (.1)
Paralogs
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10017045
169 / 8e-53
AT2G26060 486 / 5e-174
embryo defective 1345, Transducin/WD40 repeat-like superfamily protein (.1.2)
Lus10021365
169 / 1e-52
AT2G26060 482 / 2e-172
embryo defective 1345, Transducin/WD40 repeat-like superfamily protein (.1.2)
Lus10020214
56 / 1e-09
AT5G13480 738 / 0.0
Transducin/WD40 repeat-like superfamily protein (.1.2)
Lus10026837
56 / 1e-09
AT5G13480 759 / 0.0
Transducin/WD40 repeat-like superfamily protein (.1.2)
Lus10029493
54 / 3e-09
AT5G52820 786 / 0.0
WD-40 repeat family protein / notchless protein, putative (.1)
Lus10039597
54 / 3e-09
AT5G52820 792 / 0.0
WD-40 repeat family protein / notchless protein, putative (.1)
Lus10020585
54 / 7e-09
AT3G44530 1395 / 0.0
homolog of histone chaperone HIRA (.1.2)
Lus10024972
53 / 9e-09
AT5G67320 819 / 0.0
high expression of osmotically responsive genes 15, WD-40 repeat family protein (.1)
Lus10004893
53 / 1e-08
AT3G44530 1389 / 0.0
homolog of histone chaperone HIRA (.1.2)
Lus10024447
52 / 1e-08
AT5G25150 1006 / 0.0
TBP-associated factor 5 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0186
Beta_propeller
PF00400
WD40
WD domain, G-beta repeat
Representative CDS sequence
>Potri.014G136700.3 pacid=42763712 polypeptide=Potri.014G136700.3.p locus=Potri.014G136700 ID=Potri.014G136700.3.v4.1 annot-version=v4.1
ATGATCAGGCAGTTCTGGAAGAAACACACTAGAACTGTTCGGTCGTGTGCCTGGTCTCCTTCTGGGAAATTGTTGGCAACGCCCAGCTTCGATGACACAA
CTGCCACTTGGGAAATTAATGGCGGTGATTTCGAACGTGTAGCTGCTTTAGAGGGTCATGAAAATGAAGTGAAAAGCGTGCCTTGGAATGCATCTGGTTC
GCTCCTTCAGACGTGTAGTAGGGATAAAACTGTTTGGATTTGGGAAGTAATGCCTGGGAATGAATCCGAGTGTGTTTCGGTTTTTCCAAGGACACTTAAA
ATGGTTAAATGGCATCCAGCTATGGATGTTTTGTTTAGCTATGACAAGGTAAAATTGCTCATCTCTGCTCTTATTTCCACTGAAATAAGCTTCTTTAAAA
ATTCCATTTCTGCTTTCGAATAA
AA sequence
>Potri.014G136700.3 pacid=42763712 polypeptide=Potri.014G136700.3.p locus=Potri.014G136700 ID=Potri.014G136700.3.v4.1 annot-version=v4.1
MIRQFWKKHTRTVRSCAWSPSGKLLATPSFDDTTATWEINGGDFERVAALEGHENEVKSVPWNASGSLLQTCSRDKTVWIWEVMPGNESECVSVFPRTLK
MVKWHPAMDVLFSYDKVKLLISALISTEISFFKNSISAFE
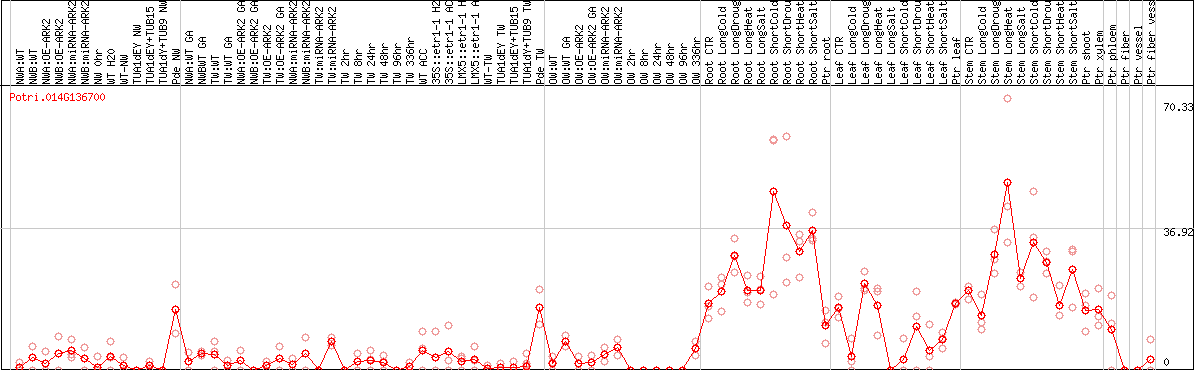
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.014G136700 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.