External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G19090 80 / 5e-17
Heavy metal transport/detoxification superfamily protein (.1.2.3)
AT5G27690 78 / 1e-16
Heavy metal transport/detoxification superfamily protein (.1)
AT3G06130 78 / 2e-16
Heavy metal transport/detoxification superfamily protein (.1.2)
AT1G23000 70 / 8e-14
Heavy metal transport/detoxification superfamily protein (.1)
AT1G56210 67 / 6e-13
Heavy metal transport/detoxification superfamily protein (.1)
AT5G66110 62 / 6e-12
HIPP27
heavy metal associated isoprenylated plant protein 27, Heavy metal transport/detoxification superfamily protein (.1)
AT5G37860 63 / 1e-11
Heavy metal transport/detoxification superfamily protein (.1)
AT3G05220 64 / 2e-11
Heavy metal transport/detoxification superfamily protein (.1.2)
AT4G35060 61 / 2e-11
HIPP25
heavy metal associated isoprenylated plant protein 25, Heavy metal transport/detoxification superfamily protein (.1)
AT1G71050 60 / 3e-11
HIPP20
heavy metal associated isoprenylated plant protein 20, Heavy metal transport/detoxification superfamily protein (.1)
Paralogs
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10041228
76 / 8e-16
AT3G06130 149 / 3e-41
Heavy metal transport/detoxification superfamily protein (.1.2)
Lus10031495
74 / 5e-15
AT5G27690 124 / 2e-32
Heavy metal transport/detoxification superfamily protein (.1)
Lus10015174
72 / 2e-14
AT5G27690 124 / 2e-32
Heavy metal transport/detoxification superfamily protein (.1)
Lus10029942
71 / 8e-14
AT5G19090 125 / 1e-31
Heavy metal transport/detoxification superfamily protein (.1.2.3)
Lus10032445
70 / 1e-13
AT1G23000 129 / 2e-34
Heavy metal transport/detoxification superfamily protein (.1)
Lus10004465
69 / 4e-13
AT5G19090 122 / 2e-30
Heavy metal transport/detoxification superfamily protein (.1.2.3)
Lus10013911
66 / 5e-13
AT1G06330 103 / 2e-28
Heavy metal transport/detoxification superfamily protein (.1)
Lus10001892
65 / 6e-13
AT1G06330 101 / 1e-27
Heavy metal transport/detoxification superfamily protein (.1)
Lus10031514
67 / 1e-12
AT5G19090 118 / 1e-28
Heavy metal transport/detoxification superfamily protein (.1.2.3)
Lus10014120
58 / 2e-10
AT4G38580 253 / 5e-88
HEAVY METAL ASSOCIATED ISOPRENYLATED PLANT PROTEIN 26, farnesylated protein 6 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF00403
HMA
Heavy-metal-associated domain
Representative CDS sequence
>Potri.014G140600.1 pacid=42763757 polypeptide=Potri.014G140600.1.p locus=Potri.014G140600 ID=Potri.014G140600.1.v4.1 annot-version=v4.1
ATGGCTAAGGAAGCAGAATTGAAGAAGATTGAGCTCAAGGTATCTGTCAACTGCTGTGATGGCTGCAAGAGGAAAGTTAAAAAGGCCTTGCAAGGTGTTG
AAGGTGTTCTGAAGACCGAAATCGATCCACAGCATCCCAAGGTGACGGTCCTGGGAAATGTGAATCCACAAATTTTAATCAAAAGACTCTTGAAAACTGG
AAAACAAGCAGAACTGTGGAGTAGTGGGAATCAGAATGCGGGAAAAGAAAAGAAAGAAGCGGATATGCTGGTTGAAAAAGAGAAAGACAAGTCAAAATCT
GAGTGTGAGCAAACAAAGTCTTCAGATTCATGTGTCAAGGTAACTGACAAGAACAGAGAGACTAAAAATGGTGGGGATGGAGGTGAGAATAAAGCTTCAA
AGGACTGTAATGAGACAGACGTCAGTGTTAAGTCTAGTAATCCTGAAGTGGTTAGAAGTGAAAATCCTGTCCCCCCACATCCAGAAGTCGGCAATTTCAG
GACTTACAATCAATATTGTTACAAGGTTGAGCCTTACGCAATTGCACTTCCCTTTTATGCCATTCCTTCATACACTGTTCCTCCTGTCAACCCCACAGGT
TATGGTCAGGAATACCTCCTTTACGAGAGGCCAGTATTTCAACCACCAGTCCAGGCGCCAACAGCAAGAGTTGAGGATTACTTCAGCGATGAAAATACTG
TGGGATGCCATGTGATGTGA
AA sequence
>Potri.014G140600.1 pacid=42763757 polypeptide=Potri.014G140600.1.p locus=Potri.014G140600 ID=Potri.014G140600.1.v4.1 annot-version=v4.1
MAKEAELKKIELKVSVNCCDGCKRKVKKALQGVEGVLKTEIDPQHPKVTVLGNVNPQILIKRLLKTGKQAELWSSGNQNAGKEKKEADMLVEKEKDKSKS
ECEQTKSSDSCVKVTDKNRETKNGGDGGENKASKDCNETDVSVKSSNPEVVRSENPVPPHPEVGNFRTYNQYCYKVEPYAIALPFYAIPSYTVPPVNPTG
YGQEYLLYERPVFQPPVQAPTARVEDYFSDENTVGCHVM
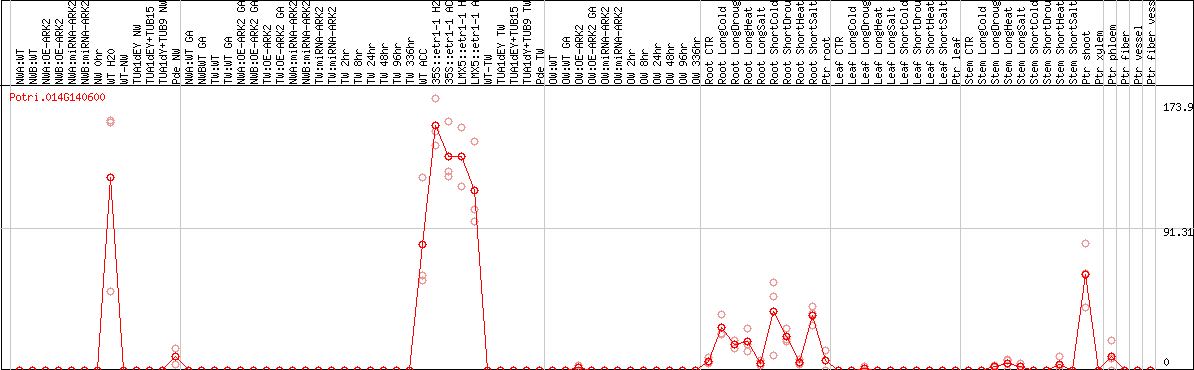
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.014G140600 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.