Potri.014G144600 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.014G144600.1 pacid=42764618 polypeptide=Potri.014G144600.1.p locus=Potri.014G144600 ID=Potri.014G144600.1.v4.1 annot-version=v4.1
ATGGTAGTAGTAGCAGCAGCCTGGTTTCTCTTTGTTGGTGGCTTAGCAGCAACATTTTCAGCTGAAGCTCTCTCTTTTAATGACTCAACTAGTGGTACTC
TCTTAAGTTTCAAGAATTCAGTTCTCGGGGACCCTTCAAATCTTCTCTCCTCCTGGAACCTGACTACCAATCCTGATTACTGCACCTGGTATGGTGTCAC
CTGTCAAAAACCTTCTAACACTACTACAGAAGTAGTAGTTATAGCTTTGAACTTCTCTGGCACTTCCACTACTCGCCTTTCTGGAACTCTGCCGGAGTCC
ATCCAAAACTTGCCCTACCTGCGAACTCTTGTCCTTTCTCACAACTGTTTTTCCGGTGAGATCCCAGCTGGTAGTATCGCCAAGCTGAGCTTCTTGGAGG
TTTTGGAGCTTCAAGGGAACAATTTTTCGGGCAAGATTCCTCAACAGATAAGCACTGACCTTCACTCTCTTCGCTTCCTTAATTTGTCTTTTAATTCTTT
CACTGGTGATATCCCTGCTACTTTGATTGGTTTTGGAAAGTTGAGAGTTATTGATTTGTCGAATAATAGGCTGACTGGTGGAATGCAACTTGTTAGTTTA
AGTAAGTGCTTGTTTTTAAGGCACTTGAAGCTTTCTAATAATCTCTTGGAAAATAATATTCCAAAGGATATTGGTCATTGCAAGAACTTGAGGACTTTGT
TGCTTGATGGGAACATTCTACAAGGTCCCATTCCAGCAGAGATAGGTCAGATTCCGGAACTTCGTGTTCTTGATGTTTCTACTAACAGTTTGACTCAGAC
CATCCCCAAGGAGTTGGGTTATTGCAGGAAACTGTCTGTTCTTGTATTGACTAATTCAAGTAATTTTGTTGGTGATAATGGGGGGACTGGTGGTAACTTG
GATGGTTTTAGATTAGAATTCAATGCTTTTGAAGGGGGTGTTCCTCAGGAGGTGTTGATGCTTCCGAGTTTGCAGATTTTGTGGGCGCCTAGGGCCAATC
TTGATGGGCGGTTGCCTGACAATTGGAGTGACTCGTGCTCGCTGAGAGTTTTGCATTTGGGGCAGAATTCTCTCAGAGGTGTTGTGCCAAAGGGGCTGGT
GATGTGCAAGAACCTTACTTTCTTGGATCTGAGTTCTAATTATTTGACTGGTGACCTGCCAATGCAGCTGCAAGTTCCTTGCATGATGTATTTCAATGTT
AGTCAGAATAACATATCCGGTGCTGTTCCAACTTTTGGAAAAGGCAGTTGTGATACTAGCATAATCTCTTATGGTCAAGATCCTAATTTTTTTTATGTGG
AGGATATACAGATTGCATACGCTAACATTCCTGTTTGGGGTTCCCATACCCTTTTGGGGTCCATGGCCGGTGCAGACTTCGTGATTGTCCATGATTTCAG
TTGGAATCATTTTGTCGGTTCATTGCCTTCGTTCTCGGTGGGAGAAGAATTCTTGGTGAGTAAAAATAGAACTTCATACAGATTGTTACTGAGTAGTAAC
GGGTTTACCGGGTCTCTTCCTGGTAAACTAGTTTCAAACTGTAATGATCTGCTGAGTTTCTCTGTTAACTTGAGTGCGAACCATATATCAGGTGAGATAC
CAGATATGCTTCTCAATTGTCTACCAATAAGGGAATTTGAAGCAGCAGATAATGAGATCAGTGGTTTTCTAGCTCCCAGTATTGGCAACTTGAGAATGCT
TCGATGTTTAGACTTGAGAAGAAACAGACTCTCTGGTTCTCTTCCTAATGAGTTGGGAAATTTGAGGTTTCTGAGATCAGTTCTTCTTGGAATGAACAAT
TTAACAGGAGAAATTCCATCTGAATTCGGTCAATTGTCCTCTCTTACAGTTTTGGATCTTTCTCATAATGCTGTAACAGGATCTATACCTATGAGTTTGA
CGAGCGCTAAGAACCTGGAGATTGTGTTACTCAATAACAATGATCTTTCTGGGGCGATACCTCCACCTTTCTCGAATATTTCCAGTCTCGTTGTCCTAAA
TGTTTCTTTCAATAACCTCTCTGGTCATATACCCCACCTTCAACACCCAATTGATTGTGATTGGTTTAGAGGGAATATTTTTTTGGATAAGTGCTTGGAT
CAATCCTCCAACACACCACCTGGAGAGGTTCAGCAGTCACATGGAGATAGAAAATGGCGCAATCACAGAAAGAAATCCTTTTTGATTGCAGTGGTGACTT
CTGCTTCAGTTGTACTCTGTGTATCATTGGTTGTAGTTTTGTTCAGTTTTTATGGCAAGAAAAAGTCTTGGAGACTCTCTATCTTGAGGGGAAAAGTAGT
TGTGACTTTTGCAGATGCTCCGGCTGAATTGACTTATGATAGTGTGGTCAGAGCTACTGGGAATTTCAGTATGAGAAACCTGATAGGTACAGGTGGCTTT
GGGTCAACTTACAAAGCTGAATTGGTCCCAGGTTACTTCATAGCCGTAAAGAGGCTCTCCATTGGCAGATTTCAAGGCATTCAACAGTTTGATGCAGAGA
TCAGAACATTGGGCAGAATTCGACACAAGAACCTTGTGACTCTGATTGGGTATTATGTGGCAGAGGCTGAAATGTTCTTAATTTACAACTATCTTTCTGG
TGGAAACCTTGAAACCTTCATACATGACAGGCCAGACACAAATGTACAGTGGCCAGTGATTCACAAGATAGCTCTTGATATTGCACAAGCTCTAGCCTAC
CTCCACTACTCATGCGCACCCCGCATTCTTCATCGTGACATCAAGCCTAGTAACATCCTGTTGGATGAGGAGCTTAACGCCTATCTGTCTGACTTTGGGT
TGGCCAAACTTTTAGAAGTTTCTCAGACCCATGCCACTACAGATGTAGCAGGCACATTTGGGTATGTTGCACCAGAATATGCAACCACGTGCAGGGTATC
TGACAAGTCAGATGTTTATAGCTTTGGGGTTGTGTTGTTGGAATTGATGTCAGGGAAGAAGTCTCTTGATCCATCATTTTCTGAGTATGGTAACGGTTTC
AATATTGTTGCTTGGGCCAAGTTGTTGATCAAGGAGAGACGCTCCTCAGAGCTCTTTGCGCCAGAACTGTGGGAAGCAGGACCTAATGAGAATTTGTTGG
GAATGTTAAAGCTTGCTTCAAGCTGCACAGTAGACTCACTTTCAGTGCGCCCATCCATGAAGCAGGTCCTTGAAAAACTGAAACAACTAAAACCATGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.014G144600.1 pacid=42764618 polypeptide=Potri.014G144600.1.p locus=Potri.014G144600 ID=Potri.014G144600.1.v4.1 annot-version=v4.1
MVVVAAAWFLFVGGLAATFSAEALSFNDSTSGTLLSFKNSVLGDPSNLLSSWNLTTNPDYCTWYGVTCQKPSNTTTEVVVIALNFSGTSTTRLSGTLPES
IQNLPYLRTLVLSHNCFSGEIPAGSIAKLSFLEVLELQGNNFSGKIPQQISTDLHSLRFLNLSFNSFTGDIPATLIGFGKLRVIDLSNNRLTGGMQLVSL
SKCLFLRHLKLSNNLLENNIPKDIGHCKNLRTLLLDGNILQGPIPAEIGQIPELRVLDVSTNSLTQTIPKELGYCRKLSVLVLTNSSNFVGDNGGTGGNL
DGFRLEFNAFEGGVPQEVLMLPSLQILWAPRANLDGRLPDNWSDSCSLRVLHLGQNSLRGVVPKGLVMCKNLTFLDLSSNYLTGDLPMQLQVPCMMYFNV
SQNNISGAVPTFGKGSCDTSIISYGQDPNFFYVEDIQIAYANIPVWGSHTLLGSMAGADFVIVHDFSWNHFVGSLPSFSVGEEFLVSKNRTSYRLLLSSN
GFTGSLPGKLVSNCNDLLSFSVNLSANHISGEIPDMLLNCLPIREFEAADNEISGFLAPSIGNLRMLRCLDLRRNRLSGSLPNELGNLRFLRSVLLGMNN
LTGEIPSEFGQLSSLTVLDLSHNAVTGSIPMSLTSAKNLEIVLLNNNDLSGAIPPPFSNISSLVVLNVSFNNLSGHIPHLQHPIDCDWFRGNIFLDKCLD
QSSNTPPGEVQQSHGDRKWRNHRKKSFLIAVVTSASVVLCVSLVVVLFSFYGKKKSWRLSILRGKVVVTFADAPAELTYDSVVRATGNFSMRNLIGTGGF
GSTYKAELVPGYFIAVKRLSIGRFQGIQQFDAEIRTLGRIRHKNLVTLIGYYVAEAEMFLIYNYLSGGNLETFIHDRPDTNVQWPVIHKIALDIAQALAY
LHYSCAPRILHRDIKPSNILLDEELNAYLSDFGLAKLLEVSQTHATTDVAGTFGYVAPEYATTCRVSDKSDVYSFGVVLLELMSGKKSLDPSFSEYGNGF
NIVAWAKLLIKERRSSELFAPELWEAGPNENLLGMLKLASSCTVDSLSVRPSMKQVLEKLKQLKP
|
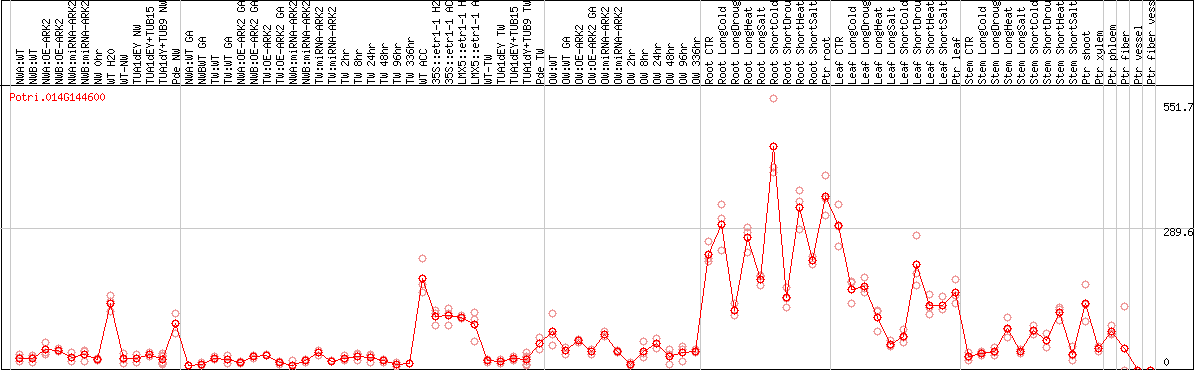
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.014G144600 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
