External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G46860 263 / 2e-88
SGR3, ATVAM3, ATSYP22, VAM3
SHOOT GRAVITROPISM 3, ARABIDOPSIS THALIANA VACUOLAR MORPHOLOGY 3, ARABIDOPSIS THALIANA SYNTAXIN OF PLANTS 22, Syntaxin/t-SNARE family protein (.1)
AT5G16830 218 / 2e-70
ATSYP21, ATPEP12, SYP21, PEP12P
syntaxin of plants 21 (.1)
AT4G17730 208 / 1e-66
ATSYP23, SYP23
syntaxin of plants 23 (.1.2)
AT1G32270 145 / 7e-41
ATSYP24
SYNTAXIN 24, syntaxin, putative (.1)
AT3G05710 51 / 3e-07
ATSYP43, SYP43
syntaxin of plants 43 (.1.2)
AT3G03800 50 / 4e-07
ATSYP131, SYP131
syntaxin of plants 131 (.1)
AT5G26980 50 / 6e-07
ATSYP41, ATTLG2A, SYP41
syntaxin of plants 41 (.1.2)
AT2G18260 49 / 9e-07
ATSYP112, SYP112
syntaxin of plants 112 (.1)
AT3G52400 49 / 2e-06
ATSYP122, SYP122
syntaxin of plants 122 (.1)
AT3G11820 48 / 3e-06
PEN1, AT-SYR1, ATSYR1, ATSYP121, SYP121
PENETRATION1, SYNTAXIN RELATED PROTEIN 1, syntaxin of plants 121 (.1.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.003G095300
254 / 8e-85
AT5G46860 368 / 2e-129
SHOOT GRAVITROPISM 3, ARABIDOPSIS THALIANA VACUOLAR MORPHOLOGY 3, ARABIDOPSIS THALIANA SYNTAXIN OF PLANTS 22, Syntaxin/t-SNARE family protein (.1)
Potri.001G138500
249 / 7e-83
AT5G46860 377 / 3e-133
SHOOT GRAVITROPISM 3, ARABIDOPSIS THALIANA VACUOLAR MORPHOLOGY 3, ARABIDOPSIS THALIANA SYNTAXIN OF PLANTS 22, Syntaxin/t-SNARE family protein (.1)
Potri.005G020800
55 / 2e-08
AT3G05710 462 / 1e-164
syntaxin of plants 43 (.1.2)
Potri.013G011100
54 / 4e-08
AT3G05710 457 / 7e-163
syntaxin of plants 43 (.1.2)
Potri.016G068600
50 / 4e-07
AT3G11820 421 / 5e-149
PENETRATION1, SYNTAXIN RELATED PROTEIN 1, syntaxin of plants 121 (.1.2)
Potri.007G023100
49 / 1e-06
AT2G18260 352 / 3e-122
syntaxin of plants 112 (.1)
Potri.006G202200
49 / 2e-06
AT3G11820 412 / 3e-145
PENETRATION1, SYNTAXIN RELATED PROTEIN 1, syntaxin of plants 121 (.1.2)
Potri.019G036700
47 / 4e-06
AT3G03800 391 / 1e-137
syntaxin of plants 131 (.1)
Potri.002G200400
47 / 9e-06
AT5G26980 397 / 1e-139
syntaxin of plants 41 (.1.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10030953
251 / 3e-83
AT5G46860 417 / 9e-149
SHOOT GRAVITROPISM 3, ARABIDOPSIS THALIANA VACUOLAR MORPHOLOGY 3, ARABIDOPSIS THALIANA SYNTAXIN OF PLANTS 22, Syntaxin/t-SNARE family protein (.1)
Lus10040094
251 / 3e-83
AT5G46860 419 / 1e-149
SHOOT GRAVITROPISM 3, ARABIDOPSIS THALIANA VACUOLAR MORPHOLOGY 3, ARABIDOPSIS THALIANA SYNTAXIN OF PLANTS 22, Syntaxin/t-SNARE family protein (.1)
Lus10000311
151 / 1e-45
AT5G46860 217 / 5e-72
SHOOT GRAVITROPISM 3, ARABIDOPSIS THALIANA VACUOLAR MORPHOLOGY 3, ARABIDOPSIS THALIANA SYNTAXIN OF PLANTS 22, Syntaxin/t-SNARE family protein (.1)
Lus10011064
154 / 4e-42
AT1G45616 366 / 2e-111
receptor like protein 6 (.1)
Lus10029864
52 / 1e-07
AT3G05710 510 / 0.0
syntaxin of plants 43 (.1.2)
Lus10013589
50 / 1e-06
AT3G11820 437 / 2e-155
PENETRATION1, SYNTAXIN RELATED PROTEIN 1, syntaxin of plants 121 (.1.2)
Lus10021263
50 / 1e-06
AT3G11820 436 / 3e-154
PENETRATION1, SYNTAXIN RELATED PROTEIN 1, syntaxin of plants 121 (.1.2)
Lus10016173
49 / 1e-06
AT3G11820 432 / 2e-152
PENETRATION1, SYNTAXIN RELATED PROTEIN 1, syntaxin of plants 121 (.1.2)
Lus10020683
49 / 2e-06
AT3G05710 361 / 7e-126
syntaxin of plants 43 (.1.2)
Lus10029373
49 / 2e-06
AT3G11820 441 / 4e-157
PENETRATION1, SYNTAXIN RELATED PROTEIN 1, syntaxin of plants 121 (.1.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF05739
SNARE
SNARE domain
CL0445
SNARE-fusion
PF14523
Syntaxin_2
Syntaxin-like protein
Representative CDS sequence
>Potri.014G145200.1 pacid=42763647 polypeptide=Potri.014G145200.1.p locus=Potri.014G145200 ID=Potri.014G145200.1.v4.1 annot-version=v4.1
ATGAGCTTCCAAGATTTTCAAAATGGCAAGAGACCTTCCTCCAGTTCTTCCACTTCCAGAAGCCCATCCCAAGCGGTGGCGGCCGGTATTTTCCAAATCA
ACACAGCCGTTGCCGGCTTCCGTCGTCTTGTCGATGCCATCGGAACCGACAAGGACACTCCTGAACACCGACATAAACTGCATAATTCAAGGCAACGGAT
TCTGCAGTTAGTTAAAGAGACTTCTGCTAAACTCAAGTCGTTGAGCGAACTCGACCACGATCCTGACATCAATCCGAGCAAGAAAATTGAAGATGCGAAG
CTAGCAAGAGATTTTCAAATCACATTGCAAGAGTTCCAGAAAGTCCAACAGCTTGCCTCTGAGCGCGAGTCCACTTACTCTCCTTCTCTACCTCCTCAAT
CTTCTTTGCCTCCTTCCTCTGGCTCTGGTGAATATGTGATCGCTAGTATGGATCAAGATAACCAACCTTTTTTAAGGGAACAGAGAAGGCAGGAGGTAAT
ACTTCTGGATAATGAAGTCGCATTTAATGAAGCCATAATTGAGGAAAGGGAACAGGGTATTAGAGACATAGAGGAACAAATTGGAGAAGCAAATGAAATA
TTCAAGGATCTTGCTGTTCTAGTTCATGACCAGGGTGTGGTTATTGATGATATTCACTCAAACATTGATTCTTCTGCTACTGCAACAACGCAAGCAAGAG
TTCAGCTATCCAAGGCTTCGAAAACCGTGAAATCTAAATGCTCATGGTGCTGGTGGCTACTGGGAATTGCTGTCGTTGTACTCGTTGTCCTTCTCCTCAT
CCTCATTTTATAG
AA sequence
>Potri.014G145200.1 pacid=42763647 polypeptide=Potri.014G145200.1.p locus=Potri.014G145200 ID=Potri.014G145200.1.v4.1 annot-version=v4.1
MSFQDFQNGKRPSSSSSTSRSPSQAVAAGIFQINTAVAGFRRLVDAIGTDKDTPEHRHKLHNSRQRILQLVKETSAKLKSLSELDHDPDINPSKKIEDAK
LARDFQITLQEFQKVQQLASERESTYSPSLPPQSSLPPSSGSGEYVIASMDQDNQPFLREQRRQEVILLDNEVAFNEAIIEEREQGIRDIEEQIGEANEI
FKDLAVLVHDQGVVIDDIHSNIDSSATATTQARVQLSKASKTVKSKCSWCWWLLGIAVVVLVVLLLILIL
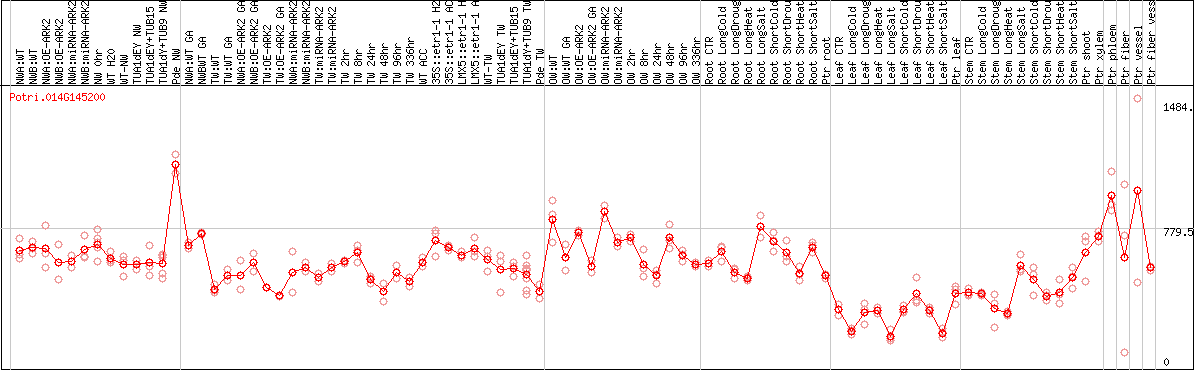
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.014G145200 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.