External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G18290 182 / 2e-57
SIP1B, SIP1;2
SMALL AND BASIC INTRINSIC PROTEIN 1B, Aquaporin-like superfamily protein (.1.2)
AT3G04090 176 / 7e-55
SIP1A, SIP1;1
small and basic intrinsic protein 1A (.1)
AT3G56950 100 / 2e-25
SIP2;1, SIP2
small and basic intrinsic protein 2;1 (.1.2)
AT3G61430 58 / 1e-09
ATPIP1, PIP1;1, PIP1A
PLASMA MEMBRANE INTRINSIC PROTEIN 1;1, ARABIDOPSIS THALIANA PLASMA MEMBRANE INTRINSIC PROTEIN 1, plasma membrane intrinsic protein 1A (.1.2)
AT2G45960 57 / 2e-09
PIP1;2, ATHH2, TMP-A, PIP1B
TRANSMEMBRANE PROTEIN A, NAMED PLASMA MEMBRANE INTRINSIC PROTEIN 1;2, plasma membrane intrinsic protein 1B (.1.2.3)
AT1G01620 56 / 3e-09
TMP-B, PIP1C, PIP1;3
PLASMA MEMBRANE INTRINSIC PROTEIN 1;3, plasma membrane intrinsic protein 1C (.1.2)
AT4G23400 56 / 4e-09
PIP1D, PIP1;5
plasma membrane intrinsic protein 1;5 (.1)
AT4G00430 56 / 4e-09
PIP1;4, TMP-C, PIP1E
TRANSMEMBRANE PROTEIN C, PLASMA MEMBRANE INTRINSIC PROTEIN 1E, plasma membrane intrinsic protein 1;4 (.1.2)
AT1G73190 55 / 7e-09
ALPHA-TIP, TIP3;1
ALPHA-TONOPLAST INTRINSIC PROTEIN, Aquaporin-like superfamily protein (.1)
AT2G39010 53 / 3e-08
PIP2E, PIP2;6
PLASMA MEMBRANE INTRINSIC PROTEIN 2;6, plasma membrane intrinsic protein 2E (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.002G227500
420 / 5e-151
AT5G18290 165 / 2e-50
SMALL AND BASIC INTRINSIC PROTEIN 1B, Aquaporin-like superfamily protein (.1.2)
Potri.019G030900
241 / 3e-80
AT3G04090 236 / 3e-78
small and basic intrinsic protein 1A (.1)
Potri.013G053400
229 / 6e-76
AT3G04090 224 / 6e-74
small and basic intrinsic protein 1A (.1)
Potri.016G024900
108 / 1e-28
AT3G56950 303 / 1e-104
small and basic intrinsic protein 2;1 (.1.2)
Potri.006G027200
105 / 1e-27
AT3G56950 264 / 1e-89
small and basic intrinsic protein 2;1 (.1.2)
Potri.009G027200
56 / 4e-09
AT4G01470 367 / 7e-130
tonoplast intrinsic protein 1;3 (.1)
Potri.001G235300
56 / 4e-09
AT4G01470 375 / 8e-133
tonoplast intrinsic protein 1;3 (.1)
Potri.016G089500
56 / 4e-09
AT2G37180 458 / 1e-164
RESPONSIVE TO DESICCATION 28, PLASMA MEMBRANE INTRINSIC PROTEIN 2C, PLASMA MEMBRANE INTRINSIC PROTEIN 2;3, Aquaporin-like superfamily protein (.1)
Potri.016G113300
56 / 6e-09
AT4G00430 524 / 0.0
TRANSMEMBRANE PROTEIN C, PLASMA MEMBRANE INTRINSIC PROTEIN 1E, plasma membrane intrinsic protein 1;4 (.1.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10035281
269 / 3e-91
AT3G04090 176 / 9e-55
small and basic intrinsic protein 1A (.1)
Lus10030046
261 / 3e-88
AT3G04090 172 / 2e-53
small and basic intrinsic protein 1A (.1)
Lus10021989
180 / 2e-51
AT5G65560 498 / 2e-157
Pentatricopeptide repeat (PPR) superfamily protein (.1)
Lus10038989
98 / 2e-24
AT3G56950 248 / 5e-83
small and basic intrinsic protein 2;1 (.1.2)
Lus10027275
97 / 3e-24
AT3G56950 249 / 2e-83
small and basic intrinsic protein 2;1 (.1.2)
Lus10035395
100 / 7e-24
AT5G05070 463 / 1e-158
DHHC-type zinc finger family protein (.1)
Lus10030997
91 / 2e-20
AT3G56930 332 / 1e-107
DHHC-type zinc finger family protein (.1.2)
Lus10005885
57 / 2e-09
AT4G01470 395 / 1e-140
tonoplast intrinsic protein 1;3 (.1)
Lus10040863
56 / 5e-09
AT4G01470 394 / 2e-140
tonoplast intrinsic protein 1;3 (.1)
Lus10022577
54 / 2e-08
AT4G00430 248 / 3e-83
TRANSMEMBRANE PROTEIN C, PLASMA MEMBRANE INTRINSIC PROTEIN 1E, plasma membrane intrinsic protein 1;4 (.1.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF00230
MIP
Major intrinsic protein
Representative CDS sequence
>Potri.014G154400.1 pacid=42763906 polypeptide=Potri.014G154400.1.p locus=Potri.014G154400 ID=Potri.014G154400.1.v4.1 annot-version=v4.1
ATGGGAGCAATCAAGGGAGCCATAGTAGATGGAATCTTGACATGCATGTGGGTATTCAGCGTTCCTTTGCTGGGTGTTTTCTCCTCCATCATAGCAACAT
ATGTGGGTGTTGAAGCCATGTCCATAGCAGGTCTTTTCATAACCATTAATGTAGCCGCTCTTTTTATGCTAACTTTCAGCTTGATAGGAGCTGCCTGTGG
TGGTGCCAGTTTCAACCCTGCTACAACCATAACTCTCTACACAGCAGGCCTAAAGCCTGATGCATCTCTCATGTCCATGGCTTTGCGTTTTCCTGTTCAG
GCAGCTGGTGGGGTTGCTGGTGCCATGGCAATTACGGAAGTGATGCCAAAACAGTACAGGTATGTGCTCAGAGGTGGTCCGTCGTTGAAGGTGGACTTGC
ATACAGGGGCAATCGCTGAGGGGGTGTTAACTTTTTTGATCTGTCTTGCTTTACATTTTGTTTTGCTCAAGGGTCCTAAAAACTTTGTGTTGAAGGTTTG
GTTGCTGGCTGTGGCCACTGTGGGACTGGTTATGGCTGGAGGTAAGTATACCGGCCCTTCAATGAATCCTGCCAATGCTTATGGGTGGGCTTATTTGAGC
AATAGGCACACCACTTGGGATTTCTTTTATGTCTATTGGATTTGCCCCTTTATAGGAGCAACTTTAGCTGCTTTGATTTCTAAATTCCTTTTCAAGGCAC
CACCAATCAAGGACAAGAAAGCCTGA
AA sequence
>Potri.014G154400.1 pacid=42763906 polypeptide=Potri.014G154400.1.p locus=Potri.014G154400 ID=Potri.014G154400.1.v4.1 annot-version=v4.1
MGAIKGAIVDGILTCMWVFSVPLLGVFSSIIATYVGVEAMSIAGLFITINVAALFMLTFSLIGAACGGASFNPATTITLYTAGLKPDASLMSMALRFPVQ
AAGGVAGAMAITEVMPKQYRYVLRGGPSLKVDLHTGAIAEGVLTFLICLALHFVLLKGPKNFVLKVWLLAVATVGLVMAGGKYTGPSMNPANAYGWAYLS
NRHTTWDFFYVYWICPFIGATLAALISKFLFKAPPIKDKKA
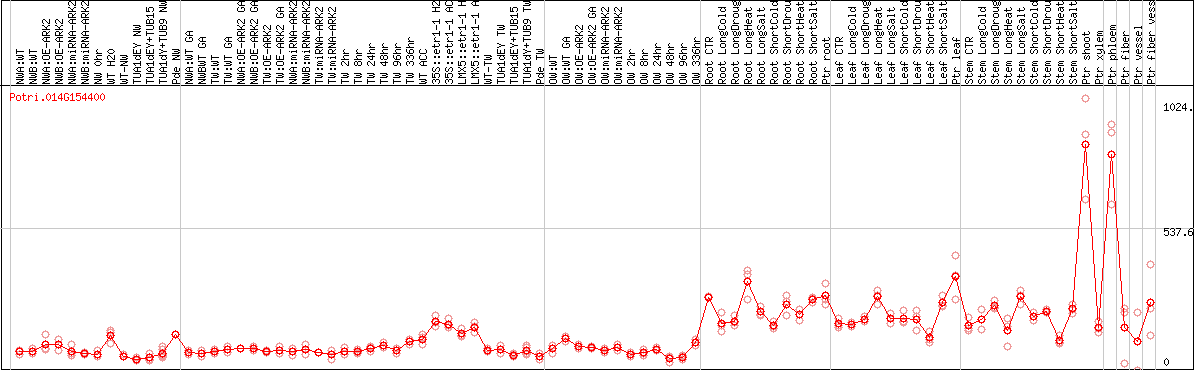
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.014G154400 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.