External link
Symbol
Pt-IBR5.1
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT2G04550 333 / 5e-116
DSPTP1E, IBR5
DUAL SPECIFICITY PROTEIN PHOSPHATASE 1E, indole-3-butyric acid response 5 (.1.3)
AT3G23610 86 / 5e-20
DSPTP1
dual specificity protein phosphatase 1 (.1.2.3)
AT3G06110 76 / 9e-17
DSPTP1B, MKP2, ATMKP2
DUAL-SPECIFICITY PROTEIN PHOSPHATASE 1B, ARABIDOPSIS MAPK PHOSPHATASE 2, MAPK phosphatase 2 (.1.2.3)
AT3G55270 66 / 7e-12
MKP1, ATMKP1
ARABIDOPSIS MITOGEN-ACTIVATED PROTEIN KINASE PHOSPHATASE 1, mitogen-activated protein kinase phosphatase 1 (.1)
AT5G23720 53 / 1e-07
PHS1
PROPYZAMIDE-HYPERSENSITIVE 1, dual specificity protein phosphatase family protein (.1.2.3)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.010G033000
83 / 2e-19
AT3G23610 249 / 4e-85
dual specificity protein phosphatase 1 (.1.2.3)
Potri.010G210900
67 / 2e-12
AT3G55270 715 / 0.0
ARABIDOPSIS MITOGEN-ACTIVATED PROTEIN KINASE PHOSPHATASE 1, mitogen-activated protein kinase phosphatase 1 (.1)
Potri.008G049900
67 / 2e-12
AT3G55270 710 / 0.0
ARABIDOPSIS MITOGEN-ACTIVATED PROTEIN KINASE PHOSPHATASE 1, mitogen-activated protein kinase phosphatase 1 (.1)
Potri.015G105000
54 / 4e-08
AT5G23720 1096 / 0.0
PROPYZAMIDE-HYPERSENSITIVE 1, dual specificity protein phosphatase family protein (.1.2.3)
Potri.012G105800
50 / 1e-06
AT5G23720 1006 / 0.0
PROPYZAMIDE-HYPERSENSITIVE 1, dual specificity protein phosphatase family protein (.1.2.3)
Potri.006G040300
49 / 2e-06
AT4G18593 148 / 5e-44
dual specificity protein phosphatase-related (.1)
Potri.016G035700
47 / 8e-06
AT4G18593 149 / 3e-44
dual specificity protein phosphatase-related (.1)
Potri.016G110700
45 / 4e-05
AT3G09100 914 / 0.0
mRNA capping enzyme family protein (.1.2)
Potri.006G096000
44 / 0.0001
AT3G09100 948 / 0.0
mRNA capping enzyme family protein (.1.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10002263
409 / 2e-145
AT2G04550 327 / 1e-113
DUAL SPECIFICITY PROTEIN PHOSPHATASE 1E, indole-3-butyric acid response 5 (.1.3)
Lus10000921
404 / 1e-143
AT2G04550 322 / 1e-111
DUAL SPECIFICITY PROTEIN PHOSPHATASE 1E, indole-3-butyric acid response 5 (.1.3)
Lus10021940
89 / 3e-21
AT3G23610 248 / 1e-84
dual specificity protein phosphatase 1 (.1.2.3)
Lus10041227
81 / 5e-18
AT3G23610 237 / 4e-80
dual specificity protein phosphatase 1 (.1.2.3)
Lus10034033
64 / 1e-12
AT3G23610 168 / 5e-54
dual specificity protein phosphatase 1 (.1.2.3)
Lus10003291
68 / 2e-12
AT3G55270 800 / 0.0
ARABIDOPSIS MITOGEN-ACTIVATED PROTEIN KINASE PHOSPHATASE 1, mitogen-activated protein kinase phosphatase 1 (.1)
Lus10006229
66 / 7e-12
AT3G23610 512 / 1e-171
dual specificity protein phosphatase 1 (.1.2.3)
Lus10030324
66 / 1e-11
AT3G55270 806 / 0.0
ARABIDOPSIS MITOGEN-ACTIVATED PROTEIN KINASE PHOSPHATASE 1, mitogen-activated protein kinase phosphatase 1 (.1)
Lus10036877
65 / 1e-11
AT3G23610 511 / 3e-171
dual specificity protein phosphatase 1 (.1.2.3)
Lus10032545
61 / 3e-10
AT5G23720 1120 / 0.0
PROPYZAMIDE-HYPERSENSITIVE 1, dual specificity protein phosphatase family protein (.1.2.3)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0031
Phosphatase
PF00782
DSPc
Dual specificity phosphatase, catalytic domain
Representative CDS sequence
>Potri.014G160500.1 pacid=42762641 polypeptide=Potri.014G160500.1.p locus=Potri.014G160500 ID=Potri.014G160500.1.v4.1 annot-version=v4.1
ATGATGATGATGAGGAAGAGGGAGAGAGAAAACCCGTGCGGGGTATGCGGTCACTACCACAAGTACGAAGAGGGTGAGGTTTGCGGGATTTGCGGCCACC
GTATGCCTGAATCCACCGCCGATAAGTCTCCCTCTGTTCACCTCAGCGCCTTTCCTTCTCAGATCCTCCCCGACTTCCTTTTCTTAGGCAGCTACGACAA
CGCCTCTCGATCCGAGCTCCTCAAAACCCAAGGAATTACTCGTGTGCTCAATACAGTTCCTGCGTGTCAAAATCTCTACAAGAATTCATTCACCTATCAC
TGCCTCCAAGATGACAAGATCTTACAATTTGACGATGCCATACAGTTTTTAGAGCAATGTGAGAGGGACAAGGCTCGCGTTCTTGTCCATTGCATGTCAG
GAAAAAATAGGTCTCCAGCTATTGTAATGGCTTACTTGATGAAGTCCAGAGGATGGAGACTTGCTCAATGTTACCAGTGGGTGAAAGAGCGGAGACCATC
CGTTGATCTAACTCAAGCCGTGCACCAGCAGTTGCATGAGTATGAACAGAAGCTTTTCGGGTCAAATGATAATAGCAACCCTGCCCTGCCAGTCTTCCTT
CCTGTAGGTGCGCCATCTTTTAGCTTTGGCTTCCCAAAGGCTAATGATCCAGTTCCTGTCCAAGCATTCATTGGTGTTGGTGCTCCCTCTATTTTCACAC
GTCCTCTAGAAGTTCCTCCACAAGAGTTCCAGTTTGGAGCTGGTCATCCCCAGAACATATCAGAAAGCTCCCTTGGCACCAATTTGCCGAACCCGAATGG
TGGGGATGTTTCAATGGATTCATGA
AA sequence
>Potri.014G160500.1 pacid=42762641 polypeptide=Potri.014G160500.1.p locus=Potri.014G160500 ID=Potri.014G160500.1.v4.1 annot-version=v4.1
MMMMRKRERENPCGVCGHYHKYEEGEVCGICGHRMPESTADKSPSVHLSAFPSQILPDFLFLGSYDNASRSELLKTQGITRVLNTVPACQNLYKNSFTYH
CLQDDKILQFDDAIQFLEQCERDKARVLVHCMSGKNRSPAIVMAYLMKSRGWRLAQCYQWVKERRPSVDLTQAVHQQLHEYEQKLFGSNDNSNPALPVFL
PVGAPSFSFGFPKANDPVPVQAFIGVGAPSIFTRPLEVPPQEFQFGAGHPQNISESSLGTNLPNPNGGDVSMDS
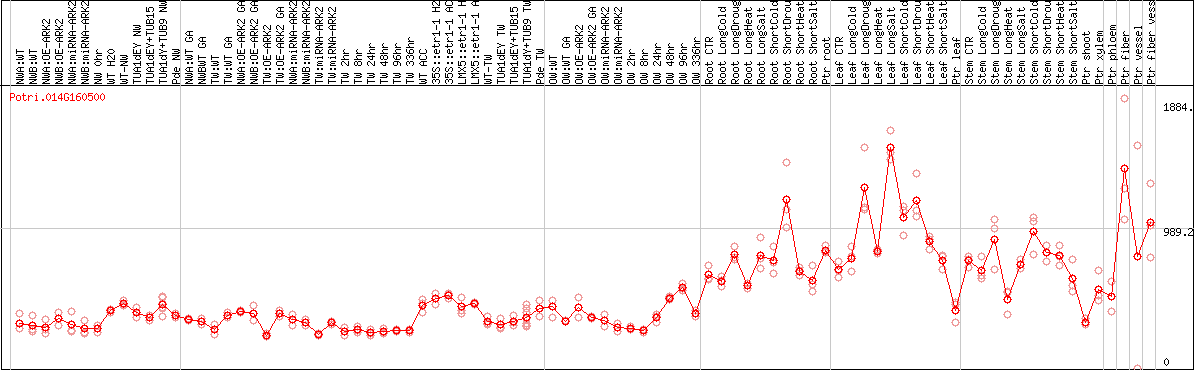
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.014G160500 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.