External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G07900 143 / 2e-43
AS2
LBD1
LOB domain-containing protein 1 (.1)
AT2G28500 143 / 3e-43
AS2
LBD11
LOB domain-containing protein 11 (.1)
AT2G30130 127 / 4e-37
AS2
PCK1, LBD12, ASL5
PEACOCK 1, Lateral organ boundaries (LOB) domain family protein (.1)
AT5G63090 118 / 7e-34
AS2
LOBB, LOB
Lateral organ boundaries (LOB) domain family protein (.1), Lateral organ boundaries (LOB) domain family protein (.2), Lateral organ boundaries (LOB) domain family protein (.3), Lateral organ boundaries (LOB) domain family protein (.4)
AT1G16530 117 / 7e-34
AS2
ASL9, LBD3
LOB DOMAIN-CONTAINING PROTEIN 3, ASYMMETRIC LEAVES 2-like 9 (.1)
AT3G27650 117 / 9e-34
AS2
LBD25
LOB domain-containing protein 25 (.1)
AT1G31320 116 / 2e-33
AS2
LBD4
LOB domain-containing protein 4 (.1)
AT2G30340 118 / 6e-33
AS2
LBD13
LOB domain-containing protein 13 (.1)
AT5G66870 116 / 8e-32
AS2
LBD36, ASL1
LATERAL ORGAN BOUNDARIES DOMAIN GENE 36, ASYMMETRIC LEAVES 2-like 1 (.1)
AT1G65620 111 / 3e-31
AS2
AS2
ASYMMETRIC LEAVES 2, Lateral organ boundaries (LOB) domain family protein (.1), Lateral organ boundaries (LOB) domain family protein (.2), Lateral organ boundaries (LOB) domain family protein (.3), Lateral organ boundaries (LOB) domain family protein (.4)
Paralogs
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10031769
139 / 2e-42
AT1G31320 193 / 8e-64
LOB domain-containing protein 4 (.1)
Lus10031191
136 / 3e-41
AT1G31320 199 / 2e-66
LOB domain-containing protein 4 (.1)
Lus10040373
137 / 1e-40
AT2G28500 211 / 7e-69
LOB domain-containing protein 11 (.1)
Lus10031285
133 / 8e-40
AT2G30130 226 / 3e-76
PEACOCK 1, Lateral organ boundaries (LOB) domain family protein (.1)
Lus10011645
132 / 6e-39
AT2G28500 205 / 4e-67
LOB domain-containing protein 11 (.1)
Lus10036130
132 / 1e-38
AT2G28500 207 / 1e-67
LOB domain-containing protein 11 (.1)
Lus10031855
128 / 4e-38
AT2G30130 220 / 4e-74
PEACOCK 1, Lateral organ boundaries (LOB) domain family protein (.1)
Lus10040638
125 / 1e-36
AT1G31320 244 / 1e-83
LOB domain-containing protein 4 (.1)
Lus10007525
124 / 2e-36
AT2G30130 204 / 1e-67
PEACOCK 1, Lateral organ boundaries (LOB) domain family protein (.1)
Lus10018275
124 / 4e-36
AT1G31320 243 / 2e-83
LOB domain-containing protein 4 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF03195
LOB
Lateral organ boundaries (LOB) domain
Representative CDS sequence
>Potri.014G167100.2 pacid=42762574 polypeptide=Potri.014G167100.2.p locus=Potri.014G167100 ID=Potri.014G167100.2.v4.1 annot-version=v4.1
ATGAGTTCTCAAGCACAGACACGAACCAGGGCTCACCAACCATGTGCTGCTTGTAGAATGCTACGAAGGAGATGCGACAGCAATTGCATGCTCGCCCCAT
ATTTTCCAGGTGATGAAGCTGAGAAATTTTTTGGGGTTCATAAGGTGTTTGGTGCCAGCAATGTTATCAGAATGATTCAGATGGTGGACGAATCAAAAAA
AGAAGATGCTGTTAAAGCAATCATCTATGAAGCAACATCAAGGCTCAGAGACCCTGTATATGGAAGTGCTGGAACAATATTCCACCTGCAAAGGATGGTT
GAAGAACTCAAGATGCAGGTGGAGTCCATGAGAGCTCAAGTAGTTCAGCTGCAAGAACAAAGAAACCAGTTGTTGGGCATTCTGATGAACGTCCGCCACC
TTGATTCCACCTCCTCGGTTCATGACAGCAAGTTTGATGGTGGCAATTTCATGTTGGATGAAGATTCAATGGCTTATGATCCCATGACATTTTTCACAGA
AAATGACTGGACTTTCTGA
AA sequence
>Potri.014G167100.2 pacid=42762574 polypeptide=Potri.014G167100.2.p locus=Potri.014G167100 ID=Potri.014G167100.2.v4.1 annot-version=v4.1
MSSQAQTRTRAHQPCAACRMLRRRCDSNCMLAPYFPGDEAEKFFGVHKVFGASNVIRMIQMVDESKKEDAVKAIIYEATSRLRDPVYGSAGTIFHLQRMV
EELKMQVESMRAQVVQLQEQRNQLLGILMNVRHLDSTSSVHDSKFDGGNFMLDEDSMAYDPMTFFTENDWTF
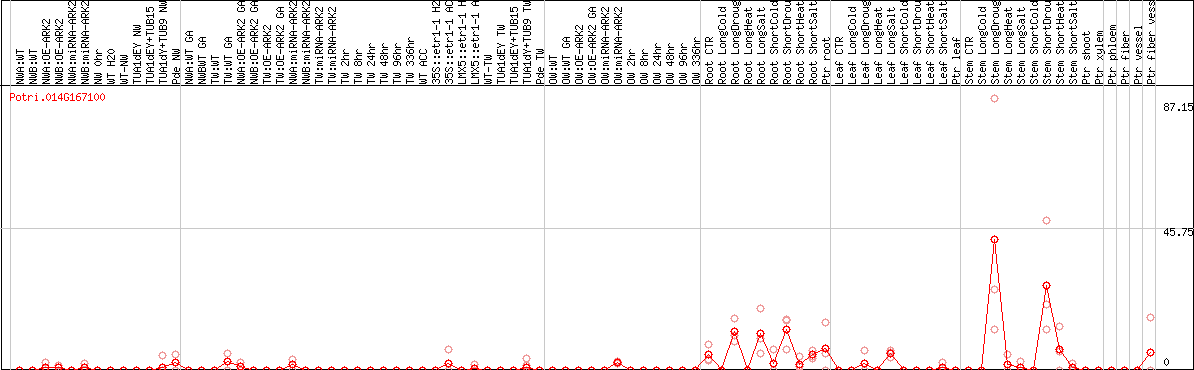
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.014G167100 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.