External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G12850 178 / 7e-57
FAR1_related
Far-red impaired responsive (FAR1) family protein (.1), Far-red impaired responsive (FAR1) family protein (.2), Far-red impaired responsive (FAR1) family protein (.3)
AT2G43280 152 / 2e-46
FAR1_related
Far-red impaired responsive (FAR1) family protein (.1)
AT3G07500 120 / 5e-34
FAR1_related
Far-red impaired responsive (FAR1) family protein (.1)
AT3G59470 115 / 1e-31
FAR1_related
Far-red impaired responsive (FAR1) family protein (.1), Far-red impaired responsive (FAR1) family protein (.2)
AT4G38180 97 / 3e-23
FAR1_related
FRS5
FAR1-related sequence 5 (.1)
AT5G18960 86 / 2e-19
FAR1_related
FRS12
FAR1-related sequence 12 (.1)
AT3G06250 85 / 6e-19
FAR1_related
FRS7
FAR1-related sequence 7 (.1)
AT2G32250 77 / 2e-16
FAR1_related
FRS2
FAR1-related sequence 2 (.1.2.3.4)
AT4G15090 70 / 7e-14
FAR1_related
FAR1
FAR-RED IMPAIRED RESPONSE 1, FRS (FAR1 Related Sequences) transcription factor family (.1)
AT1G76320 67 / 7e-13
FAR1_related
FRS4
FAR1-related sequence 4 (.1.2)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.002G239400
271 / 1e-93
AT4G12850 207 / 1e-68
Far-red impaired responsive (FAR1) family protein (.1), Far-red impaired responsive (FAR1) family protein (.2), Far-red impaired responsive (FAR1) family protein (.3)
Potri.017G029400
172 / 3e-54
AT2G43280 319 / 4e-112
Far-red impaired responsive (FAR1) family protein (.1)
Potri.014G176500
131 / 5e-38
AT3G07500 275 / 2e-94
Far-red impaired responsive (FAR1) family protein (.1)
Potri.007G129000
119 / 3e-34
AT2G43280 133 / 9e-40
Far-red impaired responsive (FAR1) family protein (.1)
Potri.007G129500
109 / 3e-29
AT3G59470 260 / 9e-88
Far-red impaired responsive (FAR1) family protein (.1), Far-red impaired responsive (FAR1) family protein (.2)
Potri.007G129101
103 / 3e-28
AT2G43280 142 / 6e-44
Far-red impaired responsive (FAR1) family protein (.1)
Potri.017G029100
105 / 7e-28
AT3G59470 323 / 1e-112
Far-red impaired responsive (FAR1) family protein (.1), Far-red impaired responsive (FAR1) family protein (.2)
Potri.002G239300
97 / 1e-24
AT2G43280 130 / 2e-37
Far-red impaired responsive (FAR1) family protein (.1)
Potri.008G076800
96 / 6e-24
AT3G59470 156 / 1e-46
Far-red impaired responsive (FAR1) family protein (.1), Far-red impaired responsive (FAR1) family protein (.2)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10026854
76 / 4e-16
AT3G59470 121 / 1e-32
Far-red impaired responsive (FAR1) family protein (.1), Far-red impaired responsive (FAR1) family protein (.2)
Lus10020226
73 / 2e-15
AT3G59470 120 / 7e-33
Far-red impaired responsive (FAR1) family protein (.1), Far-red impaired responsive (FAR1) family protein (.2)
Lus10026732
59 / 2e-11
AT3G07500 88 / 2e-22
Far-red impaired responsive (FAR1) family protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0274
WRKY-GCM1
PF03101
FAR1
FAR1 DNA-binding domain
Representative CDS sequence
>Potri.014G176600.1 pacid=42764232 polypeptide=Potri.014G176600.1.p locus=Potri.014G176600 ID=Potri.014G176600.1.v4.1 annot-version=v4.1
ATGGATTTGGACAAGGGTGTCGTTGACACTTCTACTTGTTGTGTCGAAGAAGAAGAAACTGTTGGAGATGGAATTGCAGAACCGGATGTTGGTATGGAAT
TTGAATCTGAAGATGCTGCCAGGAGATTCTACACAGAGTATGCCAGACGGGTAGGATTTGCTGTGCGTGTTATGCAACGTCGACGTTCGGGGATTGATGG
AAGAACTCTTGCTCGTCGTCTTGGATGTAACAAACAGGGTTTTTCTCCTAATCACAGGACTGATGTTGGCCCGGACAAGAAGCCTAGGCCCGGTGCACGA
GATGGTTGCAAAGCTACAATCTTGGTTAAAATGGAAAAGTCTGGAAAATGGGTGGTTACTAGATTTGAAAAGGATCATAATCATCCTTTAGTTGTCACCA
CCTCTGGATTCAGTTCATCAGGTGGCAAGGATAAGAAAATTGAAGAGCTTACTCAAGAGTTAGAGCATCAGGAGCAATTATGTGCAACTTATCGTGAAAA
ATTACTCAGTTTTATGAACAACGTTGTGGAGGAAGCAGAAGAACTGGCTTCAAAAATTCAAGTTATTGTGGACAGTGTCAGGAAAGTTGAATCGGAAGTG
CAAAAGTTTTCCCGTCATGGATAG
AA sequence
>Potri.014G176600.1 pacid=42764232 polypeptide=Potri.014G176600.1.p locus=Potri.014G176600 ID=Potri.014G176600.1.v4.1 annot-version=v4.1
MDLDKGVVDTSTCCVEEEETVGDGIAEPDVGMEFESEDAARRFYTEYARRVGFAVRVMQRRRSGIDGRTLARRLGCNKQGFSPNHRTDVGPDKKPRPGAR
DGCKATILVKMEKSGKWVVTRFEKDHNHPLVVTTSGFSSSGGKDKKIEELTQELEHQEQLCATYREKLLSFMNNVVEEAEELASKIQVIVDSVRKVESEV
QKFSRHG
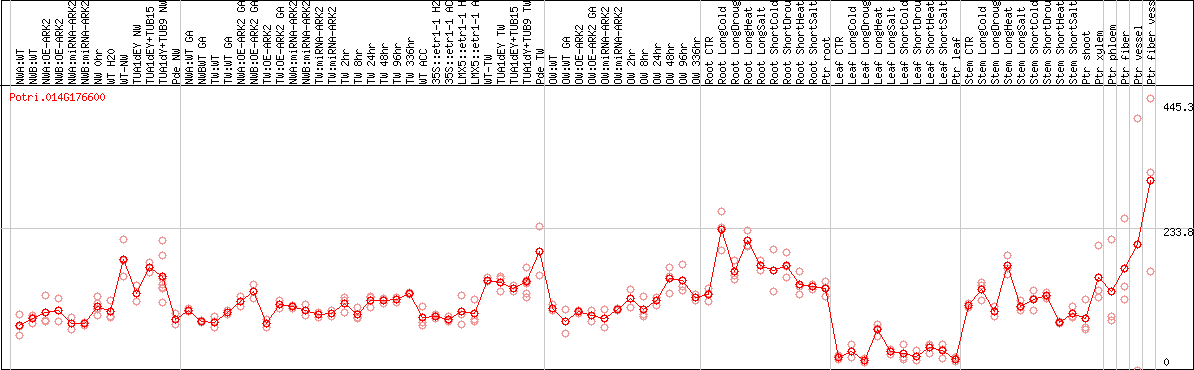
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.014G176600 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.