MRP9.1 (Potri.014G180100) [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | MRP9.1 | |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.014G180100.1 pacid=42762680 polypeptide=Potri.014G180100.1.p locus=Potri.014G180100 ID=Potri.014G180100.1.v4.1 annot-version=v4.1
ATGTTAGATTTAGGTGCTGCTGCTGCTGCTGCTGCTGCTGCTGCTGCAAATCTTAAGCTATTAATTCGGATGGATTGGCCGCAGCTACAGTCACCCTGTT
TGCGGGAGCACATAACCATAGGTGTACAGCTTGGGTTCCTCGGAATCTTGTTACTTCATCTTCTACGAAAATGCGCAGACCTAGCTTTCAATGGTGGAAC
AAAGACCACAGATCAAGGCAAGGAGAATTACCACATTGGATTGAAGTTCAGCAACTCTTACAAAGCGAGCATGGTTTGTTCGACTTGTCTGCTGGGAGTC
CATATCTCAATGCTACTGGTGCTGTTAAATGGTCAAGAGACCAGTTGCAACTCCATAGTACGAGTCTTTTCAGCTGAGGTCTTGCAAATGATATCATGGG
CAATTACACTGGTTGCAGTCTTTAGGATTTTTCCGAGTAGAAGGTATGTCAAGTTCCCTTGGATTATAAGAGCATGGTGGCTTTGCAGCTTCATGCTGTC
CATTGTCTGCACATCTTTAGATATCAACTTCAAAATTACAAACCATGGCCATCTTAGGCTACGAGATTATGCAGAGCTCTTTGCTCTCCTCCCATCCACA
TTCCTGCTAGCTATTTCATTTCGAGGGAAGACAGGCATAGTCTTTAATGCATTTAATGGCGTCACCGACCCACTTCTCCATGAAAAGAGTGATAAAGATT
CAGACACCAAGAGAGAATCACCATATGGAAAAGCTACACTCCTCCAGCTGATTACCTTCTCTTGGCTCACCCCGTTGTTTGCTGTTGGTTACAAGAAACC
TCTTGAACAGGATGAAATCCCAGATGTTTATATCAAGGATTCTGCTGGATTCCTCTCCTCCTCCTTTGATGAGAATCTAAATCAAGTTAAGGAGAAAGAT
AGAACCGCAAACCCCTCTATCTACAAGGCAATCTTTTTGTTCATCAGAAAGAAAGCAGCAATTAATGCATTATTCGCAGTTACCAGTGCAGCAGCATCAT
ATGTTGGTCCCTACCTCATTGATGACTTCGTGAATTTCCTAACCGAGAAGAAAACCAGGAGCTTACAGAGTGGCTACCTTCTTGCATTAGGGTTTCTAGG
TGCAAAGACAGTGGAGACAATAGCGCAGAGACAGTGGATATTTGGAGCCCGACAGCTCGGCCTTCGTCTTAGAGCCTCCCTGATATCTCACATTTACAAG
AAGGGCCTACTCTTGTCAAGCCAATCCCGCCAAAGCCACACCAGTGGAGAGATAATCAACTACATGAGTGTAGATATCCAACGAATTACAGATTTCATCT
GGTACTTGAACTATATATGGATGTTGCCTGTACAAATTACCTTAGCAATCTACATTCTACATACAACTCTAGGTTTGGGGTCAATGGCAGCATTAACTGC
AACTTTAGCAGTGATGGCTTGCAACATACCCATAACAAGATTCCAGAAAAGATACCAGACTAAGATCATGGAAGCCAAGGACAAAAGAATGAAAGCCACT
TCAGAAGTCCTTCGGAACATGAAGATACTAAAACTCCAAGCATGGGACACTCAATTCCTACACAAGATAGAAAGCTTGAGGAAAATAGAGTATAATTGTC
TATGGAAGTCTCTCAGGCTTTCAGCAATTTCAGCTTTTGTCTTCTGGGGATCACCCACGTTTATCTCTGTGGTCACGTTTGGTGCATGTATGCTAATGGG
AATCCAACTCACAGCAGGAAGAGTCCTATCAGCATTGGCCACCTTCCGAATGTTGCAGGACCCCATATTCAATCTACCTGACCTGCTCTCTGTGATTGCA
CAGGGAAAAGTTTCAGCAGATAGAGTTGCTTCCTTCCTTCAGGAAGGTGAAATTCAGCATGATGCCACTGAGCATGTTCCAAAAGATCAAGCAGAGTATG
CAATTAGTATTGATGATGGACGATTCTGCTGGGACTCCGATTCAAGCAACCCAACCCTTGATGAAATACGGTTGAAAGTGAAGAGAGGTATGAAAGTCGC
AATTTGTGGGACCGTGGGATCAGGCAAGTCCAGCCTGCTTTCTTGCATTCTTGGAGAAATACAGAAGTTGTCAGGGACAGTGAAGATCAGTGGCGCAAAA
GCTTACGTTCCTCAATCTCCATGGATACTGACAGGAAATATCAGGGAGAACATACTATTTGGGAATCCATATGACAGTGTCAGGTATTACAGAACAGTTA
AAGCATGTGCTTTGTTGAAGGATTTTGAACTGTTCTCCAGTGGTGACTTAACTGATATTGGAGAAAGAGGGATAAACATGAGCGGGGGACAGAAGCAGAG
GATACAGATAGCTCGTGCAGTTTACCAGGATGCTGATATATATTTATTTGATGACCCTTTCAGTGCTGTAGATGCTCATACAGGATCCCAACTCTTTCAG
GAATGCCTAATGGGAATTCTCAAAGATAAGACTATAATTTATGTCACCCATCAAGTGGAGTTTCTTCCTGCTGCAGACATCATTCTGGTGATGCAAAATG
GAAGAATAGCAGAGGCTGGAACATTTAGTGAACTTTTGAAACAGAACGTAGGATTTGAAGCTTTAGTCGGTGCTCATAGCCAAGCTCTAGAATCAGTCCT
TACAGTTGAGAATTCGAGGAGAACATCTCAAGATCCAGAACCTGACAGTGAATCTAATACAGAATCCACTTCAAACTCAAACTGTCTCTCACATTACGAG
TCGGACCATGACCTCTCTGTAGAAATAACAGAAAAGGGAGGAAAATTCGTGCAAGATGAAGAGAGAGAGAAAGGAAGCATTGGAAAAGAAGTTTACTGGT
CATACTTGACCACTGTGAAAGGTGGTGCCCTGGTCCCATGTATAATCTTAGCACAGTCATTATTCCAAATTTTACAGATAGTTAGTAACTACTGGATGGC
ATGGTCTTCTCCTCCTACAAGTGACACTGCTCCAGTATATGGGATGAATTTCATATTGCTTGTTTATACACTACTTTCTATTAGTAGTTCACTATGTGTC
CTGGTCCGAGCCACACTAGTAGCAATAGCAGGACTTTCAACAGCACAAAAGCTCTTCACAAACATGCTGCGTAGTCTACTCCGAGCTCCAATGGCATTTT
TTGATTCAACCCCAACCGGAAGGATATTAAACCGAGCATCAATGGATCAAAGCGTCATAGACATGGAAATTGCACAAAGACTCGGCTGGTGTGCTTTCTC
AATAATACAGATTCTAGGGACCATTGCAGTGATGTCCCAGGTCGCATGGGAAGTGTTTGTGATCTTCATTCCAGTAACTGCAGTCTGCATATGGTACCAA
CAATATTACACGCCAACAGCAAGAGAACTGGCTCGCTTAGCAGGGATACAACAAGCCCCGATCCTCCATCATTTTTCAGAATCACTAGCAGGAGCAGCAA
CAATTCGTGCTTTTGATCAACAAGAACGTTTTTATTGTTCAAACCTTGATCTGATAGACAACCACTCAAGGCCATGGTTTCATAATGTATCTGCAATGGA
ATGGCTTTCTTTCAGGCTGAATTTGCTATCCAATTTTGTGTTTGCCTTCTCATTGGTTTTGCTGGTGAGCCTTCCTGAAGGAGTAATCAGTCCAAGCATT
GCTGGGTTAGCTGTAACATATGGAATAAACTTGAATGTTTTGCAAGCTTCCGTCATATGGAACATATGCAATGCAGAAAATAAAATGATCTCAATAGAAA
GAGTACTTCAGTACTCCAGCATAACGAGTGAAGCGCCCCTTGTGCTTGAGCAGAGCAGGCCTCCAAACAAGTGGCCAGAAGTAGGAGCGATCTGCTTCAA
AGATTTGCAGATTCGCTACGCAGAGCATCTGCCATCTGTTCTGAAAAACATTAACTGTGCATTTCCTGGAAGAAAAAAGGTTGGAGTTGTAGGAAGGACA
GGAAGTGGGAAATCAACCCTCATACAGGCAATTTTCAGAATTGTGGAACCCAGAGAGGGAAGCATTATCATCGATGATGTGGACATTTCAAAGATAGGTC
TTCAAGACTTGAGATCAAGGCTTAGCATCATCCCCCAAGACCCAACAATGTTTGAGGGAACAGTGAGGGGAAACCTTGACCCACTGGGTCAGTACTCTGA
CTATGAAATATGGGAGGCTCTAGAAAAGTGTCAACTAGGAGATTTGGTGCGCGGAAAAGATGAAAAGTTGGACTCCCCAGTGGTGGAAAATGGAGAAAAT
TGGAGTGTGGGTCAAAGGCAACTATTCTGTTTGGGAAGAGCATTGCTAAAGAAAAGCAGGATTCTTGTACTGGATGAAGCAACAGCATCAGTCGATTCAG
CAACTGATGGGGTGATACAAAAGATCATCAGTCAAGAATTCAAAGATCGAACAGTGGTCACAATAGCTCACAGAATCCACACAGTCATAGACAGTGATCT
TGTTTTGGTCCTTAGTGATGGGAGAGTTGCAGAGTTTGATACGCCAGCAAGGCTACTAGAAAGGGAGGAATCTTTCTTTTCAAAGCTTATAAAAGAGTAT
TCCATGAGATCTCAGAGCTTCAACAACTTAACAAACGTGCACGCATAA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.014G180100.1 pacid=42762680 polypeptide=Potri.014G180100.1.p locus=Potri.014G180100 ID=Potri.014G180100.1.v4.1 annot-version=v4.1
MLDLGAAAAAAAAAAANLKLLIRMDWPQLQSPCLREHITIGVQLGFLGILLLHLLRKCADLAFNGGTKTTDQGKENYHIGLKFSNSYKASMVCSTCLLGV
HISMLLVLLNGQETSCNSIVRVFSAEVLQMISWAITLVAVFRIFPSRRYVKFPWIIRAWWLCSFMLSIVCTSLDINFKITNHGHLRLRDYAELFALLPST
FLLAISFRGKTGIVFNAFNGVTDPLLHEKSDKDSDTKRESPYGKATLLQLITFSWLTPLFAVGYKKPLEQDEIPDVYIKDSAGFLSSSFDENLNQVKEKD
RTANPSIYKAIFLFIRKKAAINALFAVTSAAASYVGPYLIDDFVNFLTEKKTRSLQSGYLLALGFLGAKTVETIAQRQWIFGARQLGLRLRASLISHIYK
KGLLLSSQSRQSHTSGEIINYMSVDIQRITDFIWYLNYIWMLPVQITLAIYILHTTLGLGSMAALTATLAVMACNIPITRFQKRYQTKIMEAKDKRMKAT
SEVLRNMKILKLQAWDTQFLHKIESLRKIEYNCLWKSLRLSAISAFVFWGSPTFISVVTFGACMLMGIQLTAGRVLSALATFRMLQDPIFNLPDLLSVIA
QGKVSADRVASFLQEGEIQHDATEHVPKDQAEYAISIDDGRFCWDSDSSNPTLDEIRLKVKRGMKVAICGTVGSGKSSLLSCILGEIQKLSGTVKISGAK
AYVPQSPWILTGNIRENILFGNPYDSVRYYRTVKACALLKDFELFSSGDLTDIGERGINMSGGQKQRIQIARAVYQDADIYLFDDPFSAVDAHTGSQLFQ
ECLMGILKDKTIIYVTHQVEFLPAADIILVMQNGRIAEAGTFSELLKQNVGFEALVGAHSQALESVLTVENSRRTSQDPEPDSESNTESTSNSNCLSHYE
SDHDLSVEITEKGGKFVQDEEREKGSIGKEVYWSYLTTVKGGALVPCIILAQSLFQILQIVSNYWMAWSSPPTSDTAPVYGMNFILLVYTLLSISSSLCV
LVRATLVAIAGLSTAQKLFTNMLRSLLRAPMAFFDSTPTGRILNRASMDQSVIDMEIAQRLGWCAFSIIQILGTIAVMSQVAWEVFVIFIPVTAVCIWYQ
QYYTPTARELARLAGIQQAPILHHFSESLAGAATIRAFDQQERFYCSNLDLIDNHSRPWFHNVSAMEWLSFRLNLLSNFVFAFSLVLLVSLPEGVISPSI
AGLAVTYGINLNVLQASVIWNICNAENKMISIERVLQYSSITSEAPLVLEQSRPPNKWPEVGAICFKDLQIRYAEHLPSVLKNINCAFPGRKKVGVVGRT
GSGKSTLIQAIFRIVEPREGSIIIDDVDISKIGLQDLRSRLSIIPQDPTMFEGTVRGNLDPLGQYSDYEIWEALEKCQLGDLVRGKDEKLDSPVVENGEN
WSVGQRQLFCLGRALLKKSRILVLDEATASVDSATDGVIQKIISQEFKDRTVVTIAHRIHTVIDSDLVLVLSDGRVAEFDTPARLLEREESFFSKLIKEY
SMRSQSFNNLTNVHA
|
DESeq2's median of ratios [POPLAR]
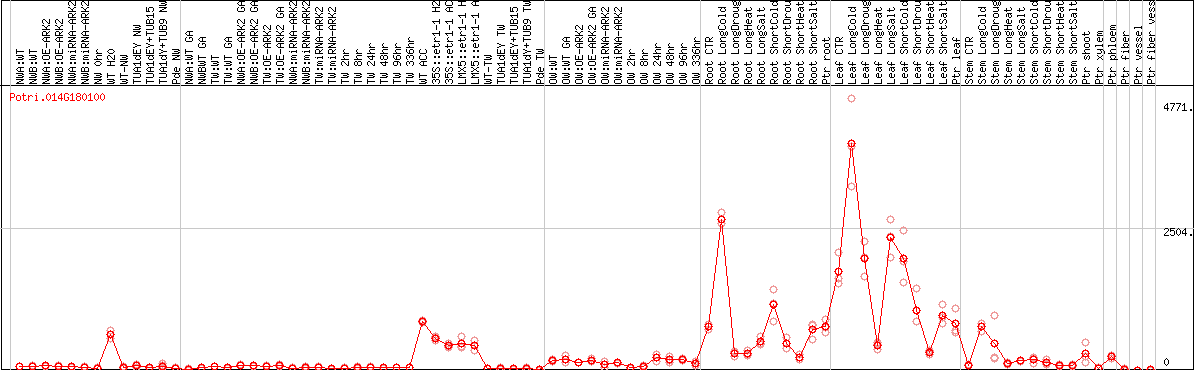
Coexpressed genes
Potri.014G180100 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
