Potri.014G180200 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.014G180200.1 pacid=42764319 polypeptide=Potri.014G180200.1.p locus=Potri.014G180200 ID=Potri.014G180200.1.v4.1 annot-version=v4.1
ATGCCTCCAAAGAAGAAATCCAAGTCGTCTTCTTCTTCAACCTCATCAGCGAAGAAAAACCATCAAAACTCACAGCAGCAGCAGCAGCAGCAGCAGCAGC
AACAACCTTCCAAGTTTGGAATCCAACATTTCTTCAATCTCCACACACAAAACGCACTTTCCCTCTCTCAAAATCCCCAACCTCCACCCACTCCTCCAAT
TCCCCAAACCCTCTCTCAAAACCCTATAATTTCCCCTAATTCTCTAAACCCTAAATTTGCGCCACCCTCCAATTCTCACCACCTCGATGCCGACGATAAC
GTGATGGATGTCTCACCTGAAATCACCAAGTCGGTGTCTCTCCAACGCTTCAAGTTCTCTCCTGGAATGTTGATAAAGCAGAGCCAGGATGATGGTGGCG
ATGAGGTCACATGGAAGATATCTCCGGTTAGTGAAAGACTCCAAGCTGTCTCCAAGCAAATGCCCCAACTCATTCAACTCTTAGCTGAAACTTCAAAGCT
TAATTCTTTCAGCATTCGTCCTTGTTCCTCTTTAAATGACGAGATTTCATCTGGTACTGCTGGCAAGGTTGATAAGCAGGTTCCTTCACCTCCATGGAAG
GGTGCTGACAGATCTTTGGTGCCTGCTAGTAGAGTTGGATTGAAAAGAATAAACCCACGTCAGGATGTAAATTTGACTGATGTTAATTGCTCGCTTGCTG
GTCGACAAAGTCCTTTCAGAACTCCGCCGTCCCTTTCGTATTGTCACGAAAAGCTTTCTAATGTTGCTGAATGCAATGGAGCATCAGATCAACTGGATCA
AAGGCAACACAAAAAGGCATTGCTTGAGCTTTTGGATCAAGTAGAAGATGCAATTTCTGTTGAAGATTCAGAACCTAGTACTGTCAATTCATCTAAAACT
CAAGATGGAAATGGTTATGACATGCCCTGCAATGCTGGATCTCTAGTAAAAGAAGCAGCAATAGATCCACCAGAAAGTGTTGCTGTGCCATTGTCAAATT
ATAATTTTCTTGTATTGGAGGTATCTGAGAAGCACAGGCCCGCGGATTTATCTGGTGCTCAATGCCCTTATAAGGTTCTTCGATTGTTGAATGAGCAAAG
TGGAGAAGAGCGAACTGTCTATTTGTGGGAAGAGTGGTTCTATAGTGTCATTTCACCTGGAGATACTGTAAATGTGATTGGTGAATTTGATGACCAAGGA
AAGTGTGACGTGGATCGTGACAATAATCTCTTAATTGTTCATCCTGACATTCTAGTCTCTGGAACTCGGGTTGCTGCTAGTTTCAGTTGTCCACGTCGAA
CTGTTTTAGATGAGAGACTAAAATGCAGTGAGCATTCAACTGCTGCATTAATTGGTACCTTGCTGCACCAAATTTTTCAGGCTGGACTCATGCAAGATAA
TCCTACCATAAACTTTTTGGAGGAATATGCTAGAATAGTACTCCAGAAAAATATTGAAAGCCTGCATGCATGTGGAGTAAATGAAAATGATATCTTCAAT
ACTTTGGTTGAGGCAATTCCTAAACTAATAAATTGGATCAACCTGTTCAAAGATTCACAGGATTCAAAAGCCCCCATTATTGATTTTGGTCCTGACAATG
GATTGAAAAAGCTAAGTATATCTGAGGTGATTGATATCGAGGAGATGGCATGGGCCCCAAAATATGGGTTGAAAGGAATGATTGATGCTTCTGTCCGCGT
TAAAGTAGAGTCAGGCAGAAATAAAGCAGATGAGAAGATTGTGCCTCTAGAGTTTAAAACTGGGAAAGTTTCCAATGGGCAGTCATCCATGGAACATGTA
GCCCAAGTGATTTTGTACACTCTTCTCATGTCTGAGAGGTATCTGAAACATATTGATTCTGGTCTTCTGTATTACCTCCAATCAGATCAGACACGGGGAA
TCACGGTTCAAAGGTCTGATGTGGTTGGGCTAATCATGCGTCGAAATGAGCTTGCAAATGACATCCTTAAGGCATCAAGAACTCAACAATTGCCTCCAAT
GCTACAGAGCCTAAATATGTGCAGAAGTTGCCGCCATCTTGATGTTTGTACAATCTATCATAAGGTGCATGGTGGAAGCAAAGAGAGTAGTGGACTAGGT
GATCTGTTTGATTCACATGTTCATCATCTGACAACTGCTCATTATGTTTTTCTTCGACGCTGGGATCAGTTGATTGACTTAGAGGCTAAAGAGACACAGC
TTGTAAAAAACAGAATATGGCGTCCACATAGTTTGAAGAGTGATCGTTCCACTAGTTGCCTTTCTTCTGTTGTTCTTGATACTTCAGATCGAGTTCCATA
TCAGAAATCTCTCAAAGACAACCGATTCATTTATCGTTTTGTCCATAAAAAAATGCCTCTTCATGACGTGCATGCTTCTGGTGGAGAATCTTTGAGTTTT
CCTTCATCTTCGGCAGAAGATTTCGATTATACACTTAAAAGTGGAGACTATGTGATTATTAGTACCAAATTTGGCCATCAAACTGTAGCAAGTGGGTTCA
TAACAGATATCAGTCGGTCCCATGTTTCTGTTTCTTTTCCTAAGCACTTAAGACTCCCTGGCAGCAATTCTTCGTCAGAAGCACATAATCTCTTTAGGGA
GGTCTGGCAGATTGACAAGGACGAGTTTATGACTTCATTTTCAGTTATGCGGTTCAACCTTGTACAACTGTTTCTCCAAAGTGAACAAAGTTCTCACCTT
AGGAAGATGATTGTTGATCTTGAGGCTCCTCGATTTGACAGTGGATGTATATTTAGTCAAGATCCAGCCCTATCTTATATATGGTCAGTGAAGAATTTAA
ATGGAGATCAGCGTCGGGCAATTCTGAAGACTCTTACAGCTAAGGATTATGCTCTAATACTAGGGATGCCTGGAACTGGAAAGACATCGACCTTGGTACA
TGCTGTCAAGGCCATGTTGATGAGAGGTGCATCCATTTTGCTCACGTCCTACACAAACTCTGCAATTGATAATTTACTAATCAAATTAAAAGCTCAGGGT
ATTGATTTTCTACGAATTGGAAGACATGAAGTTGTCCATGAGGAAGTCCGTGCCAATTGCGTTTCTGCAATGGATGTACACAGTGTAGAAGACATTAAAT
TAAGATTAGAGCAAGTCAAGGTTGTTGCAGTAACTTGCTTGGGAATTTCTAGTCCCTTGCTTGCCAACAAGAAATTTGATGTATGCATCATGGATGAAGC
TGGACAAATTACCCTCCCAATAGCTTTGGGACCCTTGATGTTTGCTTCAAAGTTTGTTCTTGTTGGTGATCACTATCAACTGCCTCCACTTGTCCAGAGT
ACAGAGGCTCGCGAGAATGGAATGGGGATAAGTTTGTTTTGTAGGCTTTCTGAGGCACATCCTCAAGCAATTTCAGCTTTGCAAAGCCAGTACCGCATGT
GTCAAGATATTATGGAACTATCAAATGCCTTGATATATGGTGACAGACTGCGTTGTGGTTCTTCTGAAATAGCAAATGCCAGGCTTAAATTTTCAGGTTT
ACAGTCCTGTTCATCGTGGCTAAAGGAGGTATTAAATCCTGGCAGACCCGTCATATTTATTAACACAGATATGCTGCCAGCTTACGAGGCAAAGGACTCC
AAAACTGTGAATAACCCAATAGAAGCTTACATAGTTGCAGAGGTCACAAAGGAATTACTTAATAATGGGATTGTAGGTGATGATATTGGCATTATTACCC
CTTATAATTCCCAGGCAAATCTCATCCGAGCTTCTGTTAACGTAACCTCTGTGGAGATTCATACCATTGATAAATACCAGGGAAGAGACAAGGAATGCAT
ACTGGTATCCTTTGCTAGGTCCAGTGAAAATCCAAGAAATTGTACGTCTTCGTTGCTTGGAGATTGGCATAGGATTAATGTGGCGCTCACACGAGCCAAG
AAAAAGTTGATCTTAGTAGGGTCGTGCAAAACCCTGTCAAAGGTCCCGCTGCTGAAGCTTCTTGTTGAGAAGGTTGAAGAGCAATCAGGCATTATCAATG
TGTCCAAGACAGATATCAACTACGAAGAGCACAAAGGAGAGCTGAAAAGATGTTCTCATATCAGATGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.014G180200.1 pacid=42764319 polypeptide=Potri.014G180200.1.p locus=Potri.014G180200 ID=Potri.014G180200.1.v4.1 annot-version=v4.1
MPPKKKSKSSSSSTSSAKKNHQNSQQQQQQQQQQQPSKFGIQHFFNLHTQNALSLSQNPQPPPTPPIPQTLSQNPIISPNSLNPKFAPPSNSHHLDADDN
VMDVSPEITKSVSLQRFKFSPGMLIKQSQDDGGDEVTWKISPVSERLQAVSKQMPQLIQLLAETSKLNSFSIRPCSSLNDEISSGTAGKVDKQVPSPPWK
GADRSLVPASRVGLKRINPRQDVNLTDVNCSLAGRQSPFRTPPSLSYCHEKLSNVAECNGASDQLDQRQHKKALLELLDQVEDAISVEDSEPSTVNSSKT
QDGNGYDMPCNAGSLVKEAAIDPPESVAVPLSNYNFLVLEVSEKHRPADLSGAQCPYKVLRLLNEQSGEERTVYLWEEWFYSVISPGDTVNVIGEFDDQG
KCDVDRDNNLLIVHPDILVSGTRVAASFSCPRRTVLDERLKCSEHSTAALIGTLLHQIFQAGLMQDNPTINFLEEYARIVLQKNIESLHACGVNENDIFN
TLVEAIPKLINWINLFKDSQDSKAPIIDFGPDNGLKKLSISEVIDIEEMAWAPKYGLKGMIDASVRVKVESGRNKADEKIVPLEFKTGKVSNGQSSMEHV
AQVILYTLLMSERYLKHIDSGLLYYLQSDQTRGITVQRSDVVGLIMRRNELANDILKASRTQQLPPMLQSLNMCRSCRHLDVCTIYHKVHGGSKESSGLG
DLFDSHVHHLTTAHYVFLRRWDQLIDLEAKETQLVKNRIWRPHSLKSDRSTSCLSSVVLDTSDRVPYQKSLKDNRFIYRFVHKKMPLHDVHASGGESLSF
PSSSAEDFDYTLKSGDYVIISTKFGHQTVASGFITDISRSHVSVSFPKHLRLPGSNSSSEAHNLFREVWQIDKDEFMTSFSVMRFNLVQLFLQSEQSSHL
RKMIVDLEAPRFDSGCIFSQDPALSYIWSVKNLNGDQRRAILKTLTAKDYALILGMPGTGKTSTLVHAVKAMLMRGASILLTSYTNSAIDNLLIKLKAQG
IDFLRIGRHEVVHEEVRANCVSAMDVHSVEDIKLRLEQVKVVAVTCLGISSPLLANKKFDVCIMDEAGQITLPIALGPLMFASKFVLVGDHYQLPPLVQS
TEARENGMGISLFCRLSEAHPQAISALQSQYRMCQDIMELSNALIYGDRLRCGSSEIANARLKFSGLQSCSSWLKEVLNPGRPVIFINTDMLPAYEAKDS
KTVNNPIEAYIVAEVTKELLNNGIVGDDIGIITPYNSQANLIRASVNVTSVEIHTIDKYQGRDKECILVSFARSSENPRNCTSSLLGDWHRINVALTRAK
KKLILVGSCKTLSKVPLLKLLVEKVEEQSGIINVSKTDINYEEHKGELKRCSHIR
|
DESeq2's median of ratios [POPLAR]
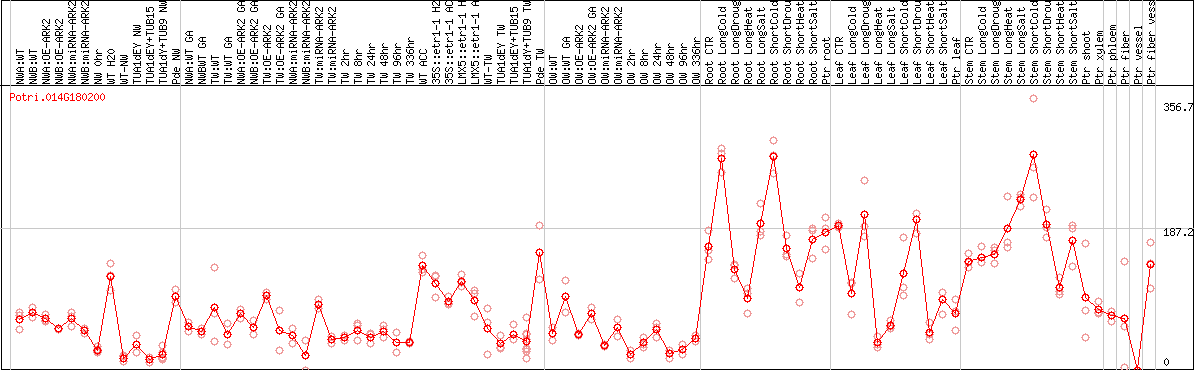
Coexpressed genes
Potri.014G180200 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
