External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G07330 429 / 3e-148
ATCSLC6, ATCSLC06
CELLULOSE-SYNTHASE LIKE C6, Cellulose-synthase-like C6 (.1)
AT3G28180 385 / 3e-131
ATCSLC4, CSLC4, ATCSLC04
CELLULOSE-SYNTHASE LIKE C4, Cellulose-synthase-like C4 (.1)
AT4G31590 383 / 6e-130
ATCSLC5, ATCSLC05
CELLULOSE-SYNTHASE LIKE C5, Cellulose-synthase-like C5 (.1)
AT2G24630 383 / 6e-130
ATCSLC8, ATCSLC08
CELLULOSE-SYNTHASE LIKE C8, Glycosyl transferase family 2 protein (.1)
AT4G07960 375 / 5e-127
ATCSLC12
CELLULOSE-SYNTHASE LIKE C12, Cellulose-synthase-like C12 (.1)
AT5G22740 261 / 3e-84
ATCSLA2, ATCSLA02
CELLULOSE SYNTHASE-LIKE A 2, ARABIDOPSIS THALIANA CELLULOSE SYNTHASE-LIKE A2, ARABIDOPSIS THALIANA CELLULOSE SYNTHASE-LIKE A02, cellulose synthase-like A02 (.1)
AT5G03760 254 / 7e-82
ATCSLA9, RAT4, CSLA9, ATCSLA09
RESISTANT TO AGROBACTERIUM TRANSFORMATION 4, CELLULOSE SYNTHASE LIKE A9, Nucleotide-diphospho-sugar transferases superfamily protein (.1)
AT1G23480 252 / 2e-81
ATCSLA3, ATCSLA03
cellulose synthase-like A3 (.1.2.3)
AT5G16190 244 / 1e-78
ATCSLA11
ARABIDOPSIS THALIANA CELLULOSE SYNTHASE LIKE A11, cellulose synthase like A11 (.1)
AT2G35650 246 / 4e-78
ATCSLA7, CSLA7, ATCSLA07
CELLULOSE SYNTHASE LIKE A7, cellulose synthase like (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.014G190900
472 / 3e-166
AT3G07330 863 / 0.0
CELLULOSE-SYNTHASE LIKE C6, Cellulose-synthase-like C6 (.1)
Potri.002G248400
464 / 6e-162
AT3G07330 957 / 0.0
CELLULOSE-SYNTHASE LIKE C6, Cellulose-synthase-like C6 (.1)
Potri.005G146900
397 / 2e-135
AT4G07960 1050 / 0.0
CELLULOSE-SYNTHASE LIKE C12, Cellulose-synthase-like C12 (.1)
Potri.006G270900
389 / 3e-132
AT4G31590 1085 / 0.0
CELLULOSE-SYNTHASE LIKE C5, Cellulose-synthase-like C5 (.1)
Potri.002G114200
389 / 3e-132
AT4G07960 1031 / 0.0
CELLULOSE-SYNTHASE LIKE C12, Cellulose-synthase-like C12 (.1)
Potri.018G009300
388 / 5e-132
AT4G31590 1077 / 0.0
CELLULOSE-SYNTHASE LIKE C5, Cellulose-synthase-like C5 (.1)
Potri.008G026400
259 / 8e-84
AT5G03760 854 / 0.0
RESISTANT TO AGROBACTERIUM TRANSFORMATION 4, CELLULOSE SYNTHASE LIKE A9, Nucleotide-diphospho-sugar transferases superfamily protein (.1)
Potri.004G189000
259 / 1e-83
AT5G22740 927 / 0.0
CELLULOSE SYNTHASE-LIKE A 2, ARABIDOPSIS THALIANA CELLULOSE SYNTHASE-LIKE A2, ARABIDOPSIS THALIANA CELLULOSE SYNTHASE-LIKE A02, cellulose synthase-like A02 (.1)
Potri.006G116900
256 / 3e-82
AT5G03760 928 / 0.0
RESISTANT TO AGROBACTERIUM TRANSFORMATION 4, CELLULOSE SYNTHASE LIKE A9, Nucleotide-diphospho-sugar transferases superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10038217
423 / 3e-148
AT3G07330 821 / 0.0
CELLULOSE-SYNTHASE LIKE C6, Cellulose-synthase-like C6 (.1)
Lus10025886
424 / 8e-146
AT3G07330 994 / 0.0
CELLULOSE-SYNTHASE LIKE C6, Cellulose-synthase-like C6 (.1)
Lus10007715
389 / 1e-132
AT4G07960 1010 / 0.0
CELLULOSE-SYNTHASE LIKE C12, Cellulose-synthase-like C12 (.1)
Lus10018651
387 / 2e-132
AT4G07960 978 / 0.0
CELLULOSE-SYNTHASE LIKE C12, Cellulose-synthase-like C12 (.1)
Lus10039440
386 / 3e-131
AT3G28180 1035 / 0.0
CELLULOSE-SYNTHASE LIKE C4, Cellulose-synthase-like C4 (.1)
Lus10039475
385 / 6e-131
AT3G28180 1035 / 0.0
CELLULOSE-SYNTHASE LIKE C4, Cellulose-synthase-like C4 (.1)
Lus10020120
380 / 1e-129
AT4G31590 1012 / 0.0
CELLULOSE-SYNTHASE LIKE C5, Cellulose-synthase-like C5 (.1)
Lus10026923
382 / 2e-129
AT4G31590 1078 / 0.0
CELLULOSE-SYNTHASE LIKE C5, Cellulose-synthase-like C5 (.1)
Lus10020539
255 / 3e-83
AT5G22740 612 / 0.0
CELLULOSE SYNTHASE-LIKE A 2, ARABIDOPSIS THALIANA CELLULOSE SYNTHASE-LIKE A2, ARABIDOPSIS THALIANA CELLULOSE SYNTHASE-LIKE A02, cellulose synthase-like A02 (.1)
Lus10008646
258 / 4e-83
AT5G03760 890 / 0.0
RESISTANT TO AGROBACTERIUM TRANSFORMATION 4, CELLULOSE SYNTHASE LIKE A9, Nucleotide-diphospho-sugar transferases superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0110
GT-A
PF13641
Glyco_tranf_2_3
Glycosyltransferase like family 2
Representative CDS sequence
>Potri.014G190701.1 pacid=42764084 polypeptide=Potri.014G190701.1.p locus=Potri.014G190701 ID=Potri.014G190701.1.v4.1 annot-version=v4.1
ATGCAGGTTTACCAACAATCTATTGCAGCCTGCTGTATCCAGGACTGGCCAAAGGAGAGAATGCTTATACAGGTTTTGGATGATTCTGATGAGTTAGATG
CCCAACTCCTTATCAAGGCTGAAGTACAGAAATGGCAACAAAGGGGTGTGCATATATTGTATAGGCACCGTCTTATACGCACAGGCTACAAGGCTGGGAA
CCCGAAATCTGCAATGAGCTGTGATTATGTAAAAGATTATGAGTTTGTAGCAATATTTGATGCTGATTTTCAGCCAGGACCTGATTTCTTGAAGAGAACT
ATTCCTCACTTCAAGGGGAAGGATGACCTAGCATTGGTCCAAGCAAGGTGGGCATTTGTAAACAAGGATGAAAACTTGCTTACAAGACTGCAGAACATAA
ACTTATCATTCCACTTTGAGGTTGAGCAACAAGTGAATGGTGTTTTCATCAACTTTTTTGGTTTTAATGGAACTGCTGGTGTATGGAGGATTAAAGCCCT
TGAAGACTGTGGTGGTTGGTTGGAACGGACAACAGTTGAAGACATGGATATTGCTGTTCGTGCTCATCTTTGTGGATGGAAGTTCATATATTTGAATGAT
GTTAAGTGCCTTTGTGAACTTCCAGAGTCCTATGAGGCATACAAAAAACAGCAACATCGTTGGCATTCAGGTCCAATGCAGTTGTTCCGTTTGTGCTTTG
TTGACATACTTCGTGCAAAGGTACTGAAGTTTTCCCCCCCTCTTGCTATCATTTATATTTGGTAG
AA sequence
>Potri.014G190701.1 pacid=42764084 polypeptide=Potri.014G190701.1.p locus=Potri.014G190701 ID=Potri.014G190701.1.v4.1 annot-version=v4.1
MQVYQQSIAACCIQDWPKERMLIQVLDDSDELDAQLLIKAEVQKWQQRGVHILYRHRLIRTGYKAGNPKSAMSCDYVKDYEFVAIFDADFQPGPDFLKRT
IPHFKGKDDLALVQARWAFVNKDENLLTRLQNINLSFHFEVEQQVNGVFINFFGFNGTAGVWRIKALEDCGGWLERTTVEDMDIAVRAHLCGWKFIYLND
VKCLCELPESYEAYKKQQHRWHSGPMQLFRLCFVDILRAKVLKFSPPLAIIYIW
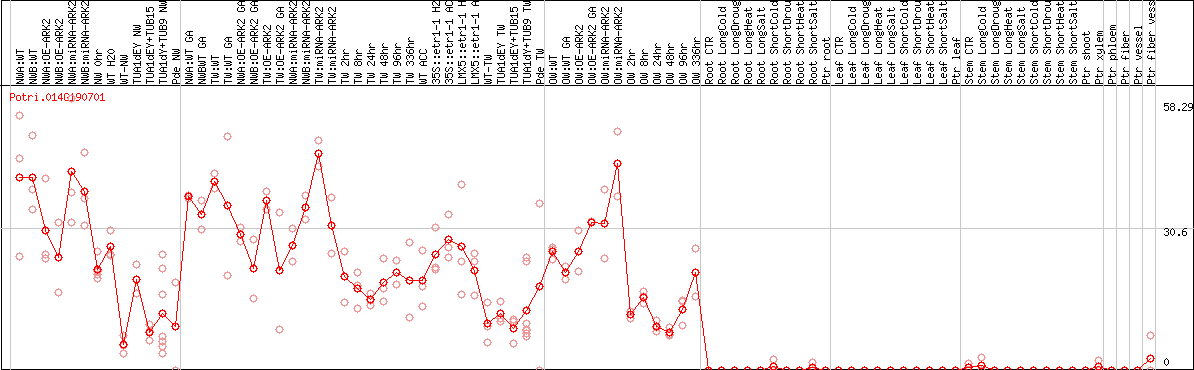
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.014G190701 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.