External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G24530 521 / 0
DMR6
DOWNY MILDEW RESISTANT 6, 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT4G10490 375 / 4e-130
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT4G10500 372 / 9e-129
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT2G36690 240 / 6e-77
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT2G44800 236 / 3e-75
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT3G60290 229 / 1e-72
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT3G51240 228 / 4e-72
TT6, F3'H, F3H
TRANSPARENT TESTA 6, flavanone 3-hydroxylase (.1.2)
AT2G38240 224 / 1e-70
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT5G05600 222 / 1e-69
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
AT3G55970 216 / 2e-67
ATJRG21
jasmonate-regulated gene 21 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.012G006300
664 / 0
AT5G24530 506 / 0.0
DOWNY MILDEW RESISTANT 6, 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Potri.011G150100
404 / 2e-141
AT4G10490 501 / 2e-179
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Potri.001G451900
399 / 2e-139
AT4G10490 498 / 3e-178
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Potri.014G106700
384 / 1e-133
AT4G10490 468 / 2e-166
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Potri.001G451700
357 / 4e-123
AT4G10500 451 / 6e-160
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Potri.001G451600
313 / 8e-106
AT4G10500 406 / 5e-142
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Potri.001G451500
290 / 2e-96
AT4G10500 325 / 2e-110
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Potri.011G150200
287 / 1e-95
AT4G10500 323 / 1e-109
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Potri.002G039500
280 / 8e-93
AT4G10500 244 / 1e-78
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10032930
541 / 0
AT5G24530 504 / 0.0
DOWNY MILDEW RESISTANT 6, 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Lus10015573
538 / 0
AT5G24530 510 / 0.0
DOWNY MILDEW RESISTANT 6, 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Lus10030185
377 / 8e-131
AT4G10490 442 / 3e-156
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Lus10035782
275 / 2e-90
AT5G24530 258 / 4e-84
DOWNY MILDEW RESISTANT 6, 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Lus10024882
270 / 4e-89
AT5G24530 271 / 2e-89
DOWNY MILDEW RESISTANT 6, 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Lus10000711
265 / 6e-87
AT5G24530 263 / 4e-86
DOWNY MILDEW RESISTANT 6, 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Lus10014398
234 / 4e-74
AT2G36690 482 / 4e-171
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
Lus10005037
231 / 3e-73
AT3G11180 471 / 3e-166
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1), 2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.2)
Lus10041913
225 / 3e-71
AT3G51240 602 / 0.0
TRANSPARENT TESTA 6, flavanone 3-hydroxylase (.1.2)
Lus10004808
225 / 4e-71
AT5G05600 457 / 5e-162
2-oxoglutarate (2OG) and Fe(II)-dependent oxygenase superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0029
Cupin
PF03171
2OG-FeII_Oxy
2OG-Fe(II) oxygenase superfamily
CL0029
Cupin
PF14226
DIOX_N
non-haem dioxygenase in morphine synthesis N-terminal
Representative CDS sequence
>Potri.015G002800.1 pacid=42776326 polypeptide=Potri.015G002800.1.p locus=Potri.015G002800 ID=Potri.015G002800.1.v4.1 annot-version=v4.1
ATGGATACAAAAGTAATTTCCTCTGGCGTCCATTACACGAACTTGCCGGCTAGCTATGTTAGGCCGGAATCTGAGCGTCCCAGGCTGTCGGAAGTTTCAA
CGTGCGAAGATGTTCCGGTTATAGATTTAGGTTGTCAGGATAGGAACCAAATTGTCCAACAAGTTGGTGATGCCTGCGAGCACTATGGCTTTTTCCAGGT
GATCAATCATGGAGTGTCCTTGGAAGCGGTGGAGAAAATGTTAGGAGTGGCCCATGATTTCTTCAGCTTGCCGGTTGAGGAGAAGCTGAAGCTGTACTCA
GATGACCCATCAAAGACAATGAGACTTTCTACAAGTTTTAATGTGAATAAGGAAAAGGTCCACAACTGGAGGGATTATCTTAGACTCCATTGCTACCCTC
TTGACAAATATGTGCCTGAGTGGCCTTCTAATCCTCCTCCTTTCAAGGAAATTGTAAGGAGTTACAGTATACAAGTTCGAGAACTGGGGTTTAGAATACA
GGAACTGATATCAGAGAGCTTAGGCCTCGAAAAGGATCACATAAAAAATGTTTTGGGTGAACAAGGACAGCATATGGCTGTAAACTTTTACCCTCCATGT
CCAGAGCCAGAGTTGACTTATGGATTGCCAGCACATACAGACCCCAATGCCCTAACAATTTTACTTCAAGACCTATCAGTGGCAGGCCTTCAAGTCCTCC
TCAAAGATGGAAAGTGGGTTGCCGTTAATCCTCATCCGGATGCATTTGTCATCAACATTGGTGACCAGTTACAGGCACTAAGCAATGGAAGATATAAGAG
TGTGTGGCACCGAGCCATTACCAATACTGACAAGGCAAGAATGTCAGTTGCTTCCTTCCTCTGTCCGTTTGATAACGCATTGATCACCCCACCTAAAGCT
CTCACCGATGATGGAACCGGAGCCATATACAGGGATTTCACGTATGCTGAATATTACAAGAAGTTTTGGAGCAGGAACTTGGATCAAGAACATTGTTTGG
AACTTTTCAAGAACTAA
AA sequence
>Potri.015G002800.1 pacid=42776326 polypeptide=Potri.015G002800.1.p locus=Potri.015G002800 ID=Potri.015G002800.1.v4.1 annot-version=v4.1
MDTKVISSGVHYTNLPASYVRPESERPRLSEVSTCEDVPVIDLGCQDRNQIVQQVGDACEHYGFFQVINHGVSLEAVEKMLGVAHDFFSLPVEEKLKLYS
DDPSKTMRLSTSFNVNKEKVHNWRDYLRLHCYPLDKYVPEWPSNPPPFKEIVRSYSIQVRELGFRIQELISESLGLEKDHIKNVLGEQGQHMAVNFYPPC
PEPELTYGLPAHTDPNALTILLQDLSVAGLQVLLKDGKWVAVNPHPDAFVINIGDQLQALSNGRYKSVWHRAITNTDKARMSVASFLCPFDNALITPPKA
LTDDGTGAIYRDFTYAEYYKKFWSRNLDQEHCLELFKN
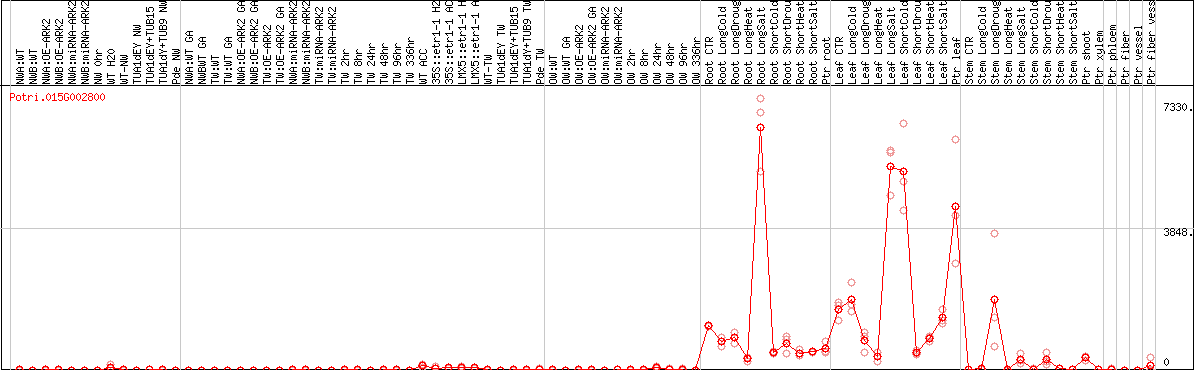
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.015G002800 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.