External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G27590 141 / 6e-44
Heavy metal transport/detoxification superfamily protein (.1.2)
AT2G37390 60 / 2e-11
NAKR2
SODIUM POTASSIUM ROOT DEFECTIVE 2, Chloroplast-targeted copper chaperone protein (.1.2)
AT3G56891 56 / 2e-10
Heavy metal transport/detoxification superfamily protein (.1)
AT5G02600 57 / 3e-10
NPCC6, NAKR1
nuclear-enriched phloem companion cell gene 6, SODIUM POTASSIUM ROOT DEFECTIVE 1, Heavy metal transport/detoxification superfamily protein (.1.2)
AT4G08570 52 / 4e-09
Heavy metal transport/detoxification superfamily protein (.1)
AT3G53530 53 / 6e-09
NAKR3
SODIUM POTASSIUM ROOT DEFECTIVE 3, Chloroplast-targeted copper chaperone protein (.1.2)
AT1G06330 51 / 1e-08
Heavy metal transport/detoxification superfamily protein (.1)
AT1G66240 48 / 7e-08
ATX1, ATATX1
homolog of anti-oxidant 1 (.1.2.3)
AT5G27690 48 / 3e-07
Heavy metal transport/detoxification superfamily protein (.1)
AT1G22990 45 / 2e-06
HIPP22
heavy metal associated isoprenylated plant protein 22, Heavy metal transport/detoxification superfamily protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.012G007200
218 / 2e-74
AT4G27590 137 / 2e-42
Heavy metal transport/detoxification superfamily protein (.1.2)
Potri.010G114600
55 / 4e-10
AT1G22990 194 / 9e-65
heavy metal associated isoprenylated plant protein 22, Heavy metal transport/detoxification superfamily protein (.1)
Potri.002G092200
55 / 4e-10
AT4G08570 193 / 3e-64
Heavy metal transport/detoxification superfamily protein (.1)
Potri.016G080400
56 / 7e-10
AT2G37390 133 / 2e-37
SODIUM POTASSIUM ROOT DEFECTIVE 2, Chloroplast-targeted copper chaperone protein (.1.2)
Potri.005G167000
54 / 8e-10
AT4G08570 229 / 1e-78
Heavy metal transport/detoxification superfamily protein (.1)
Potri.006G024800
52 / 3e-09
AT3G56891 167 / 5e-54
Heavy metal transport/detoxification superfamily protein (.1)
Potri.006G213900
54 / 4e-09
AT2G37390 127 / 2e-35
SODIUM POTASSIUM ROOT DEFECTIVE 2, Chloroplast-targeted copper chaperone protein (.1.2)
Potri.001G452400
52 / 6e-09
AT2G18196 256 / 3e-88
Heavy metal transport/detoxification superfamily protein (.1)
Potri.005G079800
52 / 6e-09
AT4G39700 189 / 2e-62
Heavy metal transport/detoxification superfamily protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10032924
123 / 5e-36
AT4G27590 122 / 4e-35
Heavy metal transport/detoxification superfamily protein (.1.2)
Lus10016436
57 / 6e-11
AT4G39700 209 / 1e-70
Heavy metal transport/detoxification superfamily protein (.1)
Lus10019676
57 / 6e-11
AT4G39700 211 / 3e-71
Heavy metal transport/detoxification superfamily protein (.1)
Lus10022508
56 / 2e-10
AT4G39700 201 / 2e-67
Heavy metal transport/detoxification superfamily protein (.1)
Lus10016812
56 / 3e-10
AT4G39700 181 / 3e-59
Heavy metal transport/detoxification superfamily protein (.1)
Lus10025294
55 / 1e-09
AT2G37390 141 / 3e-40
SODIUM POTASSIUM ROOT DEFECTIVE 2, Chloroplast-targeted copper chaperone protein (.1.2)
Lus10024435
55 / 1e-09
AT2G37390 144 / 2e-41
SODIUM POTASSIUM ROOT DEFECTIVE 2, Chloroplast-targeted copper chaperone protein (.1.2)
Lus10016708
51 / 1e-08
AT1G71050 192 / 1e-63
heavy metal associated isoprenylated plant protein 20, Heavy metal transport/detoxification superfamily protein (.1)
Lus10013911
50 / 3e-08
AT1G06330 103 / 2e-28
Heavy metal transport/detoxification superfamily protein (.1)
Lus10033250
50 / 5e-08
AT2G18196 251 / 3e-86
Heavy metal transport/detoxification superfamily protein (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF00403
HMA
Heavy-metal-associated domain
Representative CDS sequence
>Potri.015G003800.1 pacid=42775927 polypeptide=Potri.015G003800.1.p locus=Potri.015G003800 ID=Potri.015G003800.1.v4.1 annot-version=v4.1
ATGGCAAGCATGCAAATTGTTCTAGCTTCCAGCTGCAAGAATGTTGAGGCACAGCACGTGGAGATGATGGTCCCTCTTTACTCCCATGGATGCGAGAAGA
AGGTGAAGAAGACCTTATCCCATCTCAAAGGAATTTACTCCGTAAATGTAGATTATTATCAGCAAAAGGTGACGGTATGGGGAATATGCAACAAACATGA
TGTATTGGCAACCATAAAGAGCAAAAGAAAGGAAGCTCGCTTTTGGAATCCCCAAGAAATGGAAGAAGAAGAATCACAACCACCGTCTCCGCCGCCGCCA
CCACCAAAAGACTCCAGTACTATTCCTTCTTTGACTCTAATGAAGGCTCGTTCCCTAACTCGTTCCTTAAGTTGGAAAGTATGGAAAAAGGTCTTCACGA
GAACCTATTCCTTCTAA
AA sequence
>Potri.015G003800.1 pacid=42775927 polypeptide=Potri.015G003800.1.p locus=Potri.015G003800 ID=Potri.015G003800.1.v4.1 annot-version=v4.1
MASMQIVLASSCKNVEAQHVEMMVPLYSHGCEKKVKKTLSHLKGIYSVNVDYYQQKVTVWGICNKHDVLATIKSKRKEARFWNPQEMEEEESQPPSPPPP
PPKDSSTIPSLTLMKARSLTRSLSWKVWKKVFTRTYSF
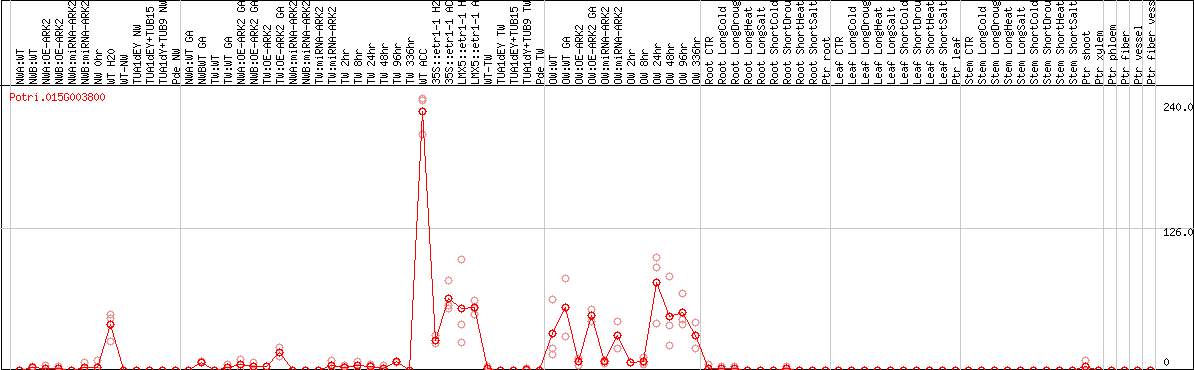
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.015G003800 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.