External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT4G18830 59 / 3e-11
OFP
ATOFP5, OFP5
ovate family protein 5 (.1)
AT5G19650 54 / 5e-10
OFP
ATOFP8, OFP8
ovate family protein 8 (.1)
AT3G52525 52 / 2e-09
OFP
ATOFP6, OFP6
ARABIDOPSIS THALIANA OVATE FAMILY PROTEIN 6, ovate family protein 6 (.1)
AT1G06920 51 / 1e-08
OFP
ATOFP4, OFP4
ARABIDOPSIS THALIANA OVATE FAMILY PROTEIN 4, ovate family protein 4 (.1)
AT2G36026 50 / 1e-08
OFP
Ovate family protein (.1)
AT1G79960 50 / 2e-08
OFP
ATOFP14, OFP14
ovate family protein 14 (.1)
AT2G18500 49 / 4e-08
OFP
ATOFP7, OFP7
ARABIDOPSIS THALIANA OVATE FAMILY PROTEIN 7, ovate family protein 7 (.1)
AT5G58360 49 / 6e-08
OFP
ATOFP3, OFP3
ARABIDOPSIS THALIANA OVATE FAMILY PROTEIN 3, ovate family protein 3 (.1)
AT5G04820 49 / 6e-08
OFP
ATOFP13, OFP13
ARABIDOPSIS THALIANA OVATE FAMILY PROTEIN 13, ovate family protein 13 (.1)
AT5G01840 47 / 3e-07
OFP
AtOFP1
ARABIDOPSIS THALIANA OVATE FAMILY PROTEIN 1, ovate family protein 1 (.1)
Paralogs
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10021302
58 / 2e-11
AT3G52525 120 / 2e-34
ARABIDOPSIS THALIANA OVATE FAMILY PROTEIN 6, ovate family protein 6 (.1)
Lus10016979
57 / 6e-11
AT5G04820 105 / 3e-27
ARABIDOPSIS THALIANA OVATE FAMILY PROTEIN 13, ovate family protein 13 (.1)
Lus10013318
57 / 6e-11
AT3G52525 82 / 4e-19
ARABIDOPSIS THALIANA OVATE FAMILY PROTEIN 6, ovate family protein 6 (.1)
Lus10021303
57 / 8e-11
AT5G04820 103 / 8e-26
ARABIDOPSIS THALIANA OVATE FAMILY PROTEIN 13, ovate family protein 13 (.1)
Lus10016978
56 / 2e-10
AT3G52525 114 / 4e-32
ARABIDOPSIS THALIANA OVATE FAMILY PROTEIN 6, ovate family protein 6 (.1)
Lus10005206
56 / 2e-10
AT3G52525 82 / 2e-19
ARABIDOPSIS THALIANA OVATE FAMILY PROTEIN 6, ovate family protein 6 (.1)
Lus10015236
53 / 3e-09
AT4G18830 112 / 1e-27
ovate family protein 5 (.1)
Lus10008589
51 / 1e-08
AT5G01840 120 / 6e-32
ARABIDOPSIS THALIANA OVATE FAMILY PROTEIN 1, ovate family protein 1 (.1)
Lus10034150
50 / 2e-08
AT5G04820 134 / 8e-38
ARABIDOPSIS THALIANA OVATE FAMILY PROTEIN 13, ovate family protein 13 (.1)
Lus10042224
50 / 2e-08
AT5G01840 114 / 4e-29
ARABIDOPSIS THALIANA OVATE FAMILY PROTEIN 1, ovate family protein 1 (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
PF04844
Ovate
Transcriptional repressor, ovate
Representative CDS sequence
>Potri.015G004300.1 pacid=42774925 polypeptide=Potri.015G004300.1.p locus=Potri.015G004300 ID=Potri.015G004300.1.v4.1 annot-version=v4.1
ATGGATGCTGGAGAAGACTACAAAAGGAAGGCAATTACAAGGAAGAAGAGTTCTTACAGGATTAGCCTTTCGGCTAGTTTACCTGAAGATGTTTGTGGTG
CTTTCTCAGGTGATACTATCTGTGCAGTTAAGCTCTCTAAGGATCCATTCTCAGACATGAGAGCATCAATATTAGAGATGATTCAGAACGTGGGTGTTCA
TGATTGGGATGAAATGGAAGAGTTAGTCTATTGTTACATTGCCCTTAACTCTCCTGACCTGCATGGCATCATTGCCAATGCATTTCTTAGTTTATCTTGT
CATTTTTCCTAG
AA sequence
>Potri.015G004300.1 pacid=42774925 polypeptide=Potri.015G004300.1.p locus=Potri.015G004300 ID=Potri.015G004300.1.v4.1 annot-version=v4.1
MDAGEDYKRKAITRKKSSYRISLSASLPEDVCGAFSGDTICAVKLSKDPFSDMRASILEMIQNVGVHDWDEMEELVYCYIALNSPDLHGIIANAFLSLSC
HFS
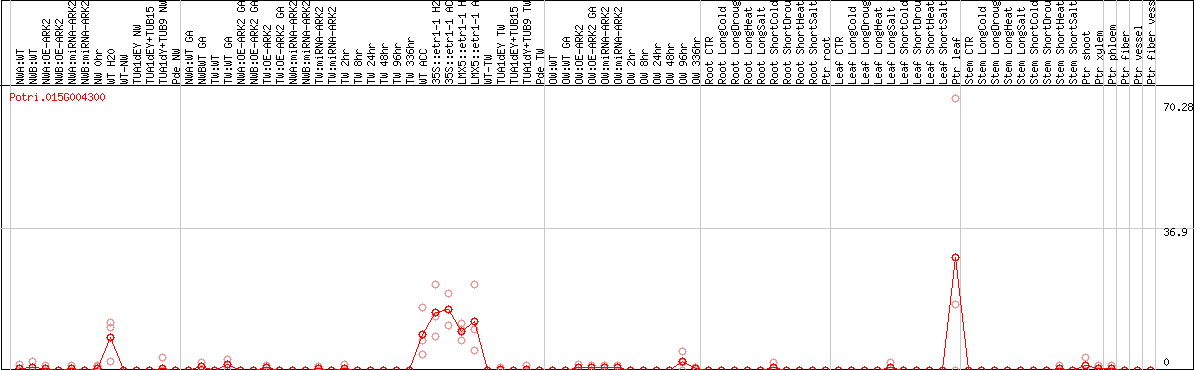
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.015G004300 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.