External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT5G24400 78 / 8e-19
PGL3, EMB2024
6-PHOSPHOGLUCONOLACTONASE 3, EMBRYO DEFECTIVE 2024, NagB/RpiA/CoA transferase-like superfamily protein (.1)
AT3G49360 49 / 2e-08
PGL2
6-phosphogluconolactonase 2 (.1)
AT5G24420 45 / 7e-07
PGL5
6-phosphogluconolactonase 5 (.1)
AT1G13700 41 / 2e-05
PGL1
6-phosphogluconolactonase 1 (.1)
AT5G24410 40 / 4e-05
PGL4
6-phosphogluconolactonase 4 (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.012G024400
94 / 9e-25
AT5G24400 384 / 3e-134
6-PHOSPHOGLUCONOLACTONASE 3, EMBRYO DEFECTIVE 2024, NagB/RpiA/CoA transferase-like superfamily protein (.1)
Potri.015G007200
92 / 3e-24
AT5G24400 395 / 1e-138
6-PHOSPHOGLUCONOLACTONASE 3, EMBRYO DEFECTIVE 2024, NagB/RpiA/CoA transferase-like superfamily protein (.1)
Potri.015G007300
76 / 2e-18
AT5G24400 357 / 2e-124
6-PHOSPHOGLUCONOLACTONASE 3, EMBRYO DEFECTIVE 2024, NagB/RpiA/CoA transferase-like superfamily protein (.1)
Potri.012G020100
61 / 7e-13
AT5G24400 229 / 9e-75
6-PHOSPHOGLUCONOLACTONASE 3, EMBRYO DEFECTIVE 2024, NagB/RpiA/CoA transferase-like superfamily protein (.1)
Potri.008G097200
55 / 2e-10
AT1G13700 386 / 6e-136
6-phosphogluconolactonase 1 (.1)
Potri.010G157200
53 / 8e-10
AT1G13700 383 / 1e-135
6-phosphogluconolactonase 1 (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10022352
82 / 3e-20
AT5G24400 382 / 2e-133
6-PHOSPHOGLUCONOLACTONASE 3, EMBRYO DEFECTIVE 2024, NagB/RpiA/CoA transferase-like superfamily protein (.1)
Lus10036919
59 / 9e-12
AT1G13700 383 / 1e-135
6-phosphogluconolactonase 1 (.1)
Lus10037065
59 / 9e-12
AT1G13700 383 / 1e-135
6-phosphogluconolactonase 1 (.1)
PFAM info
Representative CDS sequence
>Potri.015G008401.1 pacid=42775469 polypeptide=Potri.015G008401.1.p locus=Potri.015G008401 ID=Potri.015G008401.1.v4.1 annot-version=v4.1
ATGTCATTTTTTGAGGAAATAAACTTATTTGTTTTTCCCTTGAATGTTTTGCAGGTACCAATCCTCCCAGGTGATGCTTATGCCATCAACAATGCCTTGT
CGGCTGAGGGTGCAGCTGATGATTATGAAACCTGTCTCAAACTTTTGGTTCATACTGGCGTGGTAGATAAATCTTCATTGAGTGGACTCCCAAAGTTTGA
TCGGATGCTTATCGGCACGGTCTGGATGGGCATGTAG
AA sequence
>Potri.015G008401.1 pacid=42775469 polypeptide=Potri.015G008401.1.p locus=Potri.015G008401 ID=Potri.015G008401.1.v4.1 annot-version=v4.1
MSFFEEINLFVFPLNVLQVPILPGDAYAINNALSAEGAADDYETCLKLLVHTGVVDKSSLSGLPKFDRMLIGTVWMGM
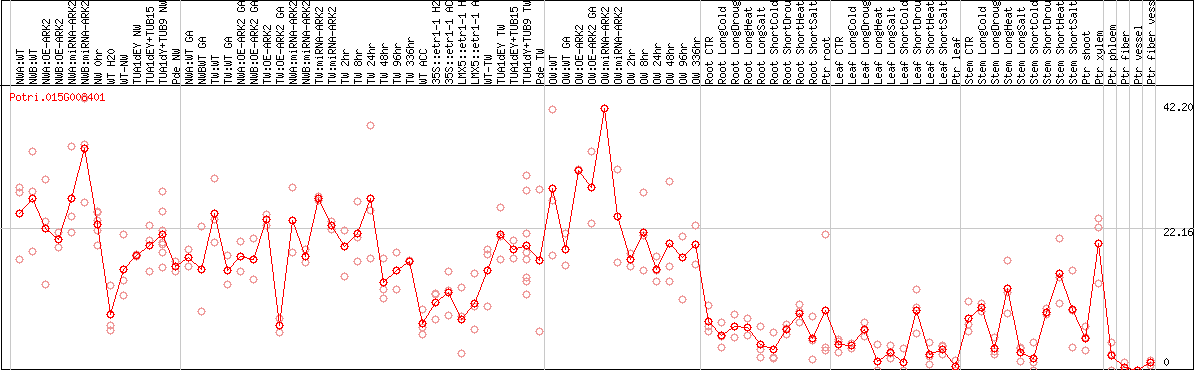
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.015G008401 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.