External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT3G17490 77 / 1e-15
F-box and associated interaction domains-containing protein (.1)
AT3G06240 67 / 3e-12
F-box family protein (.1)
AT4G12560 60 / 7e-10
CPR1, CPR30
CONSTITUTIVE EXPRESSER OF PR GENES 30, CONSTITUTIVE EXPRESSER OF PR GENES 1, F-box and associated interaction domains-containing protein (.1.2)
AT1G46984 56 / 1e-08
F-box family protein (.1)
AT3G07870 55 / 3e-08
F-box and associated interaction domains-containing protein (.1)
AT1G55070 54 / 5e-08
F-box and associated interaction domains-containing protein (.1)
AT1G32600 52 / 1e-07
F-box associated ubiquitination effector family protein (.1)
AT3G57590 52 / 2e-07
F-box and associated interaction domains-containing protein (.1)
AT5G47300 52 / 3e-07
F-box and associated interaction domains-containing protein (.1)
AT5G50220 48 / 5e-06
F-box family protein (.1)
Paralogs
Gene ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Potri.018G007900
142 / 3e-39
AT2G04920 63 / 1e-10
F-box and associated interaction domains-containing protein (.1)
Potri.006G013000
87 / 8e-19
AT4G12560 250 / 5e-79
CONSTITUTIVE EXPRESSER OF PR GENES 30, CONSTITUTIVE EXPRESSER OF PR GENES 1, F-box and associated interaction domains-containing protein (.1.2)
Potri.015G013800
82 / 2e-17
AT4G12560 105 / 7e-25
CONSTITUTIVE EXPRESSER OF PR GENES 30, CONSTITUTIVE EXPRESSER OF PR GENES 1, F-box and associated interaction domains-containing protein (.1.2)
Potri.012G014300
81 / 8e-17
AT3G16210 88 / 6e-19
F-box family protein (.1)
Potri.015G013500
76 / 3e-15
AT4G10190 70 / 4e-13
F-box and associated interaction domains-containing protein (.1)
Potri.011G121200
72 / 4e-14
AT4G12560 116 / 9e-29
CONSTITUTIVE EXPRESSER OF PR GENES 30, CONSTITUTIVE EXPRESSER OF PR GENES 1, F-box and associated interaction domains-containing protein (.1.2)
Potri.012G014700
72 / 6e-14
AT1G12170 66 / 1e-11
F-box family protein (.1)
Potri.017G058000
64 / 2e-11
AT3G06240 138 / 5e-37
F-box family protein (.1)
Potri.002G223000
63 / 8e-11
AT3G07870 467 / 5e-164
F-box and associated interaction domains-containing protein (.1)
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10006687
74 / 1e-14
AT4G12560 81 / 5e-17
CONSTITUTIVE EXPRESSER OF PR GENES 30, CONSTITUTIVE EXPRESSER OF PR GENES 1, F-box and associated interaction domains-containing protein (.1.2)
Lus10007848
62 / 1e-10
AT3G10430 73 / 5e-14
F-box and associated interaction domains-containing protein (.1)
Lus10018106
62 / 2e-10
AT3G06240 95 / 3e-21
F-box family protein (.1)
Lus10013873
62 / 2e-10
AT3G06240 104 / 2e-24
F-box family protein (.1)
Lus10037892
61 / 3e-10
AT3G06240 107 / 7e-26
F-box family protein (.1)
Lus10038606
61 / 4e-10
AT3G07870 112 / 1e-27
F-box and associated interaction domains-containing protein (.1)
Lus10038603
60 / 9e-10
AT3G06240 110 / 2e-26
F-box family protein (.1)
Lus10031020
59 / 1e-09
AT3G10430 88 / 3e-19
F-box and associated interaction domains-containing protein (.1)
Lus10026587
59 / 2e-09
AT3G06240 104 / 3e-24
F-box family protein (.1)
Lus10039024
58 / 3e-09
AT4G12560 333 / 3e-111
CONSTITUTIVE EXPRESSER OF PR GENES 30, CONSTITUTIVE EXPRESSER OF PR GENES 1, F-box and associated interaction domains-containing protein (.1.2)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0162
FBA
PF07734
FBA_1
F-box associated
Representative CDS sequence
>Potri.015G011000.2 pacid=42774753 polypeptide=Potri.015G011000.2.p locus=Potri.015G011000 ID=Potri.015G011000.2.v4.1 annot-version=v4.1
ATGCTCACACAGCGGCGAGAATTATATAGCATATCAGTGCTTCCCGGAGGAATAAATAGAGTTGATGATCGCAGCATGCCATTCGCACTGGAAGCGGGTA
AATATGCAGCAGAAATTGTGGGTTCTAGCAATGGTTTAGTTTGTTTAAGTATCCGTTCAAAGATATCTAATGATCTTAACGCTCATATATTATGGAATCC
AGCAACAAGGCAATACCGAGAGTTACCTCCTAACAGGATTTGTTACAGTCAAGCTCAAGGGTTTGGTTTCCATCACGGTATAAACGACTACAAACTGCTC
CAAGTTGCTTACAGAAAGAATGGCCAGCAGGAAGCTAAGGTTCTTGCATTGAGCACAGGTTCTTGGAGAAAGGTGGAGGACACTCTTCCTTCATATGATT
ACGGTATTGCAAAACCGGTGGTTGTTAAAGGAGTTTGGTATCACATGGCAACAGCTGCTCGAGAAACACCATTTATTATAAGGTTTGACATGGGCGACGA
CACCTTCTCCAAGGTGAAAACTGCACCTATACCTTATCATTCCTCTCCTGAATATCTCGTTAAGAAAGTGAAGTTGATGGAGTACAAAGAATTGCCTGCA
ATTTGCGTATTCGAATATAATTACTGGATCCCATATCCTAGTTTATCTTTCAAGATTGATATATGGTTAATGAACAACGAGCAGTCTTGGACAAAGGTAT
TGACATGTAGTCAGAATTCCGTTCCCATTTCTGATGTTGGCTTGCTGCCTTTAGGGTTCTGGGTTGATGAAGAAAGTATAATCTCGGAGAACTACAAGAG
GGATGGCTTGAGCTTGTTTAATCTCTCGAATAAGCAATCGAAAGTGGTTGTGGTAGATGAAGCCAAAAGGTTTGGAAGTGCTCATAGTTATGTGGAGAGC
ATGGTTTCAGTCTACCCTACCAGAAGTCAAGATAGGGACTCCACAAATTAA
AA sequence
>Potri.015G011000.2 pacid=42774753 polypeptide=Potri.015G011000.2.p locus=Potri.015G011000 ID=Potri.015G011000.2.v4.1 annot-version=v4.1
MLTQRRELYSISVLPGGINRVDDRSMPFALEAGKYAAEIVGSSNGLVCLSIRSKISNDLNAHILWNPATRQYRELPPNRICYSQAQGFGFHHGINDYKLL
QVAYRKNGQQEAKVLALSTGSWRKVEDTLPSYDYGIAKPVVVKGVWYHMATAARETPFIIRFDMGDDTFSKVKTAPIPYHSSPEYLVKKVKLMEYKELPA
ICVFEYNYWIPYPSLSFKIDIWLMNNEQSWTKVLTCSQNSVPISDVGLLPLGFWVDEESIISENYKRDGLSLFNLSNKQSKVVVVDEAKRFGSAHSYVES
MVSVYPTRSQDRDSTN
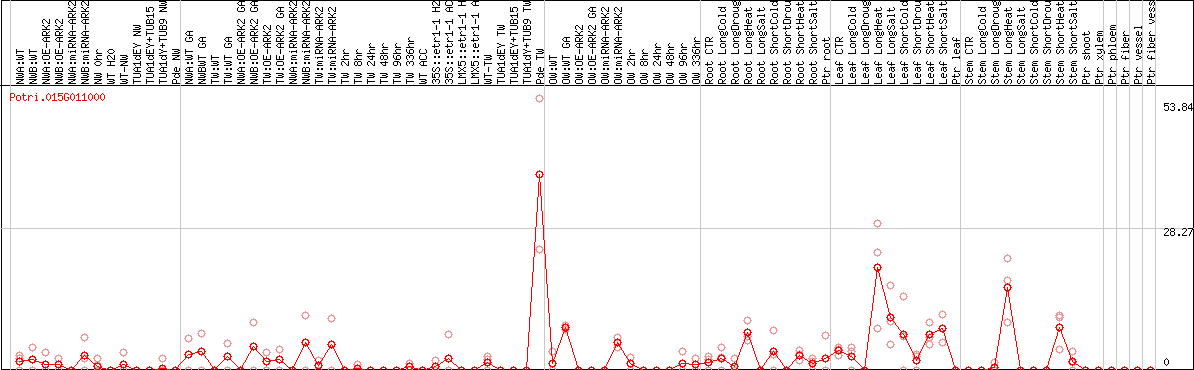
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.015G011000 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.