Potri.015G012800 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info |
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.015G012800.9 pacid=42775473 polypeptide=Potri.015G012800.9.p locus=Potri.015G012800 ID=Potri.015G012800.9.v4.1 annot-version=v4.1
ATGGTTGATGATGATGATGGGTACAGGAGGTGTTCAAGAAGTGCAGGCAAGTGGAGGTGTAGGGAGAGAGCTTTTAGTGGCAAAGCGTATTGTGAAAAAC
ACCATCTTTATTCAGTGGAGAGGGGTTTGAAGAGAAGCATGGAGAAGAAGAGTGGAAATGGAGGCGGGGTTGAAGTGGGTTCTCAGAAAAAGAAGAGACA
GAGGGAGGGAAGTGAAGAGGATTCAGGTATTTTGGCGGGAGAGAAGGAACAGAAGGTTAGTGATCATGTGGGTTTTGGAGATCAAGGGGTTCATGACTGG
TTTAGTGGAGGGAATTCAGATGTTTTGGAGTGGTTTGATGATGTAGGTGCTGGTAATGTTGGTGAAAGCTTTCAACTTTGGCAAAACGAAGCTACTGCAC
GTGGGAATAATGGGCAGGTTCTTGGTGGTGAGGGGTTTCAAGATAGTTTTGGTGGAGAATGTGGTGGTACTGAGGTTTTGGGTCTGGATGGTAAGGTTTT
TCAGGTTTGGGACGGTGAAGGATTTCGATTGGGCGAGTCTGGTCTGGGTGCTAGTGAAAATGGTGTTGTGGGTCTAGGTGATCAAGGGGCCCCAGACCTA
TTTTGTCAAGTTAGTGCTGCTAATGTTGGAAATGTGGGCGTGGGTTCTGGTGGTGGAGGAATACACGGGGTATTTGGTGAAGTCAATGGGAAAAGTGGTG
ATATAACCCTCTCTGGTGAAAGCATGGAGGGACTATTTGGTGAATGTAAAAATGGAGGTATGGCTGCTAGTGGTGAAGGGGTTCAATGCTGGTGTGGTGA
AGCTGGGTGTGCTAGTATTGACGGTGAGGGAATTCAGGGCCTTTTTGGTGAAACTACTTGTGAAAATGGGGGTGAAGGGATTGAAAGTGGATGTCATAGG
AATGGAGGTGGAGATGATGTTGAAGATGATGGGAAAACGAACAAGAAGGAAATTTTTGGTGTTGAAGCAGTTGGTATTGACGATTCAAGTGAGTTTGGTG
GTGATGACAACGGTGGAGCTGAGGTTTCTCCAGTGAAACGCAAGCGAGGGCGACCAAAGGGTTTAACGAAGAAGAAGAATGATCCTCGAGGTCAAGTAAA
ACTTGGCCTAGGTGGTGATAATGTGATCGGAAATGGTAGTGCCGTTGGGATGATTGAAATGGAAATGTTCAGTGAAACCAGTGGTGTCAATGGAGAAGGA
AATGAGAATAACCCAGTGGAGGATAAGCAAATCCTTGGTGCTGGAGAATGTATTGAGAGATCAGTTGGAAATGATATTGTGCGGCCAAAGGCAAAGCGTG
GCCGACCAAAAGGTTCAGGGAAGAAGCAAAAAGATGTTGCACCTGAAGAAAAACAGTGCTTGCCTGGTGATTTTCTGGGCAATAACAATGCTGGCATTGA
GTCTGTTACCCCGGCAACATTGGAGAATGGGAGGACTACTCTTCTATGCGCAGAGGACGATGAAACTCCTGGTGGAACTGAGACTGTCACACCAAAAAGG
CGTGGGAGACCAAAGGGATCAAAAAATAAGAATAATGGAGATACTGCAGGTGAGAATCAAGAAATGACTGGTGAAACTAAGGGTTGTATTGGTAGTGTTG
TATGGACTCCGATACTAATGGGTGTGGAGAGTGAGACTACTATTGCAGGGGCAGAGGATAGAGAAATTCCTGGTGAAGGCACTTGTGCTAATGGGAAAGT
AGAGGAGATCATCGCACCAAAGAAGCGTGGCAGACCAAAGGGTTCAAAAAAGGAGGAAATCCTTGCTGGAGATGATCGAGAAATGCCCGGTGAAACCAAG
GATACAACTGACTGTGACAATAGGACTGCTAGATCAATGCAGTTGGAGAGTGAGAGGACTACACTTGAAGGTGCAAAGGACAAAGAATTGCCTGATGAAG
TCACTGGTGGTAAAGGGATCATCACACCAAATAGGCGTGGCAGACCCAAGGGTTCAAAGAATAAGAAGGAAAATCTTGCCAGTGGTGAAGCCAAGGTGAG
TGTTGATAGTTGTGATAAGACTAATATGCCAAGGGTTTTGTGGAATGAGAGGATTACTCTTTTAGATGAAGAAGAAAATGGAGGCAGTCACGAATTGCCC
CGTGGTATTGTGGGTTATAATGAAAGTCGTGATAACAAAGTCAGGCTGGCAGGTTGTGTGAATGACATGACTTGTTTAGGGGAAAATGATGGGGCAATGG
CTGATGATAATATTGACAGTAGTTTTGGAGTGAATGATAAAAGCAAACCTAAGCGTAAGCGAGGGCGACCAAAGAACTCGAAAAATAAGCAAAAATCTCT
TGCTGGTTACAGGCAACAATTTCCCGGTGATATCATGGTTAGCAATGATGGGGCTGATAATAAAGTTAGGTTGATGGGTTTTGAGAATTGGACGACTGCT
ACTTTTGATGGATATAAAGCTTTGTGTAGTCACGCCACTATTGAAGGTGCTGACAACAAAGCTAGTCTGCTAGATTTAGGGAATGGGATGGCTGCTCTTG
TGGGTGAAAAAGCTAGGAAATTGCCTAGTGAGGCCACTGGTGTTAGTGGAGATGGTCAGGAAATCATAAAGCCTAGTCTTGACCAAAGAGTGGGCTCCAA
GAACAAGCCAAAAAATCCAGCAGGTGATAGTTGGGAATTGCCTAGTGACATTGTGGATAGAAATGGAGATGCGGGCAATGTAATCAGGGAGGTGGTTTTA
GAGAATAGGATGGTTGTTTCTTCATGTGAAGGACATCGGGTATTGTCTGTTGAGGTCACTGGCCATAGCAGAGAGGGAAATGAAATTAGAAAGCCTAGGC
CACGCGGCCGACCAAAGGGCTCAGTGAAGAAGAAAAATCTTGCTGATGGTCACCAGGAATTGCCTGGGGAAGTTATGAACGGTAATGATGGCAAGGACAC
AGTTAGTCTGATGGGTTCAGAGAATAGGATGACTGCTCTTTTAGGTGAAGAGTATTGGGTATTGCCTGGTGTGGCCACTGGCCATAGTGGAGAGGCGTAT
GATAATATAAAGTCTAGGCGTGGTCGTGGCCGACCAAAGGGCTCTGGGAACAAGAAGAAAAATCTTGCAGGTGGTGGTAATCAGAAATTGGCTGGTGTAG
TTATGTATGGTAATGATGGAGAGAATGCTGTTAGGTCGATGGGTTTAGGGAATGGGATGACTACTCTTATTGGAGAAAAACATAGAGCACTGACTGGTGA
GTCCACTGGCCATAGTGAAGAGGGGGAAATTCCTAAGCGTAAGCCAGGTCGTCCAAAGGGGTCAAAGAACAAGAAAACTCTTGCAGGTTGTATTCAGGGA
TTGCCTGGTGAAACCATGTGCGGTAATGATGCTGGTGAGAAAACACTGAGGCTAACTAGGATAGATAATGGTTGGGCAGCTCCTTTAGGCAAGGAAGTTA
AGTTGGTGCCTTGTGAGGTCAATGGTGTTAGTAGAGATGGAAATGATACCATGAAGACAAATGTTAAGCATGACCAACCTAAGAGTCTGAAAATAAGGAA
GGAGGGTGACGGAAATGAGGAAATCCAGTGCAAAATCAAGTGTAAAAGTGATGGTGAGGATAGTATTATTTGTCTAGCAGGTTCTGAGAGTGAAGGGAGT
ATGCTTGAAGGTGAGGAAGATAGGATAATAGCAACTGAAGCTGCTGGTGGTAATGAAGCTGGCACTGCAAATCCCCAGTCTAAGATTGAGTGTGCTCAAC
CTGAAGCTTCAAAGAGAAAGAAGCTAAGTATTGCTGCTAAAGAAGAAGAGAGACAGAATGGAGAATTTATGGGCAAAGACGATGGTGAAAGTAAAAGGCC
AAACAATAAGCAAGTCAGACGAAAAGTCTTAAAGAGTAAGAGAACTATTCTGTTAGCTAAATCTTTTGACCGTATATTAAGGCAGAAGTATGGAATGAAG
AAGGAAAGTGGGGAAGATTTGGGAATGGACAGGGATATTCTGGTTGAGCAAACTGGTCACTGGAGCAATATTAAGAAGAGGCCTAGGGGAAGGCCACCAA
AGCACAACCGATCAGAAAATTCAAATTTACTAGGTGCCAACAAGAAAAACGAACAGAAGACCTTGATGTGCCACCAATGCTGCAGGAATAATAGGAGTGG
TGTTGTCATTTGTTCTAATTGCAAAAGAAAGCGCTATTGCTATGAGTGCCTTGCGAAGTGGTACCCTAAGAGAACACACGAGGAGATAGAGATTGCATGC
CCATTTTGTCGTGGTAATTGCAATTGCAGGGTATGCCTCAAGGAAGATGTAGTTGTGGTCGCTGGTGATGATAAAGCTGATGCAAATGCCAAATTGCAAA
AGCTTCTTTACCTGCTACACAAGACTTTACCTCTCCTCAGGCATATTCAAAGGGAACAGAACTCTGAAATATATGTAGACTCCAGGATCCATGGTAGCCT
ATTAACAGAAGAGCATGTAACAAAATCTTTACTTGACGATGATGATCGAGTATATTGTGACAATTGCAGTACATCTATTGTTAATTTCCACAGAAGCTGT
CCCAATCCTGATTGTTCCTATGATCTCTGTCTCACTTGTTGTTCGGAACTCAGAATAGGATTTAAGCCTGGAGGTAATGAAGCAGAATCTTCTCACCTAC
GATTTTTTGAAAGAGTAGACAGTCAAGGTGCGCTTGTTCATGACCAGATAAATGAAAATGGAAAAGGACTTGGTTGCAAGACTCAAGTGTCTGATTTGGA
AAGTAAGTGCACAGCTGATATGTCATGCAAGTTTCCTGATTGGAGAGCAGAGAGTGATGGAAGGATTCCTTGCCCTCCTAAAGAACTGGGAGGCTGTGGT
AACGAGATACTCACCTTGAGACGTATCTTTGATGCTAAATTCGTTGAAGAGATGATTAAGAGTGCAGAGGAACTTACTCTTAACTATCAGTCCCCAGATA
TTAGACTTTGTGAAGAATGTTATTTGTGCCATCCTACCAGTTCCACTGAAAATGGATCTAAAGATTTTGCAGTGAGGAAAGCAGCTTATAGGGAGAACAG
TGATGATAACTTTCTGTACTGCCCAAATGCACTTCAGTTGGGTGATGATGACTTTGAACATTTTCAATTGCATTGGATGAGAGGTGAACCTGTTATAGTT
AGACATGCACTTGAAAGGACTTCTGGTCTGAGCTGGGAACCAATGGTCATGTGGAGGGCTTTTAAAGGTGCAGAAAAGATAATAAAAGAGGAAGCACACA
GGGTTAAAGCCATTGATTGCCTGGATTGGTGCGAGGTTCAGGTTAATATTTTCCAGTTTTTCAAGGGCTATTTAGAGGGTCGTAGCTATAGGAATGGTTG
GCCAGAGATGCTGAAGCTGAAGGATTGGCCCCCATCAAATTTCTTTGAAGAATGTTTGCCAAGGCATGGTGCTGAGTATGTTTCTATGCTCCCCTTCAGT
GAGTATACCCATCCAAAATCTGGCATTCTGAATATGGCTACAAAGCTTCCTGCTGTTTTGAAGCCAGACTTGGGGCCTAAAACTTACATAGCATATGGAT
TTGTGGAAGAACTTGGTAGAGGTGATTCAGTGACAAAACTGCATTGTGACATGTCTGATGCGGTCAATATACTTACTCATATGACAGAAGTGAAAGTTCC
TCGATGGCAAAGCAAAATTATAAAAAAAATACAAAAGCAACATGAAGCTGAAGATATGAATCCGGTTTGTGGTGGGATACAGAAAGTGACTCGTAAGTCT
GGAAGGAAGCCACGAAAAAGAAGGCGGAAAGTTGAGAAAATGGATCCTGAATTACCTAAAAAAGATGAAAATATCGAGAGTGACTCCTCACTAGAGCGGT
TGTATGTCCAGGAACAGAAACTGGAAGAACAAAAAAGCATGTGCCAAGAGTTGGGGGAATTTTATAGCATTGTGTGTGGGATAAGATGTTCTTCAACTAA
ATCTGAGGTGACTGCTGATACCAATTTGCAGCCAGTTGCAAAAATGAATGCAAGGGTGCAAAATTATGATACGAGCTCTGCTGATTTAAATGAAAGTGTG
AACAGAGATTGTACTGAAGGAAACCACACCTCAGAACTTGTTTATGGTGGTGCTGTTTGGGACATTTTCCGTCGGCAGGATGTGCCCAAGTTAATTGAAT
ACTTGAAGAGGCATCAGAAAGAATTTCGCCATGTCAGTAGCCTGCCTGTAAATACTGTTATTCATCCCATCCATGACCAAACCTTTTATCTAAGTGAGAA
ACACAAAAGGCAGCTGAAGGAGGAGTTCAATGTTGAACCTTGGACATTTGAGCAGCACCTTGGTGAGGCTGTTTTCATACCTGCAGGATGTCCTCATCAA
GTGAGGAATAGACAGTCATGCATAAAAGTGGCCCTTGATTTTGTATCCCCTGAGAATGTTCAAGAATGCATTCGGTTGACAGAGGAGTTTCGTTTGCTTC
CAAAGACCCACAGAGCCAAGGAAGACAAATTGGAGGTGAAAAAGATGGCACTATATGCTGCCAGCGCTGCTGTAACTGAAGCCAAAAATCTGAATTCTTG
GGTAACTGAAGCCAAAAATCTGACTTCTTGA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.015G012800.9 pacid=42775473 polypeptide=Potri.015G012800.9.p locus=Potri.015G012800 ID=Potri.015G012800.9.v4.1 annot-version=v4.1
MVDDDDGYRRCSRSAGKWRCRERAFSGKAYCEKHHLYSVERGLKRSMEKKSGNGGGVEVGSQKKKRQREGSEEDSGILAGEKEQKVSDHVGFGDQGVHDW
FSGGNSDVLEWFDDVGAGNVGESFQLWQNEATARGNNGQVLGGEGFQDSFGGECGGTEVLGLDGKVFQVWDGEGFRLGESGLGASENGVVGLGDQGAPDL
FCQVSAANVGNVGVGSGGGGIHGVFGEVNGKSGDITLSGESMEGLFGECKNGGMAASGEGVQCWCGEAGCASIDGEGIQGLFGETTCENGGEGIESGCHR
NGGGDDVEDDGKTNKKEIFGVEAVGIDDSSEFGGDDNGGAEVSPVKRKRGRPKGLTKKKNDPRGQVKLGLGGDNVIGNGSAVGMIEMEMFSETSGVNGEG
NENNPVEDKQILGAGECIERSVGNDIVRPKAKRGRPKGSGKKQKDVAPEEKQCLPGDFLGNNNAGIESVTPATLENGRTTLLCAEDDETPGGTETVTPKR
RGRPKGSKNKNNGDTAGENQEMTGETKGCIGSVVWTPILMGVESETTIAGAEDREIPGEGTCANGKVEEIIAPKKRGRPKGSKKEEILAGDDREMPGETK
DTTDCDNRTARSMQLESERTTLEGAKDKELPDEVTGGKGIITPNRRGRPKGSKNKKENLASGEAKVSVDSCDKTNMPRVLWNERITLLDEEENGGSHELP
RGIVGYNESRDNKVRLAGCVNDMTCLGENDGAMADDNIDSSFGVNDKSKPKRKRGRPKNSKNKQKSLAGYRQQFPGDIMVSNDGADNKVRLMGFENWTTA
TFDGYKALCSHATIEGADNKASLLDLGNGMAALVGEKARKLPSEATGVSGDGQEIIKPSLDQRVGSKNKPKNPAGDSWELPSDIVDRNGDAGNVIREVVL
ENRMVVSSCEGHRVLSVEVTGHSREGNEIRKPRPRGRPKGSVKKKNLADGHQELPGEVMNGNDGKDTVSLMGSENRMTALLGEEYWVLPGVATGHSGEAY
DNIKSRRGRGRPKGSGNKKKNLAGGGNQKLAGVVMYGNDGENAVRSMGLGNGMTTLIGEKHRALTGESTGHSEEGEIPKRKPGRPKGSKNKKTLAGCIQG
LPGETMCGNDAGEKTLRLTRIDNGWAAPLGKEVKLVPCEVNGVSRDGNDTMKTNVKHDQPKSLKIRKEGDGNEEIQCKIKCKSDGEDSIICLAGSESEGS
MLEGEEDRIIATEAAGGNEAGTANPQSKIECAQPEASKRKKLSIAAKEEERQNGEFMGKDDGESKRPNNKQVRRKVLKSKRTILLAKSFDRILRQKYGMK
KESGEDLGMDRDILVEQTGHWSNIKKRPRGRPPKHNRSENSNLLGANKKNEQKTLMCHQCCRNNRSGVVICSNCKRKRYCYECLAKWYPKRTHEEIEIAC
PFCRGNCNCRVCLKEDVVVVAGDDKADANAKLQKLLYLLHKTLPLLRHIQREQNSEIYVDSRIHGSLLTEEHVTKSLLDDDDRVYCDNCSTSIVNFHRSC
PNPDCSYDLCLTCCSELRIGFKPGGNEAESSHLRFFERVDSQGALVHDQINENGKGLGCKTQVSDLESKCTADMSCKFPDWRAESDGRIPCPPKELGGCG
NEILTLRRIFDAKFVEEMIKSAEELTLNYQSPDIRLCEECYLCHPTSSTENGSKDFAVRKAAYRENSDDNFLYCPNALQLGDDDFEHFQLHWMRGEPVIV
RHALERTSGLSWEPMVMWRAFKGAEKIIKEEAHRVKAIDCLDWCEVQVNIFQFFKGYLEGRSYRNGWPEMLKLKDWPPSNFFEECLPRHGAEYVSMLPFS
EYTHPKSGILNMATKLPAVLKPDLGPKTYIAYGFVEELGRGDSVTKLHCDMSDAVNILTHMTEVKVPRWQSKIIKKIQKQHEAEDMNPVCGGIQKVTRKS
GRKPRKRRRKVEKMDPELPKKDENIESDSSLERLYVQEQKLEEQKSMCQELGEFYSIVCGIRCSSTKSEVTADTNLQPVAKMNARVQNYDTSSADLNESV
NRDCTEGNHTSELVYGGAVWDIFRRQDVPKLIEYLKRHQKEFRHVSSLPVNTVIHPIHDQTFYLSEKHKRQLKEEFNVEPWTFEQHLGEAVFIPAGCPHQ
VRNRQSCIKVALDFVSPENVQECIRLTEEFRLLPKTHRAKEDKLEVKKMALYAASAAVTEAKNLNSWVTEAKNLTS
|
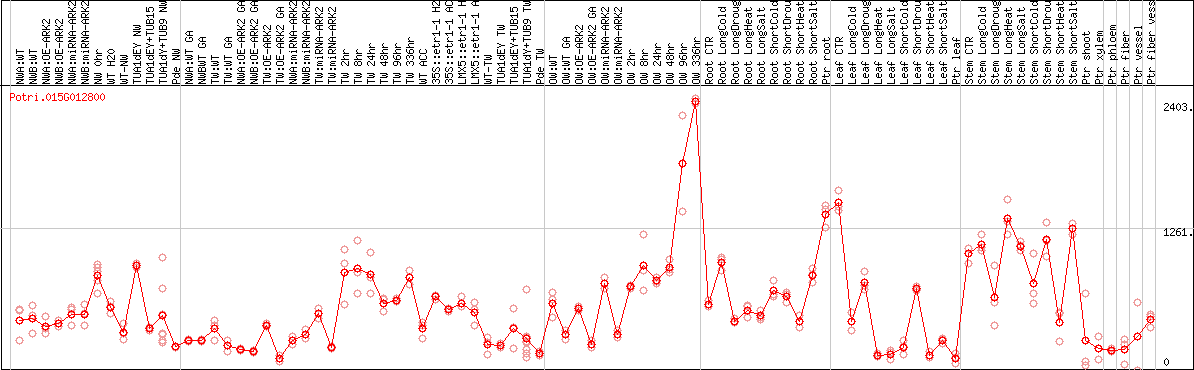
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.015G012800 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
