External link
Symbol
Arabidopsis homologues
Locus ID
BLAST score/e-value
TF class
Alias
TAIR10 short description
AT1G34390 52 / 1e-08
ARF
ARF22
auxin response factor 22 (.1)
AT4G01500 49 / 1e-07
B3
NGA4
NGATHA4, AP2/B3-like transcriptional factor family protein (.1)
AT1G43950 46 / 1e-06
ARF
ARF23
auxin response factor 23 (.1)
AT4G21550 46 / 1e-06
B3
HSI2-L2, VAL3
VP1/ABI3-like 3 (.1)
AT1G16640 44 / 2e-06
B3
AP2/B3-like transcriptional factor family protein (.1)
AT1G35540 45 / 4e-06
ARF
ARF14
auxin response factor 14 (.1)
AT1G35520 44 / 6e-06
ARF
ARF15
auxin response factor 15 (.1)
AT1G35240 43 / 1e-05
ARF
ARF20
auxin response factor 20 (.1)
AT1G13260 42 / 4e-05
AP2_ERF
EDF4, RAV1
ETHYLENE RESPONSE DNA BINDING FACTOR 4, related to ABI3/VP1 1 (.1)
AT1G77850 42 / 5e-05
ARF
ARF17
auxin response factor 17 (.1)
Paralogs
Locus ID
BLAST score/e-value
At best hit
BLAST score/e-value
TAIR10 short description
Lus10030240
47 / 8e-07
AT2G46530 381 / 1e-126
auxin response factor 11 (.1.2.3)
Lus10021326
44 / 1e-05
AT5G06250 202 / 2e-63
AP2/B3-like transcriptional factor family protein (.1.2)
Lus10005930
43 / 2e-05
AT5G62000 361 / 2e-112
ORESARA 14, HLS1 SUPPRESSOR, ARF1-BINDING PROTEIN, auxin response factor 2 (.1.2.3.4)
Lus10026510
43 / 3e-05
AT4G30080 694 / 0.0
auxin response factor 16 (.1)
Lus10019940
42 / 3e-05
AT4G30080 660 / 0.0
auxin response factor 16 (.1)
Lus10040906
42 / 3e-05
AT5G62000 367 / 2e-114
ORESARA 14, HLS1 SUPPRESSOR, ARF1-BINDING PROTEIN, auxin response factor 2 (.1.2.3.4)
Lus10007440
41 / 7e-05
AT1G59750 917 / 0.0
auxin response factor 1 (.1.2.3.4)
Lus10024754
41 / 8e-05
AT1G77850 505 / 1e-173
auxin response factor 17 (.1)
Lus10004226
41 / 8e-05
AT5G06250 183 / 4e-55
AP2/B3-like transcriptional factor family protein (.1.2)
Lus10005264
41 / 9e-05
AT2G33860 580 / 0.0
ETTIN, AUXIN RESPONSE TRANSCRIPTION FACTOR 3, Transcriptional factor B3 family protein / auxin-responsive factor AUX/IAA-related (.1)
PFAM info
Clan ID
Clan name
Pfam ID
Pfam name
Pfam description
CL0405
DNA_b-psBarrel
PF02362
B3
B3 DNA binding domain
Representative CDS sequence
>Potri.015G017000.1 pacid=42775178 polypeptide=Potri.015G017000.1.p locus=Potri.015G017000 ID=Potri.015G017000.1.v4.1 annot-version=v4.1
ATGGCAGCGTTCTCCAAGATCCTCAGAGAGACTGATACTAAGAAGAGGCTGTCAGTTCCTATCAGGTTTCTCAGGTCTCTTCCACCTTTCAAACTAGGCA
GCCATGCCGTCACGTTTGAAGCTACAGACGAGAAAGGCGAAGCTTGGCCCTTTCAATGCTCCATCCGCAAGAGAGGACACCCGAAGCCAGTCCTCACAAA
AGGTTGGGTTGCGTTCGTTAAGAGCAAGAAGCTACAAGTTGGTGACAAGGTCAGGTTTATTAAACGTAAGAACCGAGCTACTGCAGCAATCTCTTACATA
GTTCGTGCTGAAAAGGCAATCAAGATCTTTGGTGTTACATTTGGCTATGCCCGGATTGATCTAGTCCATTTCCCTAATTAA
AA sequence
>Potri.015G017000.1 pacid=42775178 polypeptide=Potri.015G017000.1.p locus=Potri.015G017000 ID=Potri.015G017000.1.v4.1 annot-version=v4.1
MAAFSKILRETDTKKRLSVPIRFLRSLPPFKLGSHAVTFEATDEKGEAWPFQCSIRKRGHPKPVLTKGWVAFVKSKKLQVGDKVRFIKRKNRATAAISYI
VRAEKAIKIFGVTFGYARIDLVHFPN
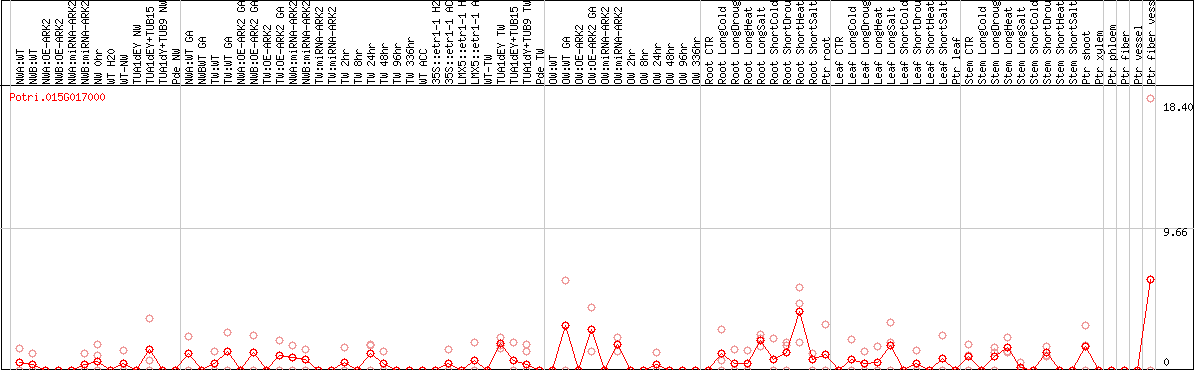
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.015G017000 coexpression network
*The number of genes in the network is adjusted within 50 genes.
*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.