Potri.015G020400 [POPLAR]

| External link |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Symbol | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Arabidopsis homologues
|
| |||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Paralogs
|
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Flax homologues |
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
| PFAM info | ||||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
Representative CDS sequence |
>Potri.015G020400.1 pacid=42775470 polypeptide=Potri.015G020400.1.p locus=Potri.015G020400 ID=Potri.015G020400.1.v4.1 annot-version=v4.1
ATGGGTGGTCTAGAGGGCTGGGTTCAGCCCAGTGGCTTTTCCCCTAATGGTCTCTTGCCTAATGAAGTTGCCTCTGTAACTCAAGCACTTGAGCCAGAGA
GATGGGCAACTGCTGAAGAGAGAACTGCAGAGCTAATTGCTTGTATTCAGCCCAATCAACCATCCGAGGAGCGCCGAAATGCTGTTTTATGCTATGTTCA
GCGCCTTATAATGAACTGCTTCCCCTGTCAGGTGTTCACCTTTGGGTCTGTGCCCCTCAAGACCTATCTACCTGATGGAGATATTGATTTAACAGTGTTT
ACTGAGAACCAGAATTTGAAGGAAACATGGGCCAATGAGGTACGGGATATCTTAGAGCACGAGGAGAAGAATGAGAACGCTGAATTCCATGTGAAAGAGG
TTCAGTATATCCAGGCAGAGGTGAAGATCATAAAGTGTCTTGTGGAAAATATTGTAGTAGACATATCCTTTGACCAGCTTGGTGGTTTGTGTACCCTTTG
CTTCCTTGAGGAGGTTGATCAATTGATAGACCAAAACCATCTATTCAAGCGCAGCATTATATTGATTAAAGCTTGGTGTTACTATGAGAGCCGCATCTTG
GGTGCTCACCATGGCCTTATTTCAACTTACGCTCTGGAGACATTGGTTTTGTACATATTCCACGTGTTCAATAATAAATTTGCTGGGCCACTTGAGGTGC
TCTACCGCTTTCTGGAATTTTTCAGTAAATTTGACTGGGAGAACTATTGCATTAGCCTCTGGGGCCCAGTTCCAATTAGTTCACTTCCTGATATGACTGC
GCTCTCTCCACGGAAGGATGGTGGCCAGATACTTCTCAGCAAATTGTTTCTAGATGTCTGCAGCTCTGTGTATGCTGTCTTTCCTAGCCGGCAGGAAAAT
CAGGAGCAATCCTTTGTTTCCAAATATTTTAATGTCATCGATCCTTTGCGTGCAAATAACAATCTTGGCCGTAGTGTCAATAAGGGTAACTTCTTTAGGA
TTCGCAGTGCTTTTGCTTTTGGAGCTCAACGTCTGGCCAGATTGCTTGACTGTCCAAAAGAGAATCTGCTTGCTGAATTCAACCAGTTTTTCTTGAACAC
ATGGGACAGGCATGGAAAAGGTCATCGCCCTGATGCTCCAAGTTCCAATCACGTTGTTCAGCGACCAATCAGGTCTAATGTAATTGATGGGTCTGAGACC
ATCATAAATTATTCAAGCAGCAAAAAGACAAAAGAGGATCCTTCTGGCCATCAATCTAAGGTCGGGGTGACTTATGCTGCTCATGCATCTGACAGTGTTT
CCTCCCAGCATGATAACCGCAGTTTAAAACGAACATCCAGGCCTGGTAATATATCAGCTATTTCTGGCACTCGAGGCCAGAGCATGCAAGCCAACTTGAC
TAACTCCATGGCCTCTGATCAGAATAATAAAAAATTAATATATAATAGTTTGAATGAGAATGCACATAATGAAAAGATGACAAGTTCCGGAACATATTAC
TTGGGGAATGAAGTGAATGCTAGGTATCAATTTGCAAGGACTCAATCCAGTCCGGAACTAACAGATACATCCAGTGAAGTCTTATCTCGAGGGAGGCATA
ACAGGGCATCAGAGACAGTAAATGGGCAGACTGCTCCTGCAAGGTCACACAATAGCAGGAGAAGAAATTTGGTCCCTGAGGTTTTGGAGAACCATGGTGC
AAGATTTTCAACTGAAGACTCACTATCTTCAAGGCATAGCCTATCCCATCAGAGCATTGATGCTGCTGTTGATTCTACTAGTGCTTCAAATAGCTATTTT
GGTGATTCTGGCGAAGGAACCATGGAAGACCACCTTTCTCTCTCAGAGACAACGCAATTGCATCAGGAAGAGCAAGACCGTGTTAGCGTGGCATCTTTCA
GTGGTTATAGTGTCAGTGAACAGGGTCAGATGCCCATGAATTTAGCTTCTGGTCAACCACCTTTTGCTTTACCACCTTCTGTATTAGCTTCCCTTGGATA
TGCTCAAAAACATATGACTGGTACAGCTCCCATAAATGCTCCTTCATTTGAGTCCCCTTGGGTGTCGAACGTGCACTATCCTCAAGGTTTTATTCCAAAT
CCAGTATCCCAATATTTTCCAAGCATGGGAATGACCTCAGATCAAGAAGTAACAATTGAAATGATTGATGCTAAATTGGCATCTACAGAGTTGAGTCAAG
AAGAGAGTGATCATGGTTGGTCAAAGCCAGATGCAGATTCTGTAAGACATCAACAGAAGAACAGAAGCTCCCAATCTCGACTGCAGGAGCATAGGCAACC
TCTTGCTTCTGTTGAGTCTAACCATGTCCATTCATCTCGTGTAAGCAGGTCCGGCAGTTTCTCTCCAAGGGATCGCGGGTTAATTACAGAAGACAGAGGT
CTGATTAGGGAAAATTACAGTGACGATGCTCAATATCAAATACCTAAAGAGTCTGATGCTTATTCCTCAGCTGATTTAAGATTTGTACCTTCTTCAGAAG
CATCGTCCTCAGGAAGCAAATCTGAAGATAATGGTGATGGATTACTTTTGAGGACTTATAAGTCAACAAAGGATCGGCGGGGAAAGAAATCTGTTCCTTC
TACAGATTCTTCTATTGCATATGGGATGGATAAGAATGAAAGGCAACGTGAAGACAAATCAGTTGATCACATATCTCTTCAGCCAGATGAGGACAACAGA
GAGTGGATTTCACTTTCAACCATGGGCACTGAACTGTCAGAAAGCATGGTTTCTGGTGTTGGTGCTTCACATGTTTGGAATCATCAAATACAAAGTTATG
ATCCGGCATCAGTGAACAGATCAAATTCAATGCTTTCAGTTGCTCCAATGTTTGTGGGTCCCAATTCTCATCAGAGACCTAATGATAACCATGGAGCACT
ACCCTTTGCATTTTATCCCACAGGACCACCGGTGCCTTTTCTAGCCATGGTTCCAGTATACAATGTTCCAACCGAAGCAGCAACATCTAGTGTATCCACC
AGAAAACTCGATATGGATGAAGAATTTGATAGCTCACAAAACAATCACTCGAACCAAAGTTTGGATTCATCTGAGAATGTTGATTGGTCAGAGATTTTAA
ATACTTATACATCTGTGAATAATGCTTCTTCCTCGGTGCATTCTGAACAGGGCAGATCTGAAATTTTGAATAGTGATTTTGCTAGCCACTGGCAGAATCT
ACAATATGGGCGGTTTTGCCAGAATGCTCGAAATAATGATTCCTTACCTCATCCATCTCCTGTTGTAGCACCGCCAATGTACTTGCAGGGACATTTTCCA
TGGGATGGTCCTGGAAGACCTCCTGCAGACATGAATCTCTTAACTCAACATATGAATGGTCCTCATCTTATTCCCGTTGCTCCTGTCCATCCAGGATCTA
TCAGGCAGCCTGGCTTTTATCAACATTATGCTGATAATATTCCTAGATACCGTGCTGGAACTGGAACTTACCTGCCCAATCCGAAAATATCTTACAGGGA
TCGACAATTTTCCAATTCCAGAAACCATAGGGGAAATAACAACTATGACAGGAAGGACCACCATGATGACAGGGAAGGAAACTGGAATAACAACTCAAGA
CCACGATTTGGTGGTCGCAGCCAAAACCAGAATCAAGTTGAGAAGCCATCCTTTAGGATGGATAGGAGTACACCAAATAATCGTCGTTCTGATAGGTCAT
GGAACTCAAAGCAAGATCCTCTTCCACCACACCATCCCCGGAATGGTTCATTTAGTTTCTCAAACTCCACAAACCGAGGTTCTACCAATGCGGCATATGG
CATGTATCCGCCAATACCAGTTGTGAATCATAGTGAAGGTTCTGCATCAGTAGTTATGCTTTATCCATATGACCGAAACATGGGCTACAGTTCTGCTGGT
GAGCATGTAGAATTTGGGTCCCTTGGGCCAGCTCATTTAACCGGTGACAAGGCATCACCTCATCTTGGTGAAGAGTCATCAAGAGGCATTAATGAGCAAC
AAGATTTTCAAAGAGTTTCTGATCTTTCTTCTCCAGATCAGCCAGCATCACCCCTGGTTCGCAGGAGGGTTTAA
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||
|
AA sequence
|
>Potri.015G020400.1 pacid=42775470 polypeptide=Potri.015G020400.1.p locus=Potri.015G020400 ID=Potri.015G020400.1.v4.1 annot-version=v4.1
MGGLEGWVQPSGFSPNGLLPNEVASVTQALEPERWATAEERTAELIACIQPNQPSEERRNAVLCYVQRLIMNCFPCQVFTFGSVPLKTYLPDGDIDLTVF
TENQNLKETWANEVRDILEHEEKNENAEFHVKEVQYIQAEVKIIKCLVENIVVDISFDQLGGLCTLCFLEEVDQLIDQNHLFKRSIILIKAWCYYESRIL
GAHHGLISTYALETLVLYIFHVFNNKFAGPLEVLYRFLEFFSKFDWENYCISLWGPVPISSLPDMTALSPRKDGGQILLSKLFLDVCSSVYAVFPSRQEN
QEQSFVSKYFNVIDPLRANNNLGRSVNKGNFFRIRSAFAFGAQRLARLLDCPKENLLAEFNQFFLNTWDRHGKGHRPDAPSSNHVVQRPIRSNVIDGSET
IINYSSSKKTKEDPSGHQSKVGVTYAAHASDSVSSQHDNRSLKRTSRPGNISAISGTRGQSMQANLTNSMASDQNNKKLIYNSLNENAHNEKMTSSGTYY
LGNEVNARYQFARTQSSPELTDTSSEVLSRGRHNRASETVNGQTAPARSHNSRRRNLVPEVLENHGARFSTEDSLSSRHSLSHQSIDAAVDSTSASNSYF
GDSGEGTMEDHLSLSETTQLHQEEQDRVSVASFSGYSVSEQGQMPMNLASGQPPFALPPSVLASLGYAQKHMTGTAPINAPSFESPWVSNVHYPQGFIPN
PVSQYFPSMGMTSDQEVTIEMIDAKLASTELSQEESDHGWSKPDADSVRHQQKNRSSQSRLQEHRQPLASVESNHVHSSRVSRSGSFSPRDRGLITEDRG
LIRENYSDDAQYQIPKESDAYSSADLRFVPSSEASSSGSKSEDNGDGLLLRTYKSTKDRRGKKSVPSTDSSIAYGMDKNERQREDKSVDHISLQPDEDNR
EWISLSTMGTELSESMVSGVGASHVWNHQIQSYDPASVNRSNSMLSVAPMFVGPNSHQRPNDNHGALPFAFYPTGPPVPFLAMVPVYNVPTEAATSSVST
RKLDMDEEFDSSQNNHSNQSLDSSENVDWSEILNTYTSVNNASSSVHSEQGRSEILNSDFASHWQNLQYGRFCQNARNNDSLPHPSPVVAPPMYLQGHFP
WDGPGRPPADMNLLTQHMNGPHLIPVAPVHPGSIRQPGFYQHYADNIPRYRAGTGTYLPNPKISYRDRQFSNSRNHRGNNNYDRKDHHDDREGNWNNNSR
PRFGGRSQNQNQVEKPSFRMDRSTPNNRRSDRSWNSKQDPLPPHHPRNGSFSFSNSTNRGSTNAAYGMYPPIPVVNHSEGSASVVMLYPYDRNMGYSSAG
EHVEFGSLGPAHLTGDKASPHLGEESSRGINEQQDFQRVSDLSSPDQPASPLVRRRV
|
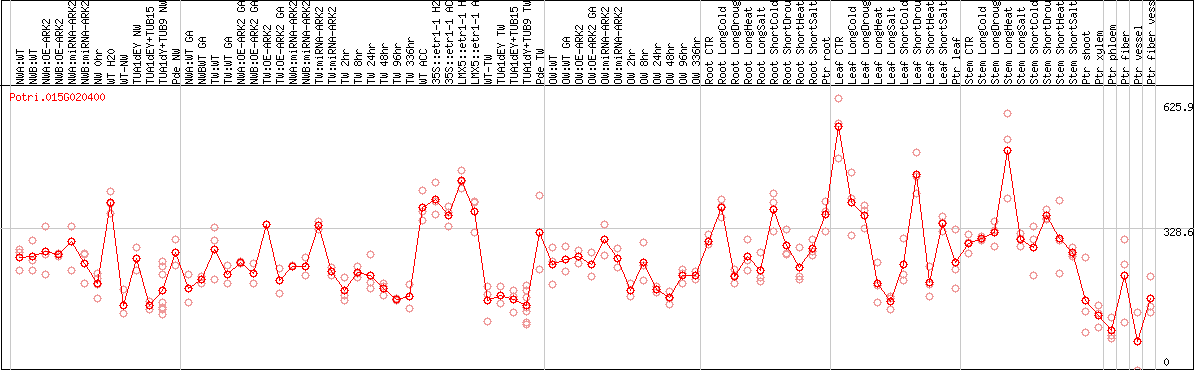
DESeq2's median of ratios [POPLAR]
Coexpressed genes
Potri.015G020400 coexpression network
*The number of genes in the network is adjusted within 50 genes.*Gene name represents symbol(s) of closest Arabidopsis gene if symbol(s) for the gene itself doesn't exist.
*Circle diameter represents the number of connection with other genes within this network.
*Color for gene name represents subnetwork based on the result of network clustering.
